External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G12480 95 / 1e-23
PEARLI 1 1, PEARLI 1, pEARLI1
EARLY ARABIDOPSIS ALUMINUM INDUCED 1, Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12490 95 / 1e-23
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12500 94 / 3e-23
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12510 92 / 4e-23
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12520 92 / 4e-23
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT2G45180 92 / 4e-23
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12470 92 / 1e-22
AZI1
azelaic acid induced 1 (.1)
AT4G12530 91 / 1e-22
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12550 90 / 2e-22
AIR1
Auxin-Induced in Root cultures 1 (.1)
AT4G00165 89 / 6e-22
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G128800
153 / 2e-46
AT4G12480 109 / 5e-31
EARLY ARABIDOPSIS ALUMINUM INDUCED 1, Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.002G200100
115 / 2e-32
AT4G12470 69 / 8e-16
azelaic acid induced 1 (.1)
Potri.018G025900
94 / 6e-24
AT2G45180 127 / 8e-39
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G158400
91 / 1e-22
AT1G62510 132 / 1e-40
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G121900
91 / 2e-22
AT1G62510 99 / 5e-27
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.003G111400
89 / 8e-22
AT1G62510 93 / 7e-25
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.014G059800
89 / 8e-22
AT4G00165 127 / 1e-38
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
Potri.001G121800
86 / 1e-20
AT1G12100 143 / 5e-45
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.006G256100
83 / 8e-20
AT2G45180 116 / 2e-34
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032259
102 / 3e-27
AT4G12520 133 / 2e-41
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10032253
99 / 2e-25
ND 139 / 2e-43
Lus10024616
99 / 2e-25
AT4G12520 139 / 2e-43
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10024623
99 / 3e-25
AT4G12490 152 / 2e-47
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10032254
98 / 3e-25
AT4G12520 139 / 3e-43
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10024615
99 / 5e-25
ND 139 / 6e-43
Lus10004348
97 / 6e-25
AT4G12520 154 / 4e-49
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10028929
97 / 1e-24
AT4G12520 147 / 9e-47
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10024626
96 / 3e-24
AT4G12520 157 / 7e-50
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10032261
96 / 4e-24
AT4G12470 157 / 9e-50
azelaic acid induced 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0482
Prolamin
PF14547
Hydrophob_seed
Hydrophobic seed protein
Representative CDS sequence
>Potri.001G122100.2 pacid=42793443 polypeptide=Potri.001G122100.2.p locus=Potri.001G122100 ID=Potri.001G122100.2.v4.1 annot-version=v4.1
ATGGCTTCAGGAGCACTAGCCTCCAATGCCCTTCTCCTCTCCCTCTACATCTTATTTTTGGCTTTGCTGAGCTCCGCTGAGTATTGCCCACCATCACCAC
CCAAGGTCAAGTCACCGCCTCCACCACCACCCAAGGCCAAGTCACCACCTCCACCACCCAAGGTCAAGTCACCACCTCCACCACCACCCAAGGTCAAGTC
ACCTCCACCACCCAAGGTCAAGTCACCACCTCCACCACCCAAGGTCAAGTCACCACCTCCACCACCACCCAAGGTCAAGTCACCTCCACCACCCAAGGTC
AAGTCACCACCTCCACCACCCAAGGTCAAGTCACCTCCACCACCCAAGGTCAAGTCACCACCTCCACCCAAGGTCAAGTCACCACCCCCACCACCCAAGG
TCAAGTCACCACCCCCACCGCCACCAATGGTCATGTCACCACCACCACCAATTGTCAAGTCACCACCGCCACCAATGGTCATGTCACCGCCACCACCAAT
TGTCAAGTCACCGCCACCACCAATGGCCAAGTCACCACCACCACCAATTGACAATTCACCGCCACCACCACCAATGTTCAAGTCACCACCACCACCACCT
CCATCACTACCAAAAGGCACTTGCCCTAGAGATACCTTGAAGTTGCAAGCATGCGCAAATGTGCTGAATTTGTTGAAAATCTTTGTTGGTGAAAAAGAAA
AGGCCAAATGCTGCAGCCTCATTGATGGTCTTGTTGATCTTGATGCTGCTGTTTGCCTTTGCACTCGAATCAAAGTTGATCTCCTGGGCCTTATTAAGTT
AGACGTTCCTGTTGCCGTGGAGTTGTTGCTCAACGAATGCGATAGGAAGGTCGCGGAAGATTTCAAGTGTCCGCCCTCCTAA
AA sequence
>Potri.001G122100.2 pacid=42793443 polypeptide=Potri.001G122100.2.p locus=Potri.001G122100 ID=Potri.001G122100.2.v4.1 annot-version=v4.1
MASGALASNALLLSLYILFLALLSSAEYCPPSPPKVKSPPPPPPKAKSPPPPPKVKSPPPPPPKVKSPPPPKVKSPPPPPKVKSPPPPPPKVKSPPPPKV
KSPPPPPKVKSPPPPKVKSPPPPKVKSPPPPPKVKSPPPPPPMVMSPPPPIVKSPPPPMVMSPPPPIVKSPPPPMAKSPPPPIDNSPPPPPMFKSPPPPP
PSLPKGTCPRDTLKLQACANVLNLLKIFVGEKEKAKCCSLIDGLVDLDAAVCLCTRIKVDLLGLIKLDVPVAVELLLNECDRKVAEDFKCPPS
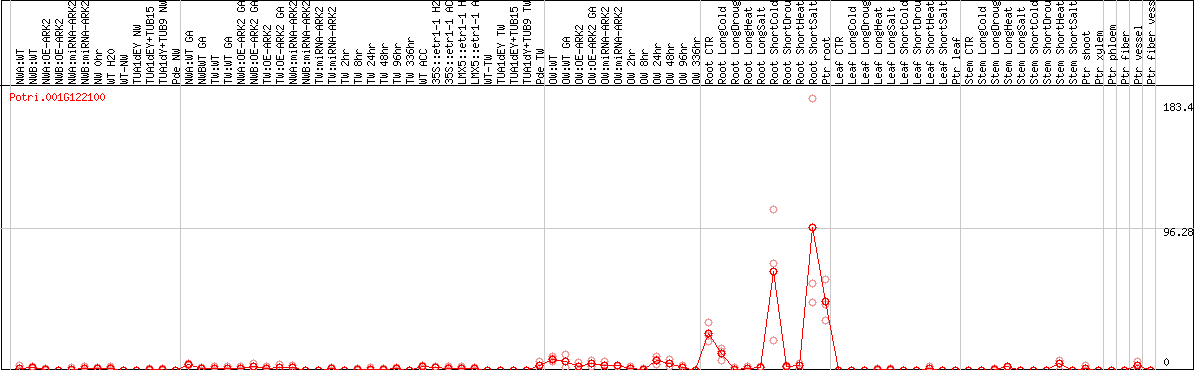
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G122100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.