External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G07840 209 / 8e-70
PIA1
phytochrome interacting ankyrin-repeat protein 1, Ankyrin repeat family protein (.1)
AT5G61230 207 / 5e-69
PIA2, ANK6
phytochrome interacting ankyrin-repeat protein 2, ankyrin repeat protein 6, Ankyrin repeat family protein (.1)
AT2G03430 64 / 8e-13
Ankyrin repeat family protein (.1)
AT2G43850 56 / 3e-09
Integrin-linked protein kinase family (.1.2)
AT3G04710 55 / 5e-09
TPR10
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
AT4G19150 53 / 9e-09
Ankyrin repeat family protein (.1.2)
AT5G02620 53 / 2e-08
ATANK1, ANK1
ankyrin-like1 (.1)
AT2G31800 52 / 3e-08
Integrin-linked protein kinase family (.1)
AT3G59830 52 / 6e-08
Integrin-linked protein kinase family (.1)
AT5G60070 50 / 2e-07
ankyrin repeat family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033816
235 / 4e-80
AT5G07840 232 / 1e-78
phytochrome interacting ankyrin-repeat protein 1, Ankyrin repeat family protein (.1)
Lus10018958
227 / 6e-77
AT5G07840 228 / 3e-77
phytochrome interacting ankyrin-repeat protein 1, Ankyrin repeat family protein (.1)
Lus10007432
61 / 3e-11
AT2G31800 69 / 3e-13
Integrin-linked protein kinase family (.1)
Lus10002621
59 / 2e-10
AT5G13530 99 / 6e-22
KEEP ON GOING, protein kinases;ubiquitin-protein ligases (.1.2)
Lus10020272
57 / 1e-09
AT5G13530 97 / 9e-21
KEEP ON GOING, protein kinases;ubiquitin-protein ligases (.1.2)
Lus10027107
54 / 2e-08
AT2G31800 709 / 0.0
Integrin-linked protein kinase family (.1)
Lus10040829
53 / 2e-08
AT2G28840 546 / 0.0
XB3 ortholog 1 in Arabidopsis thaliana (.1.2)
Lus10016561
52 / 3e-08
AT2G28840 640 / 0.0
XB3 ortholog 1 in Arabidopsis thaliana (.1.2)
Lus10008350
52 / 4e-08
AT2G31800 753 / 0.0
Integrin-linked protein kinase family (.1)
Lus10001048
51 / 6e-08
AT4G19150 230 / 6e-76
Ankyrin repeat family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0465
Ank
PF12796
Ank_2
Ankyrin repeats (3 copies)
Representative CDS sequence
>Potri.001G129800.1 pacid=42787645 polypeptide=Potri.001G129800.1.p locus=Potri.001G129800 ID=Potri.001G129800.1.v4.1 annot-version=v4.1
ATGCCACAGGAAGATTGTGTTGCTGTTTCGTTGAGGCGTAACTTGTCGAGAAGACGGTCGTTTAGGTCAGTTGGGGGTGTTGACAGAGATGATAGAGGTT
GGACTTCGCTTCATATTGGTGCTAGAAAAGGTGATCTCAAACAGGTGAAACATTTACTCGATGAAGGAATGGATGTTAATGTGCCTGCATGGGGCCCAAA
ATCAAAAGGCCTGACCGCGCTCCACCTTGCTGCTCAGGGTGGTCACCTTGAGATTATGGATGAATTGCTTGCTCGTGGTGCTAATATTGATGCTAGAACC
CTGGGTGCCTGTGGTTGGACACCACTTCACCGTGCTGCAAAGGAAAGGAAGAAAGAAGCAGTCAAATTTCTAATAGAGAACGGTGCATTCTTGCCAGATG
ATATGAATGATAGCAGATTTAACCCGCCACTCCATTACTGCACCGGTCTTGAATGGGCATATGAGGAGATGAAGCGTCATCAGAGAGAGAATTTGTCAGC
AGGCGAAGCATCCTACAGCTCTGAAAGCTAA
AA sequence
>Potri.001G129800.1 pacid=42787645 polypeptide=Potri.001G129800.1.p locus=Potri.001G129800 ID=Potri.001G129800.1.v4.1 annot-version=v4.1
MPQEDCVAVSLRRNLSRRRSFRSVGGVDRDDRGWTSLHIGARKGDLKQVKHLLDEGMDVNVPAWGPKSKGLTALHLAAQGGHLEIMDELLARGANIDART
LGACGWTPLHRAAKERKKEAVKFLIENGAFLPDDMNDSRFNPPLHYCTGLEWAYEEMKRHQRENLSAGEASYSSES
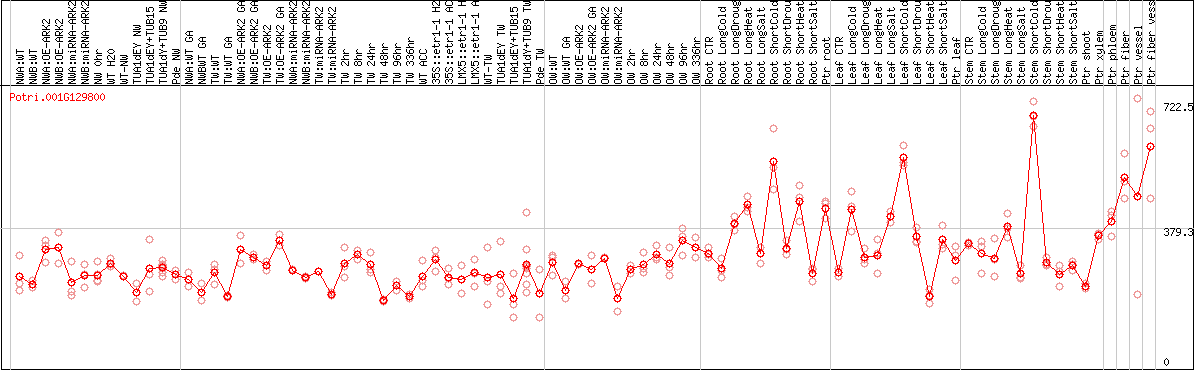
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G129800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.