External link
Symbol
Pt-ATMKK10.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G32320 232 / 4e-75
ATMKK10
MAP kinase kinase 10 (.1)
AT1G73500 183 / 4e-56
ATMKK9
MAP kinase kinase 9 (.1)
AT1G18350 177 / 5e-54
ATMKK7, BUD1
MAP KINASE KINASE7, BUSHY AND DWARF 1, MAP kinase kinase 7 (.1)
AT3G06230 156 / 5e-46
ATMKK8
MAP kinase kinase 8 (.1)
AT3G21220 155 / 6e-45
ATMAP2K_ALPHA, ATMKK5, ATMEK5
ARABIDOPSIS THALIANA MITOGEN-ACTIVATED PROTEIN KINASE KINASE 5, MAP kinase kinase 5 (.1)
AT1G51660 151 / 4e-43
ATMKK4, ATMEK4
ARABIDOPSIS THALIANA MITOGEN-ACTIVATED PROTEIN KINASE KINASE 4, mitogen-activated protein kinase kinase 4 (.1)
AT4G26070 131 / 4e-36
NMAPKK, MKK1, ATMEK1, MEK1
MITOGEN ACTIVATED PROTEIN KINASE KINASE 1, MAP kinase/ ERK kinase 1 (.1.2.3)
AT4G29810 131 / 6e-36
MK1, ATMKK2
MAP KINASE KINASE 1, MAP kinase kinase 2 (.1.2.3)
AT5G56580 126 / 7e-34
ANQ1, ATMKK6
ARABIDOPSIS THALIANA MAP KINASE KINASE 6, ARABIDOPSIS NQK1, MAP kinase kinase 6 (.1)
AT1G69220 106 / 2e-25
SIK1
Protein kinase superfamily protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G030700
191 / 7e-59
AT1G73500 392 / 9e-138
MAP kinase kinase 9 (.1)
Potri.012G043200
190 / 9e-59
AT1G73500 381 / 2e-133
MAP kinase kinase 9 (.1)
Potri.008G009800
156 / 3e-45
AT3G21220 474 / 9e-169
ARABIDOPSIS THALIANA MITOGEN-ACTIVATED PROTEIN KINASE KINASE 5, MAP kinase kinase 5 (.1)
Potri.010G249300
155 / 5e-45
AT3G21220 474 / 1e-168
ARABIDOPSIS THALIANA MITOGEN-ACTIVATED PROTEIN KINASE KINASE 5, MAP kinase kinase 5 (.1)
Potri.008G183700
155 / 6e-45
AT1G18350 289 / 5e-97
MAP KINASE KINASE7, BUSHY AND DWARF 1, MAP kinase kinase 7 (.1)
Potri.010G049500
152 / 5e-44
AT1G18350 288 / 1e-96
MAP KINASE KINASE7, BUSHY AND DWARF 1, MAP kinase kinase 7 (.1)
Potri.006G146500
137 / 6e-38
AT4G29810 495 / 7e-177
MAP KINASE KINASE 1, MAP kinase kinase 2 (.1.2.3)
Potri.018G068500
135 / 3e-37
AT5G56580 588 / 0.0
ARABIDOPSIS THALIANA MAP KINASE KINASE 6, ARABIDOPSIS NQK1, MAP kinase kinase 6 (.1)
Potri.018G050800
135 / 3e-37
AT4G29810 540 / 0.0
MAP KINASE KINASE 1, MAP kinase kinase 2 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035764
145 / 4e-41
AT1G51660 500 / 7e-179
ARABIDOPSIS THALIANA MITOGEN-ACTIVATED PROTEIN KINASE KINASE 4, mitogen-activated protein kinase kinase 4 (.1)
Lus10012938
137 / 5e-38
AT5G56580 621 / 0.0
ARABIDOPSIS THALIANA MAP KINASE KINASE 6, ARABIDOPSIS NQK1, MAP kinase kinase 6 (.1)
Lus10034986
137 / 8e-38
AT5G56580 605 / 0.0
ARABIDOPSIS THALIANA MAP KINASE KINASE 6, ARABIDOPSIS NQK1, MAP kinase kinase 6 (.1)
Lus10001081
125 / 1e-34
AT1G73500 296 / 2e-101
MAP kinase kinase 9 (.1)
Lus10027623
122 / 6e-32
AT4G29810 520 / 0.0
MAP KINASE KINASE 1, MAP kinase kinase 2 (.1.2.3)
Lus10011945
110 / 3e-27
AT4G29810 407 / 4e-140
MAP KINASE KINASE 1, MAP kinase kinase 2 (.1.2.3)
Lus10036915
107 / 6e-26
AT1G69220 917 / 0.0
Protein kinase superfamily protein (.1.2)
Lus10040127
100 / 2e-25
AT1G73500 164 / 1e-50
MAP kinase kinase 9 (.1)
Lus10037070
105 / 3e-25
AT1G69220 909 / 0.0
Protein kinase superfamily protein (.1.2)
Lus10022265
100 / 1e-23
AT5G40440 861 / 0.0
mitogen-activated protein kinase kinase 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF07714
PK_Tyr_Ser-Thr
Protein tyrosine and serine/threonine kinase
Representative CDS sequence
>Potri.001G138800.2 pacid=42792281 polypeptide=Potri.001G138800.2.p locus=Potri.001G138800 ID=Potri.001G138800.2.v4.1 annot-version=v4.1
ATGACTTTGGTGAGAGAGAGAAGGCACCAACAACCACTAAGGCTATCCCTACCACCACCCATACCTGCAGCTGACTTCCGGCATCAGATCCATTCGCCAT
CATTATCATTGACCATTAGTCCAGATTCTCCAAGCATAGAGAAGCTCTCAGACCTAGAAAAACTGGCGGTTCTTGGCCATGGCAATAGTGGCACGGTCTA
TAAAGTACGGCACAAACGCAGCTCCTCCATATTTGCATTGAAAACCCTCCGTTTCGACCGAAACTCCACCATAATTCGCCAACAAGCAGGAAGGGAGGCA
GAGATCCTTAGACGCGTGGATTCACCTTACGTTGTCCAGTGCCATGCGGTTTTTGATAGTGAGGATGACTTGTACTTTGCAATGGAGCACATGGAAAGAG
GATCACTGCATGATGTTCTGCTTGTACACAGAATATTGCCTGAGGATGTCATATCTGGTGTGGCACGGTGCATTCTGAATGGGTTACAGTATCTCCATGA
GAAGCAGATAGTACATGGGGACATAAAACCCTCAAATCTACTGATAAATGCTGAAGGAGTGGTTAAGATTGCAGATTTTGGAGTGAGTAGGGTAGTGGTG
GGAAAACATGATTCATATGAGACTTATATGGGAACATGTGCTTACATGAGTCCTGAGAGAATTGATCCTGAGAGATGGGATTGGAAATGGAGATCACGGA
TTCGCAGGAGATGTTTGGTCACTTGGGGTGGTAGTTTTGGAGTGCCTGGTGGGTCATTATCCATTGATTGGTTGTGGAGAGAAACCAGACTGGGCAGCAT
TGGTGTGTGCTATATGCTTTGGGGAGAGATTGCAAATGCCGAAGAGTGCATCCAGCAGGATTCAAAGCTTTGTTAG
AA sequence
>Potri.001G138800.2 pacid=42792281 polypeptide=Potri.001G138800.2.p locus=Potri.001G138800 ID=Potri.001G138800.2.v4.1 annot-version=v4.1
MTLVRERRHQQPLRLSLPPPIPAADFRHQIHSPSLSLTISPDSPSIEKLSDLEKLAVLGHGNSGTVYKVRHKRSSSIFALKTLRFDRNSTIIRQQAGREA
EILRRVDSPYVVQCHAVFDSEDDLYFAMEHMERGSLHDVLLVHRILPEDVISGVARCILNGLQYLHEKQIVHGDIKPSNLLINAEGVVKIADFGVSRVVV
GKHDSYETYMGTCAYMSPERIDPERWDWKWRSRIRRRCLVTWGGSFGVPGGSLSIDWLWRETRLGSIGVCYMLWGEIANAEECIQQDSKLC
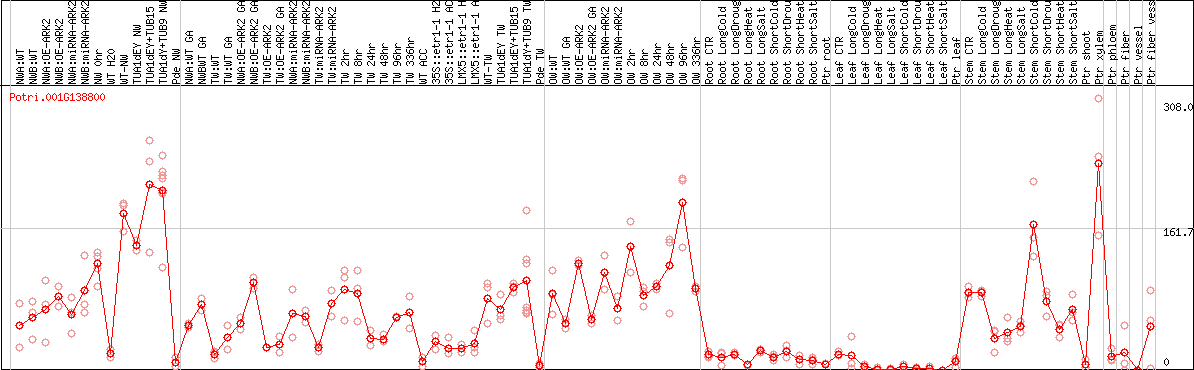
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G138800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.