External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G44610 298 / 1e-103
RAB6, AtRABH1b, AtRab6A
Ras-related small GTP-binding family protein (.1)
AT5G10260 279 / 7e-96
AtRABH1e
RAB GTPase homolog H1E (.1)
AT2G22290 265 / 1e-90
ATRAB6, ATRAB-H1D, AtRABH1d
ARABIDOPSIS RAB GTPASE HOMOLOG 6, ARABIDOPSIS RAB GTPASE HOMOLOG H1D, RAB GTPase homolog H1D (.1)
AT4G39890 236 / 8e-79
AtRABH1c
RAB GTPase homolog H1C (.1)
AT5G64990 234 / 3e-78
AtRABH1a
RAB GTPase homolog H1A (.1.2)
AT3G54840 116 / 2e-32
ARA6, AtRABF1, Ara-6, AtRab5C
Ras-related small GTP-binding family protein (.1.2)
AT5G45130 111 / 2e-30
ATRAB-F2A, RHA1, AtRab5A, AtRABF2a
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
AT4G19640 109 / 2e-29
ATRAB-F2B, ARA7, Ara-7, AtRABF2b, AtRab5B
ARABIDOPSIS RAB GTPASE HOMOLOG F2B, Ras-related small GTP-binding family protein (.1)
AT1G07410 107 / 1e-28
ATRAB-A2B, AtRABA2b
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
AT1G09630 107 / 2e-28
ATRAB-A2A, ATRAB11C, ATRABA2A
ARABIDOPSIS RAB GTPASE A2A, RAB GTPase 11C (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019501
299 / 7e-104
AT2G44610 407 / 5e-147
Ras-related small GTP-binding family protein (.1)
Lus10043349
299 / 7e-104
AT2G44610 407 / 5e-147
Ras-related small GTP-binding family protein (.1)
Lus10014336
283 / 1e-97
AT5G10260 402 / 3e-145
RAB GTPase homolog H1E (.1)
Lus10041702
282 / 4e-97
AT5G10260 404 / 7e-146
RAB GTPase homolog H1E (.1)
Lus10024046
282 / 4e-96
AT5G10260 405 / 3e-145
RAB GTPase homolog H1E (.1)
Lus10019502
154 / 8e-48
AT2G44610 218 / 5e-74
Ras-related small GTP-binding family protein (.1)
Lus10034903
116 / 6e-32
AT5G45130 258 / 2e-88
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Lus10028820
113 / 5e-31
AT3G54840 354 / 3e-126
Ras-related small GTP-binding family protein (.1.2)
Lus10040255
113 / 1e-30
AT1G07410 363 / 2e-129
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
Lus10004687
113 / 1e-30
AT1G07410 363 / 2e-129
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF08477
Roc
Ras of Complex, Roc, domain of DAPkinase
Representative CDS sequence
>Potri.001G147900.2 pacid=42793712 polypeptide=Potri.001G147900.2.p locus=Potri.001G147900 ID=Potri.001G147900.2.v4.1 annot-version=v4.1
ATGGCACCAGTCTCAGCTCTTGCAAAGTACAAGCTGGTCTTCTTGGAAGACCAATCAGTCAGTAAAACTAGCATCATCACTCGCTTCATTCAAGGAATTT
CGATTATTTTCGGGGTTCTTTTTTTTTTTGATATAGAATGTGGTGCTACCATTGGCATGGATTTTCTATCAAAAACCATGTATCTTGAAGACAGAACTAT
TCATTTGCAGTTGTGGGATACAGCTGGACAGGAAAGATTCAAAAGCCTCATTCCAAGTTACATTAGAGATTCCTCAGTTGCTGTCATTGTATTTGATGTT
GCAAGTCGGCAATCTTTCCTGAACACTTCAAAATGGATTGAAGAGGTTCGCACTGAGAGGGGCAGCGATGTCATCATTGTCCTTGTTGGGAACAAAACCA
ACCTTGTGGAAAAGAGCCACTCATTTTGGCTTGTAAATTTGTGGTCGGAACAAATACTGACAGATCTGAATTATTTGATGTTTTCCCTTTCTGCACAAGT
TTCTATAGAAGAAGGAGAAGCTAAAGCTCGTGACCTTAACGTCATGTTTATCGAAACTAGTGCAAAAGCTGGTTTTAATATCAAGGCACTTTCTTCAACA
AAGCAAGAGGACATGGTTGATGTGAACCTAAAGTCTTCCGGTGGAGCTTCATCTCAGACACAGCCTCAGGCAGGGGGATGTGATTGTTGA
AA sequence
>Potri.001G147900.2 pacid=42793712 polypeptide=Potri.001G147900.2.p locus=Potri.001G147900 ID=Potri.001G147900.2.v4.1 annot-version=v4.1
MAPVSALAKYKLVFLEDQSVSKTSIITRFIQGISIIFGVLFFFDIECGATIGMDFLSKTMYLEDRTIHLQLWDTAGQERFKSLIPSYIRDSSVAVIVFDV
ASRQSFLNTSKWIEEVRTERGSDVIIVLVGNKTNLVEKSHSFWLVNLWSEQILTDLNYLMFSLSAQVSIEEGEAKARDLNVMFIETSAKAGFNIKALSST
KQEDMVDVNLKSSGGASSQTQPQAGGCDC
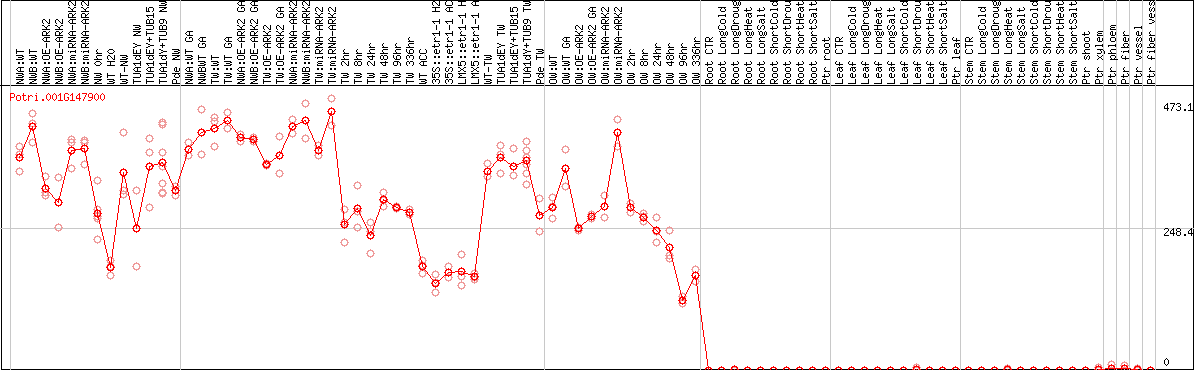
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G147900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.