Potri.001G148300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G148300.1 pacid=42790404 polypeptide=Potri.001G148300.1.p locus=Potri.001G148300 ID=Potri.001G148300.1.v4.1 annot-version=v4.1
ATGTGGGTGGTAATGTGTCCTTTCCTAATGCAATGGCAACTTTTTTGTTGCATTCTTTTAGGGTTTTTTTGTGCGTCCACCATTTCTTTTGCGCCAGGGC
AGTATGAGGGTGGGGGCATTTGGTCAGGAAATGGGTTACATAGTTCTGGATCAGTTTCAAGCAATCACTCTAGAAGTGGCACCTCTAGCTATGTTAAAAC
ATTAAAATTTTCTCTCCCATTAAACAGTTCCGTATCCTGCGAGGAACTAGAAGGAGTGGGATCGCTTAATACCACTTGCGTGGTGAACTCAAATTTGTAT
TTGAATTCTGATCTTTATATTTATGGAACAGGAAACTTAGAAATAATTCCACATGTTTCAATTGTATGCCCAATAGAAGGATGCATGGTGACGGTTAATA
TGACTGGAAATGTTAACATTGGTCAACATGCAGCAATCATTGCTGGTTCAGTTGTTTTTTCTGCTGCCAACTTGACCATGGATAGTCACTCTTCTATAAA
TACAACAGCCTTGGGTGGATCACCGCCGCCTCAAACTAGTGGTACACCAGTTGGGGATGATGGAGGTGGTGGGGGTCACGGTGGGCGAGGTGCTTCCTGT
TTGAAAAGGAATAAAACAAGTAATTGGGGTGGTGATGTTTATGCTTGGTCAACCTTGGCTGAGCCATGGAGCTATGGTAGTAAGGGTGGTGGCACATCGT
CTCAGAATAAATGTGGAGGGAATGGTGGGGGGCGTGTCAAGCTTCAAGTGAAGGAAATATTATACTTGAATGGATCTGTCGCTGCAGAAGGTGGGGATGG
GGGGCTGAATGGGGGAGGTGGGTCTGGTGGAAGCATATTTGTGCATGCTGTGAAGCTGAAGGGATATGGAACTATTAGTGCAGCAGGTGGAAGAGGATGG
GGTGGAGGTGGCGGTGGAAGAGTTTCTTTGGATTGTTATAGCATACAGGAAGATGTGAAAGTTACTGTGCATGGTGGTTTGAGTATTGGCTGTCCTGGGA
ATGCTGGAGCAGCTGGTACCTTCTTTAATGCAGATTTACTCAGTTTAAGAGTGAGCAATGACTATGTGATGACAGAAACAGAAACCCCTTTGCTTGATTT
TCCTACTATGACCCTGTGGTCAAATGTTTTTGTGGAGAATTATGCAAAAGTTTTGGTTCCACTGGTTTGGAGCAGGGTTCAGGTTAGAGGTCAGATTAGT
TTATATCGTGGAGGCAGCATTGTCTTTGGATTGTCTGAATTTCCAGTATCAGAATTTGAGCTTGTAGCAGAAGAACTTTTGATGAGCGATTCTATCATAA
AGGTTTTTGGTGCATTTAGAGTTGCTATCAAAATGCTACTCATGTGGAATTCCAAAATTGAAATAGATGGAGGTGGAAATACTGTTGTTACTGCTTCTGT
CCTTGAAGTCAGGAATCTGATTGTTCTAAGGGCAGGCTCTGTCCTCGGCTCAAATGCAAATTTGGGGTTATATGGTCAGGGACTACTAAAGTTGACAGGT
CATGGTGATACAATTCGAGGCCAGCGACTTTCCCTGTCTCTTTTCTATAACATAACTGTTGGACCTGGTTCTTTACTTCAAGCTCCGTTGGATGATGATG
CTAGCAGAAGTGTGGTTACAAAATCCCTTTGTGAGAGTCATACATGCCCAATAGATTTGATAACTCCCCCAGATGATTGCCATGTTAATTACACGCTGTC
GTTTTCACTTCAAATTTGTCGTGTTGAAGGCCTCCTTGTAAATGGCATTATAAAGGGGAGTATTATCCACATCCACAGGGCAAGAACTATCATTATTGAT
ACTGATGGCTTGATTACTGCATCGGAGTTAGGTTGCAATGATGGTATTGGAAAAGGGAACTATTCCAAAGGAGCTGGTAGTGGTGCTGGGCATGGGGGAA
GAGGGGGCTCAGGTTGTTTTAACGGGATCGTGAGTAATGGCGGTAATAAATACGGTAATGCTGATCTTCCTTGTGAATTGGGAAGCGGAACCCAAGGTCC
CAATCAGTCCTATGGAAATGTTATTGGAGGGGGAATGATTGTGATGGGCTCCATCCAGTGGCCACTGTTAAGGCTCAATCTTTATGGATCTTTGATGGTA
GATGGTCAAAGCTTTGATAAAGCGTCAGTAAATAGTAATGCTTCTTTGATTGGTGGACTTGGTGGAGCTTCTGGAGGGACGGTTCTTCTTTTCCTGCAGG
AGCTTATGCTTGCAGAAAAATCATCTCTATCTGTTCGCGGGGGCAATGGCAGCCCTCTTGGTGGCGGGGGAGGTGGGGGTGGAAGAGTTCATTTTCATTG
GTATAAGATAGACACCGGGGATGAATATGTTCCAGTTGCAAGTATAAGTGGTTCAATTAACAACAGTGGAGGAGCTGGAGAAAATGGTGGTCTTTTTGGG
GAGGAAGGCACCGTCACTGGGAAAAAGTGCCCAAAGGGCCTCTATGGTACATTTTGCAAGGAATGCCCTCTTGGTACTTTCAAAGATGTTGATGGCTCTG
ATGAAAGTCTTTGCATTCCTTGCTCTCTTGATCTTCTTCCTAATCGTGCAAATTTCATACATGTACGAGGGGGGGTTAGTCAGCCATCTTGTCCCTATAA
ATGCATATCTGACAAGTACAGGATGCCAAATTGCTACACTCCTCTTGAGGAGCTGGTGTATACATTTGGAGGCCCTTGGCCTTTTGCCCTCATCCTGTCT
GTCCTTTTGGTGCTTCTAGCTCTATTGTTGAGTACAGCGAGAATCAAATTAGTAGGATCAGGTAAATGTTATGATGCAAGTTCAGTTGAGCACCAGAGTC
ATCACCACTTTCCACATCTACTTTCCCTATCAGAGGTGCGAGGAACTAGAGCTGAAGAAAGTCAGAGCCATGTCTATAGAATGTACTTTATGGGCCCCAA
TACTTTTAGGGAACCCTGGCATCTTCCATACTTTCTCCCCAATGCAATTATTGAGATTGTGTATGAAGATGCTTTTAACAGGTTTATTGATGATATAAAT
TCTGTTGCCGCATATGATTGGTGGGAAGGGTCGGTCCACAGCATACTTTCCGTGCTTGCCTATCCTTGCGCATGGTCCTGGAAACAGTGGCGGCAGAGAA
ATAAAATTCATCGACTTCAGGAATATGTTAAGTCTGAATATGACCACTTGTGTCTTCGCTCTTGTCGATCTCGAGCCCTGTACAAGGGGATGAAGGTTGG
AGCGACACCAGACTTAATGGTTGCTTACATTGACTTTTTCCTTGGTGGTGACGAGAAGCGTTTAGATATAGTGTCTATAATACAAAAGAGATTTCCCATG
TGCATAATATTTGGAGGTGATGGGAGCTATATGTCACCTTATAATCTACATAGTGACACATTATTGACCAATCTTCTTGGTCAGCATGTCCCAGCAACCG
TCTGGAATCATTTGGTGGCTGGTTTAAATGCCCAATTGAGGATTGTGAGGCATGGATCTATCCGCTCTGCTCTACTTCCTGTCATTGATTGGATATGCAG
TCATGGAAACCCCCAGCTTGAGTTCCATGGAGTTAAGATGGAGCTGGGATGGTTCCAAGCAACTGCTTCTGGTTACTATCAGTTAGGTGTCTTAGTAATG
GTTGGTGATTATTCTCTCCATTCTATTCACCAGTCTGACTGGGTAGATAAAGGCAATGGTGAACCAACAAGGAACAGTGCCTCATGTGCAAGCAGAAGTC
TTAAGCAGCTGCAGCAAGAGCGGCCATACTTGAGTCAATCTTTATCTCGCAAAAGGATGACTGGGGGAATTAATGGAGGTCTTCTGAATGAAGCTACCTT
GAAATCTCTGGATTTTAAAAGGGACTTTCTCTCTCCGCTCTCTCTCTTATTGCATAACACTAGGCCTGTTGGCCGTCAGGATGCTCTTCAGCTTTTCATT
ACTATCATGCTTTTGGCAGATCTTTCTGTCACACTTCTCACATTGCTTCAGTTCTATTGGATCTCTCTTGGTGCTTTCCTTGCTGTTCTGCTTGTTCTTC
CTCTATCCTTGCTTTCTCCATTCCCTGCAGGCTTGAATGCCTTATTCAGCAGAGAACCTAGAAGAGCCTCACATGCTCGTGTATATGCATTATGGAATGC
AACCTCCCTGTCTAACATTGCTGTAGCTTTCACTTGTGGTATTTTTCATTATGGATTTTCTTCTCTCCGACCACCTGATGAGGAAAACACATGGAACATT
AGAAGGGAGGACAATAAATGGTGGCTGTTGTCAACTATCCTTCTGCTGTTCAAGTCAGTACAAGCTCGTTTAGTAGATTGGCACATTGCTAATCTAGAAA
TCCAAGACATTTCTTTGTTCTGCCCAGATCCAGATGCCTTTTGGGCTCACGAATCTAGTTCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G148300.1 pacid=42790404 polypeptide=Potri.001G148300.1.p locus=Potri.001G148300 ID=Potri.001G148300.1.v4.1 annot-version=v4.1
MWVVMCPFLMQWQLFCCILLGFFCASTISFAPGQYEGGGIWSGNGLHSSGSVSSNHSRSGTSSYVKTLKFSLPLNSSVSCEELEGVGSLNTTCVVNSNLY
LNSDLYIYGTGNLEIIPHVSIVCPIEGCMVTVNMTGNVNIGQHAAIIAGSVVFSAANLTMDSHSSINTTALGGSPPPQTSGTPVGDDGGGGGHGGRGASC
LKRNKTSNWGGDVYAWSTLAEPWSYGSKGGGTSSQNKCGGNGGGRVKLQVKEILYLNGSVAAEGGDGGLNGGGGSGGSIFVHAVKLKGYGTISAAGGRGW
GGGGGGRVSLDCYSIQEDVKVTVHGGLSIGCPGNAGAAGTFFNADLLSLRVSNDYVMTETETPLLDFPTMTLWSNVFVENYAKVLVPLVWSRVQVRGQIS
LYRGGSIVFGLSEFPVSEFELVAEELLMSDSIIKVFGAFRVAIKMLLMWNSKIEIDGGGNTVVTASVLEVRNLIVLRAGSVLGSNANLGLYGQGLLKLTG
HGDTIRGQRLSLSLFYNITVGPGSLLQAPLDDDASRSVVTKSLCESHTCPIDLITPPDDCHVNYTLSFSLQICRVEGLLVNGIIKGSIIHIHRARTIIID
TDGLITASELGCNDGIGKGNYSKGAGSGAGHGGRGGSGCFNGIVSNGGNKYGNADLPCELGSGTQGPNQSYGNVIGGGMIVMGSIQWPLLRLNLYGSLMV
DGQSFDKASVNSNASLIGGLGGASGGTVLLFLQELMLAEKSSLSVRGGNGSPLGGGGGGGGRVHFHWYKIDTGDEYVPVASISGSINNSGGAGENGGLFG
EEGTVTGKKCPKGLYGTFCKECPLGTFKDVDGSDESLCIPCSLDLLPNRANFIHVRGGVSQPSCPYKCISDKYRMPNCYTPLEELVYTFGGPWPFALILS
VLLVLLALLLSTARIKLVGSGKCYDASSVEHQSHHHFPHLLSLSEVRGTRAEESQSHVYRMYFMGPNTFREPWHLPYFLPNAIIEIVYEDAFNRFIDDIN
SVAAYDWWEGSVHSILSVLAYPCAWSWKQWRQRNKIHRLQEYVKSEYDHLCLRSCRSRALYKGMKVGATPDLMVAYIDFFLGGDEKRLDIVSIIQKRFPM
CIIFGGDGSYMSPYNLHSDTLLTNLLGQHVPATVWNHLVAGLNAQLRIVRHGSIRSALLPVIDWICSHGNPQLEFHGVKMELGWFQATASGYYQLGVLVM
VGDYSLHSIHQSDWVDKGNGEPTRNSASCASRSLKQLQQERPYLSQSLSRKRMTGGINGGLLNEATLKSLDFKRDFLSPLSLLLHNTRPVGRQDALQLFI
TIMLLADLSVTLLTLLQFYWISLGAFLAVLLVLPLSLLSPFPAGLNALFSREPRRASHARVYALWNATSLSNIAVAFTCGIFHYGFSSLRPPDEENTWNI
RREDNKWWLLSTILLLFKSVQARLVDWHIANLEIQDISLFCPDPDAFWAHESSS
|
DESeq2's median of ratios [POPLAR]
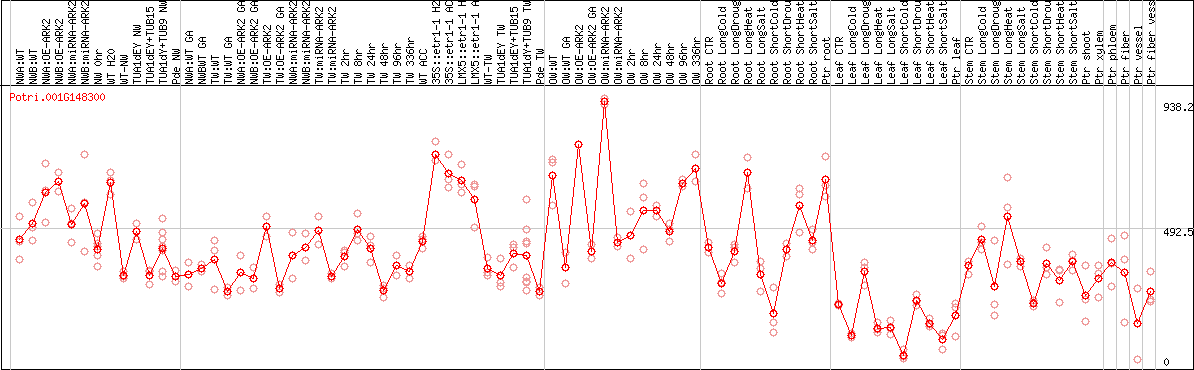
Coexpressed genes
Potri.001G148300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
