Potri.001G150001 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G150001.7 pacid=42791286 polypeptide=Potri.001G150001.7.p locus=Potri.001G150001 ID=Potri.001G150001.7.v4.1 annot-version=v4.1
ATGGAGAAAAGGGAGCCTGCTATACAGAATGTTGGTGCTCCAGTAACTCCAGAAAAGCCCCCGCCTCTGTCAGCAAGGCCACGACCGGTCTATGTTAAAA
GGTTCTCGGAACGTGGTTCACAAAGTTTTCTGGGAAGAATCGCAGCAGAAGAGGCTCGAGACCCCATTTGTGCAATTATTACTGCCGCGGCTTCCACAAG
CAAGAATGAACATCCCAACAATTGCAAGTATTTGGAAACTGGTGCTGTTTTTCAGATAGATTTGAAAGAAAGCATCTATGACTGGAACGGTAATTCTACC
GCTTCTGTTCATGGTACTCAAATCTTCGGAGACTCAGGAGGTCTGTCCGAAGAGGGTTTTCAATCTGAAAACCATAATCTTTTGTCTATTCCAGACCTGG
AGGAGAGTGATAGATACAGAGAAAAATGTTGTGTGCTTCCTTATGGTAAAGAACAAGATACAGAACATGGTACAATCTCTCGTATGAATTATGAGATGCT
TGATCATATTGGGTGCTCAAATCCAGTTGAGCAAGTTGCGATTTCGACTCCATTGCCAAAGAAAAGTGGCAGCAAAAGACCAAGAAATGGCATGGATCAG
AACAAAAGGCCAGTGAAGAAACCCAAGAGGAAGAAACATAGAACGAAAGTAGCTGTTGAAGGTAAACCAAAACGAACTCCTAAAAAGATGTCAAAGCTTG
CAACTCCGAAATGTATGCAAGAGACAAAGGAGAAGAGGAAGTGTGTTGAGAAAGCTCCTCCTCCTCTCAGAAATAGCCAAGAGAGTTCAGTGAGAGAGCA
TCATAAAGAAGTTGCTGAAATTCCAAAGATGGAAAATCTGCCAGTTGATGACAACAAGATGGCTATGGTTAGCAATGTCAACCATGATTTGAAAGATGAC
AATAAAGATTCGACTATGCAGCCTGGCATGGATAACGATAGTGGATCAATGGTTTTGTTGAGTGATGATAAATTGGGGTTCAAGTCAAATGAGAATGAAC
TAAGATGGAACGTTCGTGAAATGAGGTCTGCAGGAAAATTTTGGCCGAAACCTGGGTGTCAATATAATTCATTACAGGCTTATAGAAGGGTATTTAGAGT
AGACTCCTGTGTCGGCAATAGCAGAGAGGTTGGGCCTAATGTTCCTAAAAGATGCATGAAAAAAATGATCAAGAGGGGGCGTAAATTGAACCTGTTCAGC
TCTTTGTGGTCGTCTCACTTTAGTTGCCATGAAAGAAAGAAGAGATCTAAGGTGTTTACTCCAAGAGTCGATCCCACTTCTCTCAATGCAAGATTCGCAG
AGAAAGGGAAGGATTTAGAAACAGCTCCTCGGGATCATATACCCATTCCTGGAGAACAAGAGTTTCAAATAACCGACAAACTAAATGGAGAAAACCGCGA
AACTCTAGCAGAGGGATGCGTTGAGGAAGGGCTTTCTCTTAAACAAGCTCTAACCAATGAAGTTAATGGAGTTCAAATAGTCTGTGAGGCTAAAGCACCA
AGCTACTCTGAGATGAGACCCTCTGCTGCAAAATCAAAGTTTGAAGATGGCAAAAGCAAACAACAGTCGCGGAAAAAGAAGATCCGAATCACCGACAAAT
CAATAAAACCTGGACCGAAACTTTTCGATGACGAAAGTGGGATTCTTCTTAAGACAGTTAAGCATGCTGTAATGGATAAACTTGGTGATCTACTACGGAA
TTCAAGCATCAATGACAATAAGAGTGAAAAAAGGGGTCCTGGCAGACCAAAAAGTAAACCGGAACATGTAGTTGACGGACAGAATGCTTTAATTCCCTAT
TGGGGTCGTGGAAGAGCAAGACATGAAGTTGATCTAGATCCAGAGTCATTGAGAAGGTGGAACCAACTAATGAAAATAGATAGTGGAGAGGGTGAAGAAG
AGGATGGAAAAAGGAAAGAATGGTGGCAAAGAGAAAAAATAACTTTCAATGGACGAATAGATGCGTTTACTTCTCGGTCGCATCAGTTTCTAGGTGATAG
GAGATTCAGACAGTGGAAAGGTTCTGTTGTGGACTCAGTGGTAGGAGTTTTCCTAACCCAAAATGTGTCAGACCACCTATCAAGTGCTGCATACATGTCA
CTTGTTGAAAATTTTCCAGTTCAGTCAACGAGTAATCAACAAGCGTCAATGACCATAGATGGCCAAAAACCAATTGAGAGCAGCATGACATCCATTGGAG
CAACTTATGATGCAGATGGAAACCGATATTTTGTCACAGAAGTACCTGAACCAGATATGAATCATGGGTCCAAAAATTTACCAGCCGAAAAGGCTACATG
CTTGCTTCAAGAAGAGGTACAAAGACGGAATGGTATTGTGAGATCAGAACCTGAACCGGATATGAATCAAGGACACAAAGATTTGCCGGCCGAAAAGGAT
ACGTGCTTGCTTCAAGAAGACGAGCAAGGACAGAATAGTGTTGTGAGATCAGAATCTGAACCAGATATGAATCATGGACCCAAAGATTTACCCGCCGAAA
GGGCTATATGCTTGCTCCAGGAAGAGGTGCATAGACAGAATGGTATTGTGAGATCAGAACCTGAACCAGATATGAATCTTGTGTCTCAATATTTACTAGT
TGAAAATCCTATGTGCTTGCCTCGTGAAGTGGTGCGAAGACAGCCTCAAAATTTACCAACCGAACAGCCTATATGTTTGCCTCATGAAGTGGTGAAAAGA
CAGAGCGGGAATGAAACTGAAAAGAAAAACAGAAGCCCGAATAGGAAAACAGCTTTTAAGGAGACTAAAAAGACAATTCCGAGAGGGAAGAATAAAATTG
TTAAAGATGTGGTACTTAGAAGGAGGAACTGGGAGAGTTTAGGAAAGATCTATTCTAGACCAAGAAGCAAGGATCAGATGGATTCGGTGGATTGGGAGGC
AGTAAGACAAGCAGAAACCAGCAAAGTTGCCAGCATCATCGAAGGACGTGGTCAGCAAACTATTATTGCGGGAAGAATCAAGCAATTTCTGGATCGAGTT
GTTGACATGCATAAAAGCATAGATCTCGAATGGTTACGATATGCTCCACCTGATGATGTGAAGGATTACCTGTTGGAGTTCATGGGATTGGGCTTGAAAA
GCGTGGAATGTGTACGACTTTTGTCTCTTCAACAGGTTGCCTTTCCGGTTGATGTAAATGTTGCTCGGATAGCGGTTCGTTTAGGATGGGTTCCCCTGAA
AGCACTCCCTGGAAGCCTTCAGTTCCACCTTATAGAAGAGTTTCCGATATTGGACACCATTCAGAAATACCTCTGGCCCCGACTATGCGAGCTTGACCAT
AGAACACTCTATGAACTGCACTATCACATGATAACCTTTGGAAAGGTATTCTGCACAAAGGAAAACCCAAGGTGCCATGCTTGTCCAATGAAAGCAGAAT
GCAGACATTATGCAAGTGCACAATCAAGTGCTGCCAATCTTTCCCTACCAGGACCGTCAAAGAAAAGTGTGGACCAATCAATTGTTACCAGCAAGCCTAG
TAGAAATTCTAGTTCACTTGGAAACAATTGTGATGTCAAAAATCCAATGTCAGTTACCCTTTTAGATGCCAACAGAACTACAGAATCTCGTCGAGATGTT
CACACTTGCAAACCCATAGTTGAAGAGCCAAAATCACCACAACAAGAAGAGGACATTGAAGATTTTTCATGGGATGGATGTATTGAAGATGAAGAGATCC
CCACTATCAAACTCAATCGTGAAGAGTTCAACCCAGATGGGAAAAAGTCTCCATACAGCAATTCTATGTCGCAAGCTTTAGTTTCTGTGGGGACTACTCC
ATTGTCAGCACCTAAAATGAAGCGTGTGACCTCTTTGCGGACTGAACATCAAGTGTATGAACTTCCAGATAATCATGAGATTTTGGTTGGGCTAGACAAA
CGAGAAAGGGATGATTCAGTACCTTACCTTCTCGCTATATGGCAACCAGGTGAAACTCCGAATTCAAGCCAGCAACCAGAGAAATTATGCAGCTCGCAGG
GATCACAACTCTGTGATCAGAAAACATGTTTTGCTTGTGAAGGTATTCGAGAACAACAAGCTGGTATAGTTCGTGGAACAATTTTGATACCATGCAGGAC
AGCTCTGAAGGGAAGCTTTCCACTCAATGGCACATATTTTCAAGTTAATGAGGTATTTGCTGATCACAAATCTAGTTATGACCCCATTATTGTTCCCAGA
GAATTGCTATGGAACCTCGTAAAGAGGACTTTATATGTTGGGTCATCAACAAAATCAATATTTAGAGATTTGTCATTAAAGGAGATTCACCAAAACTTTT
GGACAGGTTTTACTTGTGTGAAAGCATTCGAGCGAGGAACAGGTGCTCCAAAACCTCTTGCAAGGAGATTCCACTGTTCAGCAAGCAAGATGGAGAAGGA
AGGAAGGAAAAAATCAGAGAAAAAATCATTTCACAGCCAGATGACCATCTTATTCAAGCCTCCAAACTTCTATGAATGCCAGTTGGGTTTGTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G150001.7 pacid=42791286 polypeptide=Potri.001G150001.7.p locus=Potri.001G150001 ID=Potri.001G150001.7.v4.1 annot-version=v4.1
MEKREPAIQNVGAPVTPEKPPPLSARPRPVYVKRFSERGSQSFLGRIAAEEARDPICAIITAAASTSKNEHPNNCKYLETGAVFQIDLKESIYDWNGNST
ASVHGTQIFGDSGGLSEEGFQSENHNLLSIPDLEESDRYREKCCVLPYGKEQDTEHGTISRMNYEMLDHIGCSNPVEQVAISTPLPKKSGSKRPRNGMDQ
NKRPVKKPKRKKHRTKVAVEGKPKRTPKKMSKLATPKCMQETKEKRKCVEKAPPPLRNSQESSVREHHKEVAEIPKMENLPVDDNKMAMVSNVNHDLKDD
NKDSTMQPGMDNDSGSMVLLSDDKLGFKSNENELRWNVREMRSAGKFWPKPGCQYNSLQAYRRVFRVDSCVGNSREVGPNVPKRCMKKMIKRGRKLNLFS
SLWSSHFSCHERKKRSKVFTPRVDPTSLNARFAEKGKDLETAPRDHIPIPGEQEFQITDKLNGENRETLAEGCVEEGLSLKQALTNEVNGVQIVCEAKAP
SYSEMRPSAAKSKFEDGKSKQQSRKKKIRITDKSIKPGPKLFDDESGILLKTVKHAVMDKLGDLLRNSSINDNKSEKRGPGRPKSKPEHVVDGQNALIPY
WGRGRARHEVDLDPESLRRWNQLMKIDSGEGEEEDGKRKEWWQREKITFNGRIDAFTSRSHQFLGDRRFRQWKGSVVDSVVGVFLTQNVSDHLSSAAYMS
LVENFPVQSTSNQQASMTIDGQKPIESSMTSIGATYDADGNRYFVTEVPEPDMNHGSKNLPAEKATCLLQEEVQRRNGIVRSEPEPDMNQGHKDLPAEKD
TCLLQEDEQGQNSVVRSESEPDMNHGPKDLPAERAICLLQEEVHRQNGIVRSEPEPDMNLVSQYLLVENPMCLPREVVRRQPQNLPTEQPICLPHEVVKR
QSGNETEKKNRSPNRKTAFKETKKTIPRGKNKIVKDVVLRRRNWESLGKIYSRPRSKDQMDSVDWEAVRQAETSKVASIIEGRGQQTIIAGRIKQFLDRV
VDMHKSIDLEWLRYAPPDDVKDYLLEFMGLGLKSVECVRLLSLQQVAFPVDVNVARIAVRLGWVPLKALPGSLQFHLIEEFPILDTIQKYLWPRLCELDH
RTLYELHYHMITFGKVFCTKENPRCHACPMKAECRHYASAQSSAANLSLPGPSKKSVDQSIVTSKPSRNSSSLGNNCDVKNPMSVTLLDANRTTESRRDV
HTCKPIVEEPKSPQQEEDIEDFSWDGCIEDEEIPTIKLNREEFNPDGKKSPYSNSMSQALVSVGTTPLSAPKMKRVTSLRTEHQVYELPDNHEILVGLDK
RERDDSVPYLLAIWQPGETPNSSQQPEKLCSSQGSQLCDQKTCFACEGIREQQAGIVRGTILIPCRTALKGSFPLNGTYFQVNEVFADHKSSYDPIIVPR
ELLWNLVKRTLYVGSSTKSIFRDLSLKEIHQNFWTGFTCVKAFERGTGAPKPLARRFHCSASKMEKEGRKKSEKKSFHSQMTILFKPPNFYECQLGL
|
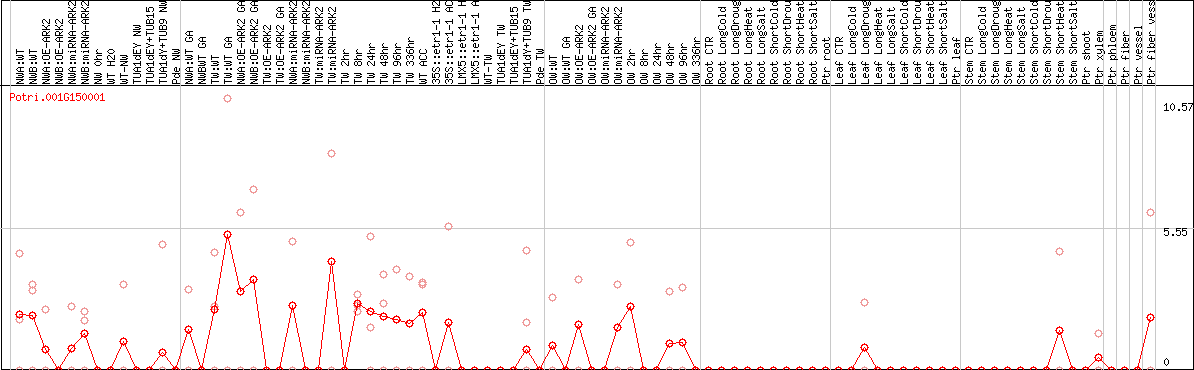
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G150001 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
