Potri.001G150900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G150900.1 pacid=42792640 polypeptide=Potri.001G150900.1.p locus=Potri.001G150900 ID=Potri.001G150900.1.v4.1 annot-version=v4.1
ATGAATGCAGAAGAGAGCAATCCGATGGTCTCGATAATCTCCTCTCTATCTGACAGCTTCAAGCAAGTCCCTTTGGCAGCTCTCCCAGCAATGCTTGATT
GCATTTTAGCATCAACTGGGCTATCTCCATCTGCCCTTTTTGCTTCACTTCTTGATTCATTCTCTAAATTTATCAAGGATGTTAGGGAGAAGGACTTGAA
GTTGGACTCTAGCATGTGTAACTACATCACATCCATGGTGGGTTCGCTTTGCCATCTTCTAAACAAATTTGGAAATAATACTGATGGTCTGCAGTCATTT
ATCTGGAAATGTTTTATTCCATTGATGAAGATGGTTCATGCTTTTGAGCGTGAAATGCTTAATGAGATTGCAGAGTCATTCTTTTGTGTCGTTAGCTCAA
CTCATTCCTGGGGTGTTTTGGAAGCAAACTTGGTACCATTTTTTCTAAGATCAGTTGGTCTTTCTATCGGCATGATTCAAAATGAAGAATCAGATGCCTT
TGAATGGGATCATTTCTCCATTTACCATGGTTTAAGTGACCTGGAAAATGATTTCGATTTGGATCAAGAGCCCATGCTGTCCCTGTCAGGGTCCTTTCCA
CTACCAATATCATGCCATATTCTAACCTTAATCCTGGATGCTGCTCTGCAAAGTTTCCAGGCAGTTTCAAGCACAAAATCAATGTTGGCAAATGGGTTCT
GTGATGTCGAAAAGCTATTTTCTAATTTGCTTTGGGATCTATGTAACATGAGTGAGCGATTGCTTTCACAAAGTTTGGAACATCGTTCTTGCACTATTGG
ATTTCTGCTTCCAATCATTTTCAAAGCATTGGGTTCTCAGCGCTCTTTGGAAATCACAGTCCATGGGAAGATGTTTATACTGTCCAGAAACGTTTTCTTT
AGGAAAATTTGGAAGCTGTGCAGATCATTGTTTTCTCTTGGACATTTAGAGAGAAGAGATGCCTATAATGTACTCTCTTTATACTTGTCCTTTTTCTCTT
TGACAGAAGGATTTGGAAATGTTGATGCAAGTGTAAAAGCAGAAGAATTTGATGTAAGAGCTGAGAGGGAGTTTTGGGATGAAATAAAAAGAGGCCTGGT
TGATGAAGAAGGCCTGGTAAGGAAGCAATCATTACACATATTGAAGACAGTACTACAGATAAGTGGTGGAAGCCAGTGCCACTCTGGGGTTTCAGAGAAG
AAATCACAAGAGAAACACCCAGTTCCACATGGTATGACAAAGAGGGAAATGTGGGCTGACAAGGAAGCTAAATCCTTAGGTGTATGGGAACCCTGCAATT
CAGCCGATTCTCCTTTAAACAGTCAGCAGCAGTGGGAAGCGTTTATACTCCTGTATGAAATGCTTCAAGAGTATGGTACGCACTTGGTAGAAGCTGCTTG
GCATCACCAGCTAAACTTGTTGCTTCAGTTTTCTGTTTCAAACAATAATTTTACAAGCTATATCTTCAGAGGATTTCATCAAAAGCAGACTGATATTTTG
AGAGAAGCATTTAGTTGGGTAACAATTTTATGGCAACTGGGCTTTCAACATGATAATCCTCAAGTTAGGTGCTTGATCATGGAGTCGTTTTTGGGCATTG
AGTGGATGAAGTATGGAAACACTGCCAAATCAGTGTCTGAAAGTTTTGTTCTTGGACCATTCATTGAAGGGTTGAATGACCCTGTGCACCACAAAGATTT
CGGTGTAAAAGGGTCTTACAATTCAAAGACTATTGAAGGTGCAGCCAGGTTCTTGCATCAATATACAAGTCACCTAAATACAAGGGAAGGGATTGCTTTT
TTGCATAGTTTAGCATCTGTTGCCAAACATCACTCTTTTGGTCGAGCTGGATTAATGGGTCTTGCAGAATGTATTGCTTCAGCTGCAAATGGAGTAGGAA
GGCATGATAGTGGAGCAAAATGGAGTGAAGATGCTTTTCCTGATGAGGTGCAAGTGGAATCTTCTCCTGAAAATTTTTCTGATTGTAGAACAGCCTTTTT
GGATGTTCTGAGATTTGTTATTGAGAGCAGCAAGCAACATTTCAACCCTTATTATCGTCTTCAAGTTTGTGAAAAAGTTCTGGAGGCTGCTACTTCGCTG
GTCAGTACACTTGATGTGCCTCTTGAGATTCTCTTGCACTTCATTGCTACACTACCACGGGCATTTACTGATTATGGGGGTTCTTTAAGATTAAAAACAC
AAGAGTGGCTCTTGGGGTCTGCTACGGAACATTGTAATGTAAATTGCTGCGGTGCTGAGATTCAACTTCTGAAAAATCTTCAGGACTTCCCAGAAAGGTT
TACAAGTTCTCAATACTTGGTTGATGGTTTTTTGAGTTTAGATGATGAAGACTTGGATGCATGGGAATCTGAATCAAAAAGATGGGCAAGAGCCCTTTTT
CTTATAATCAAGGGGGAGGATCAACTAGCACCGATATTGAGATTTATTCAAAATTGTGGCGTCAACATCTGCAAACAACAGAGTCATTTAGAATGGCTAC
CTGTGAAGTTTCTTGTCCTTGCTAGGAGCTTGGTTGCAGAGATTCAAATAATGCAGGAGAGATCAGCCCAATGTGGTATCAAAATTAAATGCAGATCAGA
GATTAGCTTGCTTGATACGGTGGATCAATTGTGTTATACAGAAGCTTCTATGATTAATGGAAGAATTCATGGCCTTTTCCTGTTCATATTGGAGGAGCTG
GTTTCTTTTGCCGACTTGTCTTCTTCCATATTTTGGTCTAGCATTACTAAGGAGACAACTCTTCCTGGTTCTGTGAGAGGAAAACTTGGAGGCCGTAGTC
AACGTCGGTTGTCAACTTCTACCACCACTGCTATATTGCAAGCTATAACATCAATACAAGCGGTTGCATCCATTTCATCATGGTGTGCACAGTTTAAAAG
TGATGTTAAGCTCAGTTCTGTGTGGAACTTTTTGTGGAAATTCTTCTGGAAGACGGTTTCCTCTCCAACTTGTGATTCTGAGGCTGGAGCAGAAATTTGT
CTGGCAGCATATGAAGCATTAGCTCCTGTTTTGAGAGCTCTAGTTTCAACATCCTCTTCACTATCTTTGGATCTCATTAGAGAAAATGATGAATTTTCGG
CACCTGTTGTAGAAGGCAAATGCTGCTTAGATTCTTTGGCTCTTTCTTTTCTTCAGAATATCAATAATCTCCTTGCAGTTGGAGTTCTGGCACGAACTCG
ACGGGCAGTTTTGCTAAACCAGAAGTGGATTTGCTTGGAGTCTCTGCTGTCTATTCCTTACTCTGCTCCTTGGAATGTGCTAAATTTAGAGGATGGCAGC
TTGTTCTTCTCGGATTCAGCAATCAGATGTATTTTCAGTGATCTTGTTGAAAGTCTTGATAATGCTGGGGAAGGTTCTGTTTTACCCATGCTGAGATCTG
TCAGATTGGCTTTGGGCCTGATTGCCTCAGGAAAGTTAGATTCACATGTTTCCTCATGCAATGGTGTGGATGCTCAGATGATGTGGCGTTTGGTTAATTC
ATCTTGGATATTGCATGTCAATTGCAACAAGCGGAGAGTGGCATCTATTGCTGCACTTTTGTCTTCTGTCTTGCATCGTTCCGTGTTTACAGATGAGGGC
ATGCATTTAATCAACAATAGGCCCGGGCCTCTGAAGTGGTTTGTTGAAAATGTTATTGAGGAAGGTACAAAAAGTCCTCGTACAATTCGCCTTGCAGCAT
TACATCTAACAGGACTGTGGCTCTCTCATCCAAAGACTATTAAGTATTACATGAAAGAACTGAAGTTGCTATCACTTTATGGTTCTGTTGCATTTGATGA
GGATTTTGAAGCTGAATTATGTGACAATCAAGATGCAAGCACTGAAGTGTCATTATTGGCTAAAAGCCCAGACCCTGAGCTGACTGAAGCATTCATCAAC
ACAGAATTGTATGCACGTGTGTCTGTAGCTGTTCTTTTTTACAAGCTTGCTGACTTGGCCAATTTGGTGGGATCGGCAAATGAGAATGAAGATTGTCACG
CTGCTCTTGAATCTGGAAAATTGTTTCTGCAGGAGCTTCTTGATTCTGCGGTGAATGACAAAGATCTTGCAAAGGAGTTGTATAAAAAGTATAGTGGGAT
CCATAGACGTAAAATACGAGCTTGGCAAATGATATGTGTTTTGTCACGATTTGTTACTGATGATATAGTTGCGCAAGTTACTCACAGCTTGCACATATCC
CTCTATAGAAATAATTTTCCCGCAGTTCGTCAATACTTGGAAACTTTTGCAATTAACATCTACCTGAAATTCCCATTGTTGGTCAGGGAACAATTAGTAC
CTATATTACGAGATTATAACATGAAGCCTCAGGCACTCTCTTCCTATGTGTTCATAGCAGCCAATGTCATCCTCCATGCTTCTAATGCAAATCAATCCAG
GCACTTCAATGAATTGCTTCCACCTATAATTCCATTATTAACCTCACACCACCACAGTTTACGTGGTTTTACGCAGTTGTTAGTCTACCAAGTTTTTTGT
AAATATTTTCCTATGTTGGACTATGGTGCCTCTGAAATGCCTTTAGAGAAAATGTGCTTTGAAGATTTGAAATCATACCTGGCAAAGAACCCTGATTGTA
GGCGCTTACGAGCATCTTTGGAAGGATATCTTGATGCCTATAATCCCATAGCCTCTGGTACTCCAGCTGGAATTTTCATTGATCGGGTTGAGGAACTTGG
GTTTGAATGTGTTCCAACATCCCTCATGGAGGAAGTGCTCAACTTTCTAAATGATGTCAGGGAAGACCTCAGATGTTCAATGGCAAAGGATGTTGTGACT
ATCAAGAATGAGAGCTTAAAAACTGATGAAGATGGTAACTGCAGGCGAACAGTAATTGATTCCCAGCTGCCTAAAGAAACATCATTTGATTTCCAGAAAA
AGCTTACTCTCTCTAAACATGAGAAACAAGACACTGATTCCAGCTCTGTTTTGGGGAACAATGAAGCATGCAAACAACTTTTAGAGATGGAAAAGGAAGA
TGAGCTTCTTGATCAGTCACTTCAATCCAGAAGATTGACCATGGAAAAAATAAGAGCAAGTCGACAACAATTTATCCTAGTTGCATCATTGCTTGATCGG
ATACCCAATCTTGCTGGATTGGCACGGACATGTGAGGTATTTAAGGTGTCAGGACTAGCCATTGCAGATGCAAGCATATTACGTGATAAACAATTCCAGC
TCATCAGCGTGACAGCAGAGAAGTGGGTTCCTATAATAGAAGTCCCAGTAAACAGCGTGAAGCATTTCTTAGAAAAAAAGAAACGAGATGGCTTCTCAAT
ATTGGGACTAGAACAGACAGCAAATAGTGTACCGCTCGATCACCATGCCTTTCCCAAAAAGACAGTACTGGTCCTTGGACGTGAAAAGGAGGGCATACCG
GTGGACATTATCCATATGCTCGATGCTTGCATTGAGATTCCGCAATTGGGAGTAGTTAGGTCGCTTAATGTTCATGTTAGTGGGGCAATTGCCCTCTGGG
AGTATACTCGACAGCAAAGATCTCAATAA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G150900.1 pacid=42792640 polypeptide=Potri.001G150900.1.p locus=Potri.001G150900 ID=Potri.001G150900.1.v4.1 annot-version=v4.1
MNAEESNPMVSIISSLSDSFKQVPLAALPAMLDCILASTGLSPSALFASLLDSFSKFIKDVREKDLKLDSSMCNYITSMVGSLCHLLNKFGNNTDGLQSF
IWKCFIPLMKMVHAFEREMLNEIAESFFCVVSSTHSWGVLEANLVPFFLRSVGLSIGMIQNEESDAFEWDHFSIYHGLSDLENDFDLDQEPMLSLSGSFP
LPISCHILTLILDAALQSFQAVSSTKSMLANGFCDVEKLFSNLLWDLCNMSERLLSQSLEHRSCTIGFLLPIIFKALGSQRSLEITVHGKMFILSRNVFF
RKIWKLCRSLFSLGHLERRDAYNVLSLYLSFFSLTEGFGNVDASVKAEEFDVRAEREFWDEIKRGLVDEEGLVRKQSLHILKTVLQISGGSQCHSGVSEK
KSQEKHPVPHGMTKREMWADKEAKSLGVWEPCNSADSPLNSQQQWEAFILLYEMLQEYGTHLVEAAWHHQLNLLLQFSVSNNNFTSYIFRGFHQKQTDIL
REAFSWVTILWQLGFQHDNPQVRCLIMESFLGIEWMKYGNTAKSVSESFVLGPFIEGLNDPVHHKDFGVKGSYNSKTIEGAARFLHQYTSHLNTREGIAF
LHSLASVAKHHSFGRAGLMGLAECIASAANGVGRHDSGAKWSEDAFPDEVQVESSPENFSDCRTAFLDVLRFVIESSKQHFNPYYRLQVCEKVLEAATSL
VSTLDVPLEILLHFIATLPRAFTDYGGSLRLKTQEWLLGSATEHCNVNCCGAEIQLLKNLQDFPERFTSSQYLVDGFLSLDDEDLDAWESESKRWARALF
LIIKGEDQLAPILRFIQNCGVNICKQQSHLEWLPVKFLVLARSLVAEIQIMQERSAQCGIKIKCRSEISLLDTVDQLCYTEASMINGRIHGLFLFILEEL
VSFADLSSSIFWSSITKETTLPGSVRGKLGGRSQRRLSTSTTTAILQAITSIQAVASISSWCAQFKSDVKLSSVWNFLWKFFWKTVSSPTCDSEAGAEIC
LAAYEALAPVLRALVSTSSSLSLDLIRENDEFSAPVVEGKCCLDSLALSFLQNINNLLAVGVLARTRRAVLLNQKWICLESLLSIPYSAPWNVLNLEDGS
LFFSDSAIRCIFSDLVESLDNAGEGSVLPMLRSVRLALGLIASGKLDSHVSSCNGVDAQMMWRLVNSSWILHVNCNKRRVASIAALLSSVLHRSVFTDEG
MHLINNRPGPLKWFVENVIEEGTKSPRTIRLAALHLTGLWLSHPKTIKYYMKELKLLSLYGSVAFDEDFEAELCDNQDASTEVSLLAKSPDPELTEAFIN
TELYARVSVAVLFYKLADLANLVGSANENEDCHAALESGKLFLQELLDSAVNDKDLAKELYKKYSGIHRRKIRAWQMICVLSRFVTDDIVAQVTHSLHIS
LYRNNFPAVRQYLETFAINIYLKFPLLVREQLVPILRDYNMKPQALSSYVFIAANVILHASNANQSRHFNELLPPIIPLLTSHHHSLRGFTQLLVYQVFC
KYFPMLDYGASEMPLEKMCFEDLKSYLAKNPDCRRLRASLEGYLDAYNPIASGTPAGIFIDRVEELGFECVPTSLMEEVLNFLNDVREDLRCSMAKDVVT
IKNESLKTDEDGNCRRTVIDSQLPKETSFDFQKKLTLSKHEKQDTDSSSVLGNNEACKQLLEMEKEDELLDQSLQSRRLTMEKIRASRQQFILVASLLDR
IPNLAGLARTCEVFKVSGLAIADASILRDKQFQLISVTAEKWVPIIEVPVNSVKHFLEKKKRDGFSILGLEQTANSVPLDHHAFPKKTVLVLGREKEGIP
VDIIHMLDACIEIPQLGVVRSLNVHVSGAIALWEYTRQQRSQ
|
DESeq2's median of ratios [POPLAR]
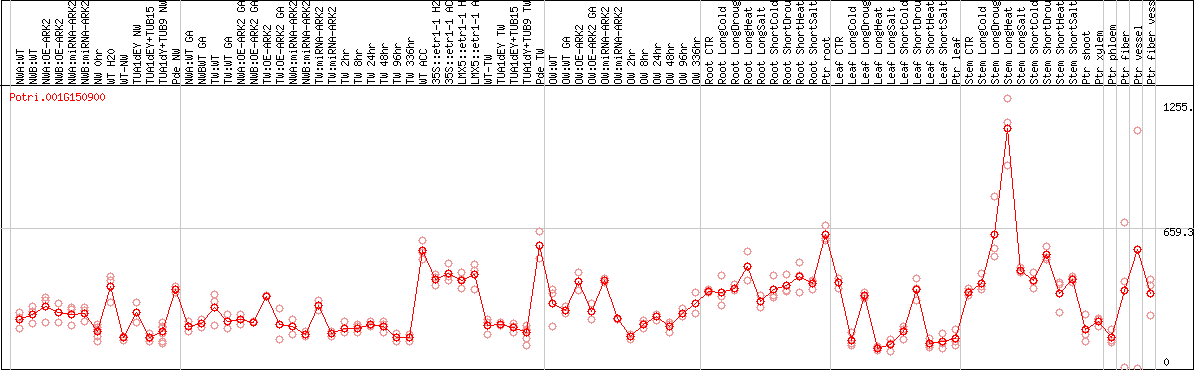
Coexpressed genes
Potri.001G150900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
