Potri.001G157200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G157200.5 pacid=42792395 polypeptide=Potri.001G157200.5.p locus=Potri.001G157200 ID=Potri.001G157200.5.v4.1 annot-version=v4.1
ATGAGGAATAGTGAAATGGATGGTTTGAGTGTAGATGGGGATGGAGAAAGAGCTAAAAATAGAGAGAATGGAGGCTCTTGGCCACCTCCGCGCATACAAC
ACAGGCGCTCTAAAAGTGCATCTGAAAGGAACTCAGTCATTTCAGGGGGTCGAGTTTTGCATTCCTTAAAGAAAGACCAGAATGAAATTCATATGTTACC
ACTTTCAACAAGGGCTCATAGAACACAAAGTCCTCTCCATGATTACCCCATTTGTAACGACAAAAATGCTTCATCTAACCACCGAGCCTCCTTGGAAAAA
GATATTGAGCAGCTGCAATTACGATTGCAGCAGGAGAAATCTATTCGCATGGTGCTCGAGAAATCAATGGGCCGAGTTTCTAGTACTTTATCTCCTGGTC
ATAGGCATTTTTCTACTCAGTGCCAGACAAAAGAGTTGATTGCAGAGATTGAGTTACTAGAAGAAGAGGTTGCAAACCGTGAGCAGCAAATGCTATCCAT
CTACAGGAATGTTTTTGAAAGTTGTGTTAGCCGGGCACCATCTAAGCAAAACTCAGGCATGCCGTCTCCTGCCCATATAAAGCATGCATCCAGGAAACAC
CCAAGTGTTATCTCAAGTGCCTTCTGCTCCTCAAAAAAGTTCCCTTTACGACCTTTGCAAGCTCTAGTTTCACCAGCTGAGTCAGGTAAAAGAACCCCTA
AAGCCAGTGATGCTCCATCATCCCATGGAAAAAGTGACATTCATTCTGAGAAGAACTGTTACAACCTGGTTAAGGTTCATGAGAGAGTACCAGCCGCAGA
GAAAACTTCCGTGCTGCAAACTCTGAAAGATCATCTTTATCAGTGCCCCAGCAAGTTGTCTGAAGAGATGGTCAGGTGTATGGCTGCAGTATATTACTGG
CTCTGTAGTGCAGAATCTGTAGCTCCTGGAAAAAACAGATCACCTTTATTGTCAAGGTCTTCCACAAATGTTATACTTCCCGGACGTGGTATCGGAGAGG
ATCGAGATTGGTCTTGCAAATCCATGGTAGAAATATCTTGGATATCAAATGATAAAAGCCAGTTCTCTCGTGCATCTTATGCCATCAACAATTATAGAGT
TCTAGTGGAGCAACTGGAAAGGGTGACAGTGACTCGGATGGAAAACAGTGCGAAAACTGCATTTTGGATCAATGTGTATAATTCGCTTGTTATGCATGCA
CATTTAGCATATGGAATTCCCCACAGCTCTCTTAGGAGGTTAGCCTTGTTTCATAAGGCTGCATACAACATTGGCGGCCATATCATTAGTGCAAATTCCA
TAGAGCAGTCAATATTTTGCCTCCGTACACCCCGAGTTGGAAGGTGGTTTGAGACCATTCTTTCCACTGCCTTGCGGAAAAAATCTGGTGAAGAAAGGCA
ACTTATTAGTTCCAAATATGGTCCATCAGATCCCCAAGCACTAGTCTGCTTTTCCCTCTGCACCGGTGCCTTTTCTGATCCCGCGCTAAAAGTCTACACA
GCATCAAATGTTAAAGAAGAACTAGAAGTCGCTAAACGAGAGTTTCTTGAAACGAATGTGGTTGTGAGGAAGTCAAAAAAAGTGTTCTTACCCAAGGTGC
TGGAAAGGTTTGCTAAGGAAGCTTCAATCAACTTGGATGATCTTCTCAAGTGGGTCGCTGAAAATGTTGACAATAAGCTTCATGATTCAATACAGAAATG
CATCGATCATAAATCAAACAAAAAGGCATCTCAGGTTGTAGAGTGGTTACCTTACAGTTCAAGGTTCCGGTACGTGTTCTCAAAGGACTTAACAGAGAAG
CCATGCTGGGTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G157200.5 pacid=42792395 polypeptide=Potri.001G157200.5.p locus=Potri.001G157200 ID=Potri.001G157200.5.v4.1 annot-version=v4.1
MRNSEMDGLSVDGDGERAKNRENGGSWPPPRIQHRRSKSASERNSVISGGRVLHSLKKDQNEIHMLPLSTRAHRTQSPLHDYPICNDKNASSNHRASLEK
DIEQLQLRLQQEKSIRMVLEKSMGRVSSTLSPGHRHFSTQCQTKELIAEIELLEEEVANREQQMLSIYRNVFESCVSRAPSKQNSGMPSPAHIKHASRKH
PSVISSAFCSSKKFPLRPLQALVSPAESGKRTPKASDAPSSHGKSDIHSEKNCYNLVKVHERVPAAEKTSVLQTLKDHLYQCPSKLSEEMVRCMAAVYYW
LCSAESVAPGKNRSPLLSRSSTNVILPGRGIGEDRDWSCKSMVEISWISNDKSQFSRASYAINNYRVLVEQLERVTVTRMENSAKTAFWINVYNSLVMHA
HLAYGIPHSSLRRLALFHKAAYNIGGHIISANSIEQSIFCLRTPRVGRWFETILSTALRKKSGEERQLISSKYGPSDPQALVCFSLCTGAFSDPALKVYT
ASNVKEELEVAKREFLETNVVVRKSKKVFLPKVLERFAKEASINLDDLLKWVAENVDNKLHDSIQKCIDHKSNKKASQVVEWLPYSSRFRYVFSKDLTEK
PCWV
|
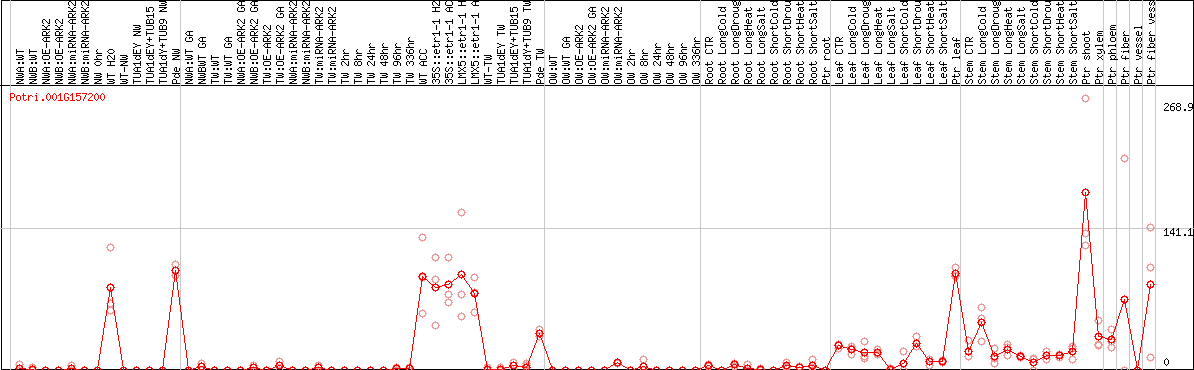
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G157200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
