Potri.001G157300 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G157300.1 pacid=42788018 polypeptide=Potri.001G157300.1.p locus=Potri.001G157300 ID=Potri.001G157300.1.v4.1 annot-version=v4.1
ATGGATTATGATGAAAATGATTTTCAAAACCACAATCTTCATTTAGCTGGTGAAGGAAGCAACAAATTTCCTTCTGTTTTACAGCCATATGCTCTTCCCA
AGTTTGATTTCGATGACAGTCTTAATGGATCTTTAAGGTTTGATAGTTTGGTTGAGACTGAGGTTTTTCTTGGCATCGAAAGTAACGAGGACAACCAGTG
GATTGAAGATTTCTCTCGTGGTACCAGTGGGATACAGTTTAGTTCAAGTGCAGCAGAGTCTTGTTCTCTATCAAGACGCAACAATGTCTGGTCTGAAGCT
ACTTCCTCAGAATCTGTTGAAATGTTATTGAAATCTGTTGGCCAGGAAGATAATACTCCTATACAAACTAATACTAAGGAGTCAGATGCCTGTGATGAAC
TGGGTTGCATACTAAAGCACATGGAGCCCATCTTGAAACAGGATAATGATACATCACCTAAAGTGGAAGATACTGCGAATTTACAGGCTACATTTCTGCC
AGGCGAGGATGTGGAAGATTTTTCTGTGTTGGATAATGATGTTGGACAGCAGCAGCCTCTTGATGATAGTTCTCAGGATCATAAAGGTGAAGCATCTGCT
GATAGTGGTTTGGGACCTTTAGTTGACCCATCTGCTGTTAGTGTAGAGGTCAGGCAACCTGTTATTGAAGGGAGTCTGTCCATTGATAGTAAATCTAATC
ATGTTACTCAAAGGGAAATCGATAATGTCGTGAATGGATCTTCAAATGATAGGCCACAGAAAGTTCCTGCTTCAGGGATGCAGGATGGTGCCTCTGTGCA
AAATATCACTACAGGAAATATTGAATTGAATGAAAAAGATGGCCCTGATGATATAAATAACACTTCTGATGATAGTAAAGATTTTCTGGAAACAGACACT
GGTGAGAATCAGAAAAAGGGACAAGTATTAAGTCAAGAGGGCCAAATGGAGGATGAGAATCCTTGTTCAGATGCAGTGGAATCTATGGAAGAAGCAAACG
TCATTGAAACTAATTCGAGTAATTTGGGGGAACCTTCTTGCAAAATACTGAAGGGGCATTCTGGTTTCCCAGAAGACGTAGTGACTAGTGATCAGTCCGA
AGTGGATACAGTTGGGGGATCAGTGATGGCTGTTGAAGGTAATACTACTTTTAAGAGGGATGAAATTGAGGACTCAAATGGTTCCCAATTGGATAACAAG
AACCTATCTAATAAGTGTGAGGGATCTCTCCTGTCTGCTGAAGACTGTGAGCCTGCTAAAGTGAAAGTTGGTGGAACTAGCAGCAGCGATACAGGTGGTG
TTTCTAGTTTGGCTACAGTGTGCTGTTCAGCTGAAGTTGTTGGAGAAGTGGCCCATGTTTCATCCTCTTTCCTTGTTGAATCATCACAAATATGTGGGAA
GTCTATGGTTTCTGCTGAGGGAAAGGAGACCACAGAGCTTCCTTCTGGCAATGTTAGCACTGAAAACAATTTCATAGCTTCGAGATTACAATCAGATGCT
GCTTCTGACAACAATTCGGCATCAGATGTTAGCTGCGAACATGCTAACATGGTAACCTGTGCTACAATGGATGGTGTTCCAGCGCCTTCTGGTGATGTCA
CTAATGTAGATGCAGTTATTGGTCACAAAGATGTCAAAATGTCACTTTTAAGTGAGATGGGGTTTAGTCCCTTAGATATAGAGAAAGAAACCGTGGACAA
GATTTCTGTGGAGGCCAGTTTATCAGGTCTGAAGACATCTTGTCAAGTGATAGCTGGATTAGATCCTGGTTCTGAATCCAAAAAAGGCGCTTCTTCTGGT
GCTGCAGGACAGATATTATGCGAGTCAGCTGAACAATCTCCATTAATGGTGGATGCTAGTAAAACAGAAGGACCTCATTCAGAAGTAATTGATAAGGTCA
GCCTGCAGAGCACAAAGGAAATGAATGTGTGTCCAGTTCTTTGTGATTCAACTGCAAACAAGGGTGATGATGCTGAAGTGTTAGTAAAAGAAAATGATGA
AAAGGAATCATCCAAAGTTTCAGAACCGACTGTAAACAAGAATGAGATGCTAGGACCTATCTCCTCAGAAAAGGAAGAGTGTCGAGAGGATACTAACCAG
AAAGGTCAGGAAGAAAATGAAGCTGCCATAGTGTCTGAAGATAATAGTGATGGGAATATTGCTGTCCCTAGCACAAACGATTGTGGAAGTTGTGCTGATG
TTGGCAAAGCAGCCAGTGGTTCCCCTACTGTCATCAGAGCTGCCAGAGATTTTCAGAGTGAAAGTGACAAGGATGGAGCCAAATGTTCAGTTGAGCAGAC
TGCTGTTGCTGATAGCAATGCCAGCAAAGCTTTGTCTGGTTCTCGGGATCCGAAACAAAATGATGCATCCAAAGATGAAAGGAGCTTCACTTTTGAGGTC
AGTCCATTGGCAAATATGCCTCAAAAAGAAGTTGGCAATAAATGGCAGCCCTTTTTGAACAAACCAGCCACTAAAGCCTACCCGATTTTGAATGCTTCTC
CATCATCGGGTTTAGTCCAAATAGATCCCAAGCTTGCCCAAGATCTTCCTCATGGAAGTCCAAAGGTGTCTGATGTAGCCATTGTGCGTAGTGGTTCTAA
AGGTACTTCTGAGCGTAAAACAAGACGATCATCTGGTAAGGCCATGGAAAAGGAAAGTGCGAGAAAGGGAAATCCTATTAAAGATACTGCTTCTGTGAGG
CTAGAAAAGGGGGCAAAAACAAACAATGTCTCCCCAAGCTCTTCTGGGATATTGCAACATGTGCAATCAAATGAGATGCAGCGCTATGGTCATGCGGATT
CCAGTACTATGAAACCATTTGTTCATGCATCTTCAAGCCTTCCAGATCTGAATTCTTCTGCTTCTCCATCTGTGATGTTTCAACAACCCTTCACGGACTT
GCAACAAGTGCAATTGCGTGCTCAGATCTTTGTTTATGGCGCTTTGATACAAGGAACAGCTCCAGATGAAGCATATATGATATCAGCATTTGGAGGATCT
GATGGTGGGAAAACCATATGGGAGAATGCTCTACGCTCTTCTATAGAACGGCTTCATGGTCAAAAACCTAATCTCACTTCTCCAGAAACCCCTTTGCAGT
CACGTCCTGGTGTCAGAGCTCCTGATCAAGCAATTAAACAGAGTACTGTTCAAAGTAAAGTAATCTCTTCACCTATTGGTCGATCCAGCAAGGGTACTCC
AACAATTGTGAACCCCATGGTACCCCTTTCTTCACCGCTCTGGAGTGTACCTACCCCCGCTGGCGACACATTTCAATCAAGCAGCATGCCAAGGGGTCCC
ATTATGGATCATCAGCGGGCACTTTCTCCTATGCATCCTCATCAAACTCCACAGATAAGAAATTTTGCTGGTAATCCTTGGCTATCTCAGGCACCTTTTT
GTGGTCCCTGGGCTACATCTCCACAAACACCTGCACTTGACACAAGTGGTCATTTTTCTGCGCAATTGCCTATCACAGAACCAGTTCAATTGACACCTGT
GAAAGACTTATCCATGCCAATTATCTCTGGTGCAAAGCATGTTTCTCCTGGTCCTGTAGCTCAGAGTGGGGCTTCCACCAGTGTTTTTACTGGGACTTTT
CCTGTTCCAGATGCAAAAAAGGCTGCAGTGTCATCCAGTCAACCTCCTGCTGATCCAAAACCTAGGAAAAGAAAAAAGAACTCGGTTTCTGAGAGTCCGG
GCCAGAATATTTTGCCTCCTCACCTTCGAACAGAATCAGTTTCTGCTCCTGTTGTTACCAGTCATCTGTCTACTTCTGTTGCCATTACAACCCCTGTCAT
CTTTGTTTCTAAAGCCCCTACAGAGAAATTTGTGACTTCTGTATCTCCTACACCTACTGATATCAGAAACGGGAATCAGAATGCAGAACAGAGGAATATT
TTGTCAGAGGAAACCCTTGATAAAGTGAAGGCTGCGAGAGTGCAGGCAGAGGATGCTGCTACTCTTGCTGCTGCTGCTGTTAGTCACAGCCTGGAAATGT
GGAATCAGTTAGATAAGCAGAGAAATTCTGGGTTATCACCAGATATTGAAACCAAACTAGCTTCTGCAGCTGTCGCAATAGCAGCAGCTGCTGCTGTTGC
AAAAGCGGCAGCGGCAGCTGCCAAAGTTGCATCTAGTGCTGCATTGCAAGCAAAACTGTTGGCTGATGAAGCAGTAAATTCTGGTGGTTACAGCAATCCC
AGTCAAGACAATACAATTTCTGTTTCTGAAGGCATGAAGAATTTGGGAAAGGCTACTCCTGCTTCCATCCTGAAGGGTGATGATGGGACAAACAGTTCCA
GTTCAATCCTTATTGTGGCAAGGGAGGCTGCTAGACGGAGAGTTGAAGTTGCTTCCGCTGCTGCAAAACGAGCTGAAAATATGGATGCTATTGTTAAAGC
TGCGGAGCTGGCTGCAGAAGCTGTGTCCCAAGCTGGGAAGATAGTTGCTATGGGCGATCCTTTGCCACTGAATGAGCTAGTAGCTGTGGGTCCAGAGGGA
TATTGGAAAGTAGCCAAAATAAATAATGAACTGATCTCTAAATCAAATGATATAGGTAGAAAAACTCTTAATATAGATAGAGTTGGAGAAAGACCACGCA
CTCCAACTGAGGGGTCCACTGAGGACCATGTTAGATTGGAAGATGGTTTCTTGAGTTCTGGTGCAGCTGCTGCAAAGGATGTTAAAGGACAAAAGGGCTA
CAAGGTGTCTGAATCTGAGAATGGTTTAAGATCTTTGGGAACTATAGAAAACTTTAATAGCATCAAGGAGGGGTCCCTTGTAGAGGTTTTCAAAGATGGG
AATGGATTTAAAGCAGCCTGGTTCTCTGCCAATGTCGTGGATTTAAAGGATGGGAGTGCATGTGTGAGTTATACTGATCTTTCATCAGTTGAAGGCTCAG
AGAAGCTAAAGGAGTGGGTGACACTTAAAGGTGAAGGAGAGAGAGCGCCAAAAATACGAATAGCTCGCCCTATTACTGCCGTGCAATTAGAAGGAACAAG
GAAGAGGCGACGAGCTGCCACGGTGGACCATATTTGGTCTGTTGGTGATAGAGTGGATGCATGGATACAAGATAGCTGGTGGGAAGGAGTGGTCATTGAA
AGGAGCAAGAAAGATGGCACCACACTAACAGTCCAATTTCCAGTTCAAGGAGAAAAATCTGTTGTTAGAGCATGGCATCTTCGGCCCTCTCTCCTATGGG
AAAATGGGGAATGGATTGAATGGTCCAGTTCAAGAGTGGGCTCTCATTCTACCAACAAGGGTGATACTCCACAGGAAAAACGACCTAGGGTGCGGAGCCC
TGCAGTGGACAACAAAGGGAACGATAAGTTGTCAAAAGGTTTTGATTCTGTGGAAACTAATAAACCTGATGAGCCAACTTTATTGGATTTAGCTGCTCAT
GAAAAATTGTTCAATATTGGTAAGAGTACCAAAGATGGTAATAAACCTGACGTGCTCAGAATGGCACGGACTGGCTTGCAGAAGGAAGGATCAAAAGTGA
TCTTCGGTGTACCAAAGCCTGGGAAAAAAAGAAAGTTTATGGAAGTGAGCAAGCACTATGTTGCAGATCAGAGCAGCAAAAACGACGACGCAAATGATTC
GGTAAAGTTTGCTAAATATTTGATGCCTCGGGGATCAGGATCCCGTGGATGGAAGAATACTTTAAGAACTGAATCAATCGCAAACCGAACAGCGGCATCG
AAGCCCAAGGTTTTCAAGTCTGGAAAGCCACAGAATGTTTCTGGTAGAACAATTACTCAGAAGGACAACTCATTGACAACTACAGTTTCTGCTTCCAATG
ATGGTGCTGTTACAGATCATGTTGCAAAGACCAAGGCTTCTATAAGCCATGTTGAAAATACATCCGAAAAACGTAACTTGACTGACTTTCAGCCCTTATC
TAGCTCCGTGGGAGCAACAGAGGGTCCAATATTTTCTTCCTTTCCTCTTCCTTCAGGTACACTTTCCTCTAAGAAAACTTCCACATCAAATGCTAAACCT
CAACGGGTAAGTAAAGGAAAACTTGCTCCTGCTGGTGGAAAGTTGGGTAGAATTGAAGAGGACAAAGTTTTCAATGGAGATTCTTCAAAATCAAACTCTG
ATGTCACAGAGCCACGAAGGTCTAACCGTAAAATGCAGCCTACATCAAGACTGTTAGAAGGGCTACAAAGCTCCTTGATGGTGTCAAAAGTTCCAGCTGT
GTCGCATGATAAAAGTCAGAAAAGTAGGACTGCTTCTAGAGGGAATAACCATGGTTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G157300.1 pacid=42788018 polypeptide=Potri.001G157300.1.p locus=Potri.001G157300 ID=Potri.001G157300.1.v4.1 annot-version=v4.1
MDYDENDFQNHNLHLAGEGSNKFPSVLQPYALPKFDFDDSLNGSLRFDSLVETEVFLGIESNEDNQWIEDFSRGTSGIQFSSSAAESCSLSRRNNVWSEA
TSSESVEMLLKSVGQEDNTPIQTNTKESDACDELGCILKHMEPILKQDNDTSPKVEDTANLQATFLPGEDVEDFSVLDNDVGQQQPLDDSSQDHKGEASA
DSGLGPLVDPSAVSVEVRQPVIEGSLSIDSKSNHVTQREIDNVVNGSSNDRPQKVPASGMQDGASVQNITTGNIELNEKDGPDDINNTSDDSKDFLETDT
GENQKKGQVLSQEGQMEDENPCSDAVESMEEANVIETNSSNLGEPSCKILKGHSGFPEDVVTSDQSEVDTVGGSVMAVEGNTTFKRDEIEDSNGSQLDNK
NLSNKCEGSLLSAEDCEPAKVKVGGTSSSDTGGVSSLATVCCSAEVVGEVAHVSSSFLVESSQICGKSMVSAEGKETTELPSGNVSTENNFIASRLQSDA
ASDNNSASDVSCEHANMVTCATMDGVPAPSGDVTNVDAVIGHKDVKMSLLSEMGFSPLDIEKETVDKISVEASLSGLKTSCQVIAGLDPGSESKKGASSG
AAGQILCESAEQSPLMVDASKTEGPHSEVIDKVSLQSTKEMNVCPVLCDSTANKGDDAEVLVKENDEKESSKVSEPTVNKNEMLGPISSEKEECREDTNQ
KGQEENEAAIVSEDNSDGNIAVPSTNDCGSCADVGKAASGSPTVIRAARDFQSESDKDGAKCSVEQTAVADSNASKALSGSRDPKQNDASKDERSFTFEV
SPLANMPQKEVGNKWQPFLNKPATKAYPILNASPSSGLVQIDPKLAQDLPHGSPKVSDVAIVRSGSKGTSERKTRRSSGKAMEKESARKGNPIKDTASVR
LEKGAKTNNVSPSSSGILQHVQSNEMQRYGHADSSTMKPFVHASSSLPDLNSSASPSVMFQQPFTDLQQVQLRAQIFVYGALIQGTAPDEAYMISAFGGS
DGGKTIWENALRSSIERLHGQKPNLTSPETPLQSRPGVRAPDQAIKQSTVQSKVISSPIGRSSKGTPTIVNPMVPLSSPLWSVPTPAGDTFQSSSMPRGP
IMDHQRALSPMHPHQTPQIRNFAGNPWLSQAPFCGPWATSPQTPALDTSGHFSAQLPITEPVQLTPVKDLSMPIISGAKHVSPGPVAQSGASTSVFTGTF
PVPDAKKAAVSSSQPPADPKPRKRKKNSVSESPGQNILPPHLRTESVSAPVVTSHLSTSVAITTPVIFVSKAPTEKFVTSVSPTPTDIRNGNQNAEQRNI
LSEETLDKVKAARVQAEDAATLAAAAVSHSLEMWNQLDKQRNSGLSPDIETKLASAAVAIAAAAAVAKAAAAAAKVASSAALQAKLLADEAVNSGGYSNP
SQDNTISVSEGMKNLGKATPASILKGDDGTNSSSSILIVAREAARRRVEVASAAAKRAENMDAIVKAAELAAEAVSQAGKIVAMGDPLPLNELVAVGPEG
YWKVAKINNELISKSNDIGRKTLNIDRVGERPRTPTEGSTEDHVRLEDGFLSSGAAAAKDVKGQKGYKVSESENGLRSLGTIENFNSIKEGSLVEVFKDG
NGFKAAWFSANVVDLKDGSACVSYTDLSSVEGSEKLKEWVTLKGEGERAPKIRIARPITAVQLEGTRKRRRAATVDHIWSVGDRVDAWIQDSWWEGVVIE
RSKKDGTTLTVQFPVQGEKSVVRAWHLRPSLLWENGEWIEWSSSRVGSHSTNKGDTPQEKRPRVRSPAVDNKGNDKLSKGFDSVETNKPDEPTLLDLAAH
EKLFNIGKSTKDGNKPDVLRMARTGLQKEGSKVIFGVPKPGKKRKFMEVSKHYVADQSSKNDDANDSVKFAKYLMPRGSGSRGWKNTLRTESIANRTAAS
KPKVFKSGKPQNVSGRTITQKDNSLTTTVSASNDGAVTDHVAKTKASISHVENTSEKRNLTDFQPLSSSVGATEGPIFSSFPLPSGTLSSKKTSTSNAKP
QRVSKGKLAPAGGKLGRIEEDKVFNGDSSKSNSDVTEPRRSNRKMQPTSRLLEGLQSSLMVSKVPAVSHDKSQKSRTASRGNNHG
|
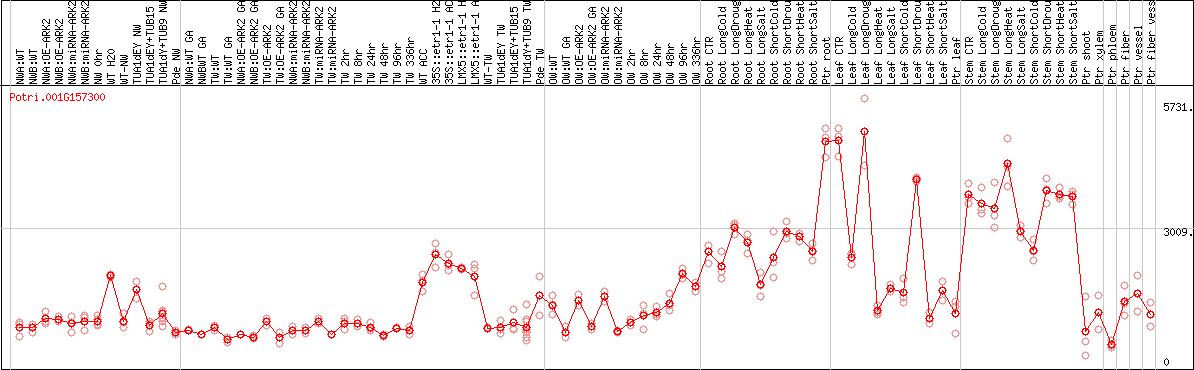
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G157300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
