Potri.001G164900 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G164900.7 pacid=42789281 polypeptide=Potri.001G164900.7.p locus=Potri.001G164900 ID=Potri.001G164900.7.v4.1 annot-version=v4.1
ATGAAATCCAATACACTTCTCGACTATGCTGTGTTTGAACTGTTGCCAAACCTGACACGATGTGACTTGTTTGTTTCAAGCAATGGAAACACAGAGAAGC
TTGTGTCTGGACCAGTAAAGCCATTTATAACTCATTTGAAGGTTGCAGAAGAACAAGCTTCACAGGCAGTCCAGTCAATTAAACTAGAGGTTGACCGGCG
TAGAAATGCTGAGACCTGGTTCACAAAAGGAACACTTGAGAGGTTTGTGCGTTTTGTCAGTTCACCGGAGGTTTTAGAGATGGTCAATACATTTGATGCA
GAGATGTCTCAGTTAGAAGCAGCCCGGAAAATATATTCTCAGGGGTCTGGAGATCAACTTTCTGGTGCTTTAGGCAGGGATGGAACAGGAACCACAGCAG
GAGCTGATGCAACAAAGAAGGAGCTTCTAAGGGCGATTGATGTGAGGCTTGCTGCAGTTAGGGAGGATTTAGCCACAGCCTATGCTAATGCATCTGCTAC
TGGATTCAACCTTGACACTGTTCCAGACCTCCAGCATTTTGCAGATCGCTTTGGTGCCCATCGCTTGAATGAGGCTTGTACCAAATTTATGTTACTGTGC
CAGAGAAGACCAGACCTGACCAATCCATGGAAGCCAATTGTGGGGGATCAAGTGGTCCGGTCCTCGTGGGGATCTGACATGTCAATTGATGACCCCACTG
AAGATGAAAGTGGTTCCTACCTGACCAGGCCCCACCAAAACCCATTCCAAAATAAACACCAACAGCAGCAAGCTAGCCAAGAACTACAACAAATAGAAAC
GACACAAACCCAATTCCACCTTAACCAATCCAAGTCCTCCACTTATCAACAGCCCAATTCCTCTCTCGCTACGCAGCAACAGACTATTCAGAACGAGAAC
AAGGAAGAAGAAAAGAAGAAAGAGGAGGCTGTGACTAATTCATCAACAAGTCTACCAAGCCAATCTTCTAGGAGACTCAGTGTTCAGGACAGGATCAACC
TATTTGAGAACAAGCAGAAGGAGAGCTCAGGTGGCAAACCTGGAGCTGTAGGTAAATCTGCAGAGCTGAGGAGACTATCATCTGATGTTTCACCTGCAAC
TGCAACTGCAACTGCAACTGAAAAGGCTGTGTTGAGGAGATGGAGTGGTGCAAGTGATATGAGCATTGACTTGGGCAATGACAAGAAGGATGATAACAAT
ATAGACAGCCCCTTGTGTACACCCTCTTCATCCTCTGTTTCTGGAACCAAAAGTAATGTGTTTCCTGTCTCATCAGATGATGATAAGGACCAGAAAGGGT
TGAATGATACAGAAAGTGCAGCAAATTTAGTTAAGTTGGAGACTAAGAGCCTTTCTGGGTTGAAAGATCAGGGTGACTTGCAAACTCATGGTGGTGGGCC
TGCGAGGAAGGAGAAGGAGGTGAATTTGAAGGGGAAAGTGAATTTGAAAGATCAAGTAGGGTCTCTGGCCCAGTTGAGGTCATCTGCAGGTAGAGGGGAG
GAGTCAGGGGTGGGTGATCAGGTGGTTTTGGAGAAGTTGAAGGGTACTTCAGGTGGGGAAGAGAGGACTGTTGGAGCTAAAGCTCAACTGAGTTTTCAGG
AGAAGTCGAGAGGTTTTCCAGATAAGGTGGAGATTGTTGCAGTGAAAAACCAGGTTGATCTGCAGACCCAGATTGGAGGTTTTGTGGGCAGGGTTGGGAA
TGTGGCGTCTGGGAATAGAATTGACGACATTAAAATAAGGGATCAGTCCTCGAGTCAGTCAAGGTCTGGGGTTTCTCAGACTCATACTAGATCTTTTTCA
GGACAGTTTGAAGGTGGTTTTGGAGTGAAGGATAAGGAATTGCCAACTAAAGTGACTGACTTGGATCTGTCAGCTTCTCAGACACAGCAGAAATTGTTTA
AGGGTGAAGTTGATCAGGCAAGGAAGGAGGACACTGAACAGATAACAGAGGATGATCTTGAAGTTTCAAAAATGAAGGTCCAGAAACAACCTTTTTTGGG
TCCAGAGCAATTTAGGAAGCTGCAGGGTAGGAGGGATGAGAGTGGATCTATTCATGGAAGCAACAAGCCTAGCTTTCCCAGTAAAAAGTATTCTGAGAGT
CAAGAGAGCATTGGTTCTCAGCAAGTACCATCAGCAGATCAGTTTCAAAGGGTGAGGCAGTCTAAGGGGAATCAAGAATTAAATGATGAGCTGAAGATAA
AGGCAAATGAGCTTGAAAAGCTTTTTGCTGAACACAAACTTCGCATTCCTGGTGATCAATCCAGTTCTGCTCGGAGAGGCAAGCCTAGTGAAGTGCAATC
TGAACAGGCAGCAAGTTTACAGTACAGAAAACCAGTAGCTGTAGAGATATCTCCTGTTCAGTTTCAGGAAAAGACGGTGCTTGAACGAACTGGGAGTTCT
AGTGATACTGGAAAGTTTAGCACTCCACCAAGGAAGATAGTAGATCATCAGGACTGTGGCAGTTCTCTGAGGCAGAGTTTCTCTGAGATTAGTTTTTCAG
ATGATTCTAGAGGGAAATTTTATGAGAGGTACATGCAGAAAAGAGATGCAAAACTGAGGGAGGAGTGGGGTACAAAAAGATTGGAGAAGGAAGCCAAGTT
GAAGGCCATGCAGGAAAGTCTTGAGCGGAGTAGAGCTGAGATGAAGGCAAAATTTTCTTGCTCTGCAGACAGACAAAATTCCTTGTCTGATACTCACAGA
TGTGCAGAAAAGCTCAGATCATTCAATTTTAATTCAAGTACGAAGAGAGAACAGCCAGTAGATTCCATTCATAGTGAAGAGGATGAAGATCTTTCCGAAT
TTCCAGAACAGATTTATTATGGAGAGGACAGATCATTCAATGAGGTGTCTTTAGGGGGCATTGCTTCTAGAAGCTCTCAAAACAAGAAGCTCTTGCTGAA
CAGAAATTCATCTTCATCCACCCCTCGCACCACAGTTGTGCCAGTTCCACGGTCATCATCCAAAATTTCCAACCCAAGTTCCGGGAGGCGGAGAGTACAA
TCAGAAAATCCTCTCGCACAGTCTGTTCCAAACTTCTCCGATTTCAGAAAAGAGAACACAAAGCCCTTGTCTGGAGTTAGCAAAGCAGCAAATCGCTTGC
AGGTGAGAACCTATGCTCGCAGCAAGAGTTCCAGTGAAGAGATACCTCTGGCCAAGGAAGAAAAGAACCAGCGGTCTCAATCCTTGCGCAAAAGCTCTGC
CGGTCCTATTGAGTTCAAGGATTTGCCCCCATTAAACTCGGATGTTGTTTTGGCACCATTGAAATTTGATAAAGAGCAAACTGAGCAGATCCCATATGAC
AAATTCTCAAAGAATGTGGAGTCAAAGCCTTTCCTCAGGAAGGGCAATGGCATAGGTCCTGGTTCTGGAGCAACTGTTGCGAAACTGAAAGCTATGGTAG
CATCTGAGACTTTAAAAAATGAAGAATTTGAAGAATCAGCCTTTGAGGCAGAAGATTCAGTTGATGAGTCAAAGGAAGAAGAAGACGAGGGACTTGAAAC
AACCGAGATTGAAGATTGTGCTAATATGGATAATGGCAAACCAAGACTAAGCCTGGATTCAGACAAGATGGGTACATCTGGATCCGAGAATGATGAATCT
CTGAGATCTATTTCTCAAATTGATCCTTCTTCAGTTGCTGAATTGCCTGCGTCTGTGCCTTCAACATTCCATGCTGTAGGGTCTCTGCAGGACTCACCTG
GGGAAAGCCCTGTGTCATGGAACTCACGCATGCAACATCCATTTTCCTACCCACATGAGACCTCAGATATTGATGCTTATGTGGACTCTCCAATTGGGAG
TCCTGCATCATGGAATTCTCATTCTCTGACCCAAACAGAGGCTGATGTAGCTCGGATGAGGAAGAAGTGGGGAAGTGCGCAGAAACCTATTCTTGTTGCC
AATTCATCCCATAATCAGTCTCGCAAGGATGTGACCAAAGGGTTCAAACGGTTGTTAAAGTTTGGAAGGAAAAGTCGTGGGGCAGAGGGTTTGGTTGATT
GGATTTCTGCTACCACTTCTGAAGGAGATGATGATACAGAAGATGGGCGAGATCCTGCCAATCGGTCATCAGAAGATTTGAGGAAGTCAAGAATGGGATT
CTCACAGGGTCACCCTTCAGATGATGGCTTTAATGAAAGTGAGTTGTTCAATGAGCAAGTTCAAGCATTACATAGCTCCATACCAGCACCTCCAGCAAAC
TTTAAATTGAGGGATGATCATTTGTCTGGAAGCTCTATTAAAGCACCACGGTCATTCTTCTCTCTGTCATCCTTCCGAAGCAAAGGTAGTGACTCAAAGC
TTAGATAA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G164900.7 pacid=42789281 polypeptide=Potri.001G164900.7.p locus=Potri.001G164900 ID=Potri.001G164900.7.v4.1 annot-version=v4.1
MKSNTLLDYAVFELLPNLTRCDLFVSSNGNTEKLVSGPVKPFITHLKVAEEQASQAVQSIKLEVDRRRNAETWFTKGTLERFVRFVSSPEVLEMVNTFDA
EMSQLEAARKIYSQGSGDQLSGALGRDGTGTTAGADATKKELLRAIDVRLAAVREDLATAYANASATGFNLDTVPDLQHFADRFGAHRLNEACTKFMLLC
QRRPDLTNPWKPIVGDQVVRSSWGSDMSIDDPTEDESGSYLTRPHQNPFQNKHQQQQASQELQQIETTQTQFHLNQSKSSTYQQPNSSLATQQQTIQNEN
KEEEKKKEEAVTNSSTSLPSQSSRRLSVQDRINLFENKQKESSGGKPGAVGKSAELRRLSSDVSPATATATATEKAVLRRWSGASDMSIDLGNDKKDDNN
IDSPLCTPSSSSVSGTKSNVFPVSSDDDKDQKGLNDTESAANLVKLETKSLSGLKDQGDLQTHGGGPARKEKEVNLKGKVNLKDQVGSLAQLRSSAGRGE
ESGVGDQVVLEKLKGTSGGEERTVGAKAQLSFQEKSRGFPDKVEIVAVKNQVDLQTQIGGFVGRVGNVASGNRIDDIKIRDQSSSQSRSGVSQTHTRSFS
GQFEGGFGVKDKELPTKVTDLDLSASQTQQKLFKGEVDQARKEDTEQITEDDLEVSKMKVQKQPFLGPEQFRKLQGRRDESGSIHGSNKPSFPSKKYSES
QESIGSQQVPSADQFQRVRQSKGNQELNDELKIKANELEKLFAEHKLRIPGDQSSSARRGKPSEVQSEQAASLQYRKPVAVEISPVQFQEKTVLERTGSS
SDTGKFSTPPRKIVDHQDCGSSLRQSFSEISFSDDSRGKFYERYMQKRDAKLREEWGTKRLEKEAKLKAMQESLERSRAEMKAKFSCSADRQNSLSDTHR
CAEKLRSFNFNSSTKREQPVDSIHSEEDEDLSEFPEQIYYGEDRSFNEVSLGGIASRSSQNKKLLLNRNSSSSTPRTTVVPVPRSSSKISNPSSGRRRVQ
SENPLAQSVPNFSDFRKENTKPLSGVSKAANRLQVRTYARSKSSSEEIPLAKEEKNQRSQSLRKSSAGPIEFKDLPPLNSDVVLAPLKFDKEQTEQIPYD
KFSKNVESKPFLRKGNGIGPGSGATVAKLKAMVASETLKNEEFEESAFEAEDSVDESKEEEDEGLETTEIEDCANMDNGKPRLSLDSDKMGTSGSENDES
LRSISQIDPSSVAELPASVPSTFHAVGSLQDSPGESPVSWNSRMQHPFSYPHETSDIDAYVDSPIGSPASWNSHSLTQTEADVARMRKKWGSAQKPILVA
NSSHNQSRKDVTKGFKRLLKFGRKSRGAEGLVDWISATTSEGDDDTEDGRDPANRSSEDLRKSRMGFSQGHPSDDGFNESELFNEQVQALHSSIPAPPAN
FKLRDDHLSGSSIKAPRSFFSLSSFRSKGSDSKLR
|
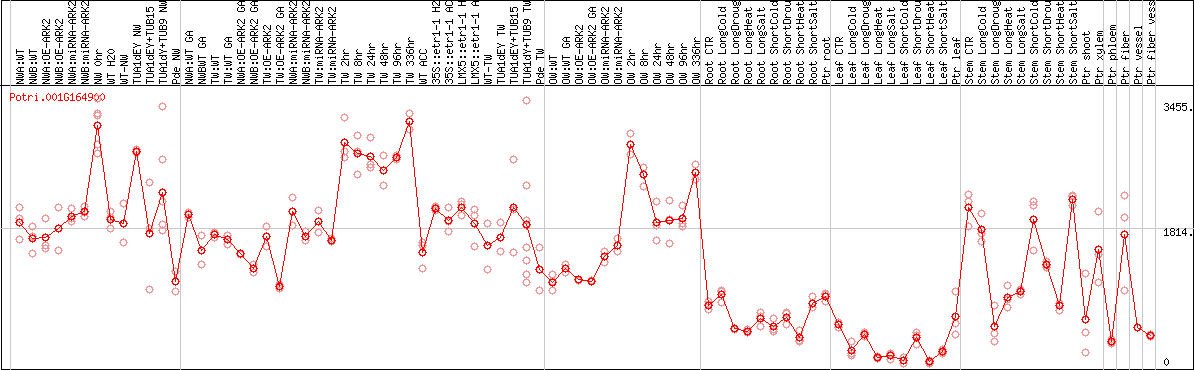
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G164900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
