Potri.001G165000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G165000.1 pacid=42792516 polypeptide=Potri.001G165000.1.p locus=Potri.001G165000 ID=Potri.001G165000.1.v4.1 annot-version=v4.1
ATGGCTGATTCCTCCTTTGAAATTGATTACCACTCCACCGGCCGCTCACGAAACTATCATCAGCGTAGGCAACTCAGGCACCCCAGCACTCCTGTTAACT
CACAGTATGATTCACGTTCATTTACACAATTCAGCTTTACAAATACTCCAAATAGGCTTCGCCAAACTCCTTCCACGCCTTTCGCCACGGACAATGATAC
GTCATGGCAAGATGAGCTTTCATGGCAGTTTCAGCCCACAGGGTGGAATGATACCCGCAGCCTGGGAGCAGCTCTTAGCCCATGGGCAGCTAGCACACCA
TCAAACAGACATATTTTTCAAAGATCAGCAAATGACTACTATCTTTCACGTACACATGGTGGATTTAGAACCTTCACAAATCCATACTACGATCAATCTA
GCTATGGTGCTGTACCTGCTGGGAGGCTTGAGCTACAAAGCTACGCTGCCAGAAATAATGAGAGGTCGGTAGTTCATGTTAGGGATTATAGCTCGGCTGC
TTACAGTAAATCCCATCATGGGATTTCAAGACCAATATCTCAGGCTATTAAGGGAGGGGCTCGCAGAAATGCAAGTCCTTTGGTTGATCAAGATGAGCTT
AGCATGATTGACTATGACAGCGAGGATGTTGAGAAACAGGGAGAGTTATTGCAGACTGACACTAATCTCCATGGTGATAAAGATTCTAGATGGATTTCAG
TGTCTCATGCTTATATGGAAGATGATGGTGTTAGTCCATTGTACCACAGTACTCCACACGGTGGGCACGATCATCATGGTCATGAATTATCCCGTAGTAG
ACATGATGATTTACTCAGTGCTTATGAAGCTAATCGATCAACCAGTCGTGATTACGTTCCCGGTAAATATCCATATGATGACATTGATCAGGCTTCGGAG
TATGAAGATGAAGATTATGATGAGGAGGATGATGACAACGAAGAAGCAGCACGAAGGGAAGTAGGGTTGTTTAGTCTGTTCAAATACTCGACGAAATGGG
ATATGGTTCTTGTGTTTTTGGGTTGTTTAGGAGCTCTCATAAATGGAGGATCTCTTCCATGGTATTCTTATTTTTTCGGAGATTTTGTGAACAGGATTGC
GAAGCACTCTGATGATAATATGATGAAAGAAGTGGAGAGGATATGTCTGCTTATGACTGGAGTAGCAGCATTAGTTGTGGTTGGAGCCTATTTAGAAATC
ACTTGTTGGAGATTAGTTGGAGAACGATCAGCTCATCGAATAAGAAACTTGTATCTGAGCGCAGTTCTAAGGCAAGATATCACTTTCTATGATACAAAAG
TCAGCACTAGTGATATTATGCATGGAATTTCAAGTGATGTTGCACAAATCCAAGAAGTGATGGGGGAGAAGATGGCACATTTCATCCATCATATATTTAC
CTTCATTTGCGGTTATTGGGTCGGGTTCTTAAGGTCATGGAAGGTATCTCTGGTGGTTTTGTCAGTCACTCCATTAACGATGTTTTGTGGCATTGCTTAT
AAGGCCATTTATGTCGGTCTAGCTACAAAAGAAGAGGTTTCTTATCGGAAAGCTGGTGGCGTAGCTGAGCAGGCAATCAGTTCAATTAGGACTGTATTTT
CTTTTGTTGCTGAGGATAAGTTGGCCAGGAAATATGCTGATTTATTGATGAAGTCGGTGCCTATCGGTGCAAAGATTGGGTTTGCTAAGGGTGCTGGAAT
GGGAGTTATTTACTTGGTTACCTACTCAACATGGGCACTGGCCTTTTGGTATGGATCTATCCTGGTTGCAAGGAAAGAGATAAGTGGAGGTGATGCCATA
GCTTGCTTCTTTGGTGTCAATGTAGGGGGAAGAGGATTGGCATTGTCTCTATCATATTTTGCTCAATTTGCGCAAGGCACAGTAGCAGCAACCAGAGTAT
ATGAAATTATAGACAGAATTCCTGATATAGATCCTTACAGCCCCCATGGAAGGATACTGTCAACTGTTGGTGGAAGGATCGAGATCAAAGGCGTTACTTT
TGCTTACCCATCGCGTCCTGAAACTGTAATTCTCCGCTCTCTTAATCTAGTTATTCCATCCGCAAAGACTCTTGCACTGGTTGGTGCTAGTGGTGGTGGC
AAGTCCACCGTTTTTGCTCTCATAGAGAGGTTCTACGATCCCATCAATGGAGTAGTAACTTTGGACGGTAATGACCTGAGGACACTGCAAGTCAAGTGGC
TAAGAGGTCAAATAGGCATGGTTGGGCAGGAGCCAGTTCTGTTTGCCACCAGCATACTAGAAAATGTGATGATGGGAAAGGAGAATGCCACAAAGAAAGA
AGCAATCAATGCTTGCATTGCTGCCAATGCTCACAGCTTTATCTCTGGCCTTCCCTTTGGCTATGACACTCAGGTTGGAGATAGGGGAACCCAACTCTCA
GGGGGGCAAAAACAGAGAATTGCACTAGCTAGAGCTATGATTAAGAATCCTAGAATCCTTCTTTTAGATGAGCCCACCAGTGCTCTGGACCAAGAGTCTG
AATCTGTTGTTCAGCAAGCCATCGATAAGATATCCACGGGCAGAACAACCATTGTCATTGCTCATAGGCTAGCGACAGTAAGAAATGCCAACACCATTGC
TGTTCTTGATCAAGGCTCTGTTGTTGAAATTGGTGACCATCGCCAGCTAATGGAAAATGCTGGTGCGTACTATGACCTTGTCAAACTTGCCACTGAAGCT
GTTTCAAAATCTGCATTGAAACAGGAGGATGCTGCCAAGGATATGGAGTTCTCTATTTACGAGAAATCTGTTGATTTGAGATCTAAGAATGCATTTGAAA
CCTCAAAATCAAGGTACTTAAAATCAATGCAAGCAGAGAATCAACAAGAGGAGGAAATGCAGGAAAGTGCAAAGCCAAGAAAGTATCAACTCTCAGAAAT
TTGGGGTCTTCAGAGACCAGAGATTGTCAAGCTTCTTCTGGGCTTTCTTTTGGGTATGCATGCCGGTGCAATACTTTCAGTCTTTCCTTACCTTCTAGGC
GAAGCCCTTACAATCTACTTTGAAGATAACAAATTCAAACTGAAGAGAGATGTTGGACGCCTCTGTTTAATACTTGTAGGTCTTGGATTTGGTTGCATTA
TTTCTATGACAGGCCAGCAAGGTTTATGCGGATGGGCAGGAACAAAACTTACCGTGAGAATACGGGACCTCTTATTTCGATCTATACTAAAGCAAGAACC
TGGTTGGTTTGATTTCGAAGAAAATTCCGTAGGAGTACTTGTTTCAAAGCTCTCAATTGATTGTATCAGTTTCCGATCCGTTCTTGGTGATAGACTCTCA
GTCTTGTTAATGGGTTTAAGTTCAGCTGCTGTTGGTCTTGGTCTGTCATTTTATCTGCAATGGCGATTGGCTCTTTTGGCTGCAGCACTAACTCCTTTCA
CTCTTGGTGCAAGCTACTTGAGTTTGATCATAAACGTTGGACCAAAACTAGATAATAGCTCTTATGCCAAAGCTAGCACAATTGCTGCTGGTGCTGTTTC
AAGTATAAGAACAGTAGCAACCTTCTCTGCTCAAGACCAAATAGTGGAGTCCTTTGATAGAGCCTTGGCTGAGCCCAAGAAGAAATCAGTGAAAAGGTCA
CAAGTTCTAGGCCTAACACTTGGATTCTCTCAAGGGGCCATGTATGGTGCATATACCTTAACACTTTGGTTTGGAGCCTACCTCGTAAAACAAGGGGAAA
CAAACATTGGCGTGGTATACAAAATTTTCCTCATTCTTGTGCTGAGTTCATTTTCTGTTGGACAACTAGCGGGGCTAGCGCCGGATACTTCCATGGCTGC
TCCTGCAATTGCTGCTATTTTTGATATCATTCATCGTAAGCCATTGATTCGTAGTGACCGGGATAGAGGTAAGAAAATTGATAGATCAAACCTATTGGAT
ATCGAGCTCAAGATGGTAACTTTTGCATATCCCTCTAGGCCTGAGATTATTGTGTTGAGGGATTTTTGCTTAAAGGTCAAAGGTGGTAGCACGGTGGCCT
TAGTTGGGGGTAGTGGGTCAGGAAAATCAACTGTTGTATGGTTGATACAAAGGTTTTATGATCCTAATCAAGGGAAGGTGACAATGGGAGGGGTGGATTT
GAGGGATTTTAATGTCAAGTGGTTAAGGAGTCAAACAGCTTTGGTCGGTCAAGAGCCTGCATTGTTTTCTGGTAGTATAAGGGAAAATATTGCATTTGGA
AATCCTAATGCTTCAAGGGCTGAGATTGAAGAGGCTGCAAGTGAAGCATACATTCACAAGTTCATCTGTAGCCTCCCTCAAGGCTATGAAACTCAGGTAG
GAGAGAGTGGGGTTCAACTATCTGGTGGCCAAAAACAAAGAATAGCAATAGCAAGGGCAATTCTAAAGAGATCAAGGGTGCTACTGCTAGATGAAGCAAG
TAGTGCACTGGACTTGGAATCAGAGAAGAATGTCCAAGAGGCACTAAGGAAGATCTCTAAGAGGGCAACGACTGTTATAGTGGCTCACAGGCTTTCTACC
ATTAGAGAAGCTGATATGATTGCTGTAGTGAAAGATGGTGCAGTGGTGGAGTATGGTAGTCATGATGCACTCTTGAATTCACATCGTAATGGTTTGTATG
CTAGCATGGTTCGTGCAGAAACTGAAACTAATGCATTTGCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G165000.1 pacid=42792516 polypeptide=Potri.001G165000.1.p locus=Potri.001G165000 ID=Potri.001G165000.1.v4.1 annot-version=v4.1
MADSSFEIDYHSTGRSRNYHQRRQLRHPSTPVNSQYDSRSFTQFSFTNTPNRLRQTPSTPFATDNDTSWQDELSWQFQPTGWNDTRSLGAALSPWAASTP
SNRHIFQRSANDYYLSRTHGGFRTFTNPYYDQSSYGAVPAGRLELQSYAARNNERSVVHVRDYSSAAYSKSHHGISRPISQAIKGGARRNASPLVDQDEL
SMIDYDSEDVEKQGELLQTDTNLHGDKDSRWISVSHAYMEDDGVSPLYHSTPHGGHDHHGHELSRSRHDDLLSAYEANRSTSRDYVPGKYPYDDIDQASE
YEDEDYDEEDDDNEEAARREVGLFSLFKYSTKWDMVLVFLGCLGALINGGSLPWYSYFFGDFVNRIAKHSDDNMMKEVERICLLMTGVAALVVVGAYLEI
TCWRLVGERSAHRIRNLYLSAVLRQDITFYDTKVSTSDIMHGISSDVAQIQEVMGEKMAHFIHHIFTFICGYWVGFLRSWKVSLVVLSVTPLTMFCGIAY
KAIYVGLATKEEVSYRKAGGVAEQAISSIRTVFSFVAEDKLARKYADLLMKSVPIGAKIGFAKGAGMGVIYLVTYSTWALAFWYGSILVARKEISGGDAI
ACFFGVNVGGRGLALSLSYFAQFAQGTVAATRVYEIIDRIPDIDPYSPHGRILSTVGGRIEIKGVTFAYPSRPETVILRSLNLVIPSAKTLALVGASGGG
KSTVFALIERFYDPINGVVTLDGNDLRTLQVKWLRGQIGMVGQEPVLFATSILENVMMGKENATKKEAINACIAANAHSFISGLPFGYDTQVGDRGTQLS
GGQKQRIALARAMIKNPRILLLDEPTSALDQESESVVQQAIDKISTGRTTIVIAHRLATVRNANTIAVLDQGSVVEIGDHRQLMENAGAYYDLVKLATEA
VSKSALKQEDAAKDMEFSIYEKSVDLRSKNAFETSKSRYLKSMQAENQQEEEMQESAKPRKYQLSEIWGLQRPEIVKLLLGFLLGMHAGAILSVFPYLLG
EALTIYFEDNKFKLKRDVGRLCLILVGLGFGCIISMTGQQGLCGWAGTKLTVRIRDLLFRSILKQEPGWFDFEENSVGVLVSKLSIDCISFRSVLGDRLS
VLLMGLSSAAVGLGLSFYLQWRLALLAAALTPFTLGASYLSLIINVGPKLDNSSYAKASTIAAGAVSSIRTVATFSAQDQIVESFDRALAEPKKKSVKRS
QVLGLTLGFSQGAMYGAYTLTLWFGAYLVKQGETNIGVVYKIFLILVLSSFSVGQLAGLAPDTSMAAPAIAAIFDIIHRKPLIRSDRDRGKKIDRSNLLD
IELKMVTFAYPSRPEIIVLRDFCLKVKGGSTVALVGGSGSGKSTVVWLIQRFYDPNQGKVTMGGVDLRDFNVKWLRSQTALVGQEPALFSGSIRENIAFG
NPNASRAEIEEAASEAYIHKFICSLPQGYETQVGESGVQLSGGQKQRIAIARAILKRSRVLLLDEASSALDLESEKNVQEALRKISKRATTVIVAHRLST
IREADMIAVVKDGAVVEYGSHDALLNSHRNGLYASMVRAETETNAFA
|
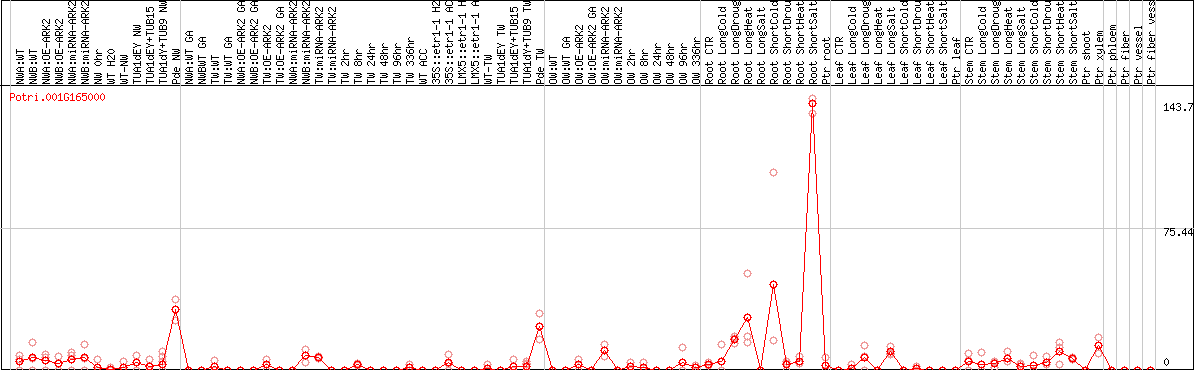
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G165000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
