External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G80160 270 / 4e-94
GLYI7
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
AT1G15380 263 / 2e-91
GLYI4
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
AT2G28420 171 / 9e-55
GLYI8
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
AT5G57040 54 / 2e-09
Lactoylglutathione lyase / glyoxalase I family protein (.1)
AT2G32090 51 / 1e-08
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G062400
287 / 1e-100
AT1G80160 261 / 2e-90
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.007G129200
228 / 1e-77
AT1G80160 225 / 1e-76
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.003G009000
194 / 2e-63
AT1G80160 190 / 8e-62
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.004G223300
190 / 6e-62
AT1G80160 187 / 1e-60
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.005G117000
187 / 6e-61
AT1G80160 166 / 2e-52
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.007G015100
185 / 5e-60
AT1G15380 167 / 7e-53
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.009G055500
165 / 2e-52
AT2G28420 229 / 1e-77
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Potri.010G087000
50 / 4e-08
AT2G32090 189 / 3e-63
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.018G056200
49 / 3e-07
AT5G57040 253 / 2e-86
Lactoylglutathione lyase / glyoxalase I family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030070
246 / 2e-84
AT1G80160 247 / 7e-85
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10014480
243 / 3e-83
AT1G80160 244 / 1e-83
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10025971
175 / 5e-56
AT1G15380 165 / 3e-52
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10014269
171 / 2e-54
AT1G15380 166 / 2e-52
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10005324
170 / 4e-54
AT2G28420 245 / 2e-83
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Lus10039579
169 / 2e-53
AT2G28420 248 / 2e-84
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Lus10017158
54 / 4e-09
AT5G57040 254 / 7e-87
Lactoylglutathione lyase / glyoxalase I family protein (.1)
Lus10006122
52 / 8e-09
AT2G32090 199 / 3e-67
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10010552
50 / 5e-08
AT2G32090 197 / 3e-66
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10021573
42 / 4e-05
AT5G57040 163 / 2e-51
Lactoylglutathione lyase / glyoxalase I family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0104
Glyoxalase
PF12681
Glyoxalase_2
Glyoxalase-like domain
Representative CDS sequence
>Potri.001G171600.1 pacid=42788944 polypeptide=Potri.001G171600.1.p locus=Potri.001G171600 ID=Potri.001G171600.1.v4.1 annot-version=v4.1
ATGAAGGACCACATGGGAAACTCACTCCATCTCAAGTCTTTGAACCACATCTCACTTTTGTGTAGATCAGTTGTGGAATCCATTGATTTCTACCAGGATG
TTCTTGGTTTTGTGCCAATAAGGAGGCCGGGATCTTTCAACTTTGATGGGGCATGGCTATTTGGTTTTGGAATTGGAATTCATTTATTGCAATCAGAAAA
CCCAGAAAAGATGCCAAAGAAAAGTGAAATTAATCCCAAGGACAATCATATCTCGTTTCAGTGTGAGAGCATGGGAGCAGTGGAGAAGAAGCTGAAAGAA
CTGGGAATTCAGCATGTGCGAGCATTAGTGGAGGAAGGTGGGATTCAGGTTGAACAGTTGTTCTTTCATGACCCTGATGGATTCATGATTGAGATTTGCA
ACTGCGATAATCTCCCAGTGATCCCTCTGGCGGGCGAAGTCGCACGATCATGCTCTTGCTTGAACCTTCAAACGATGCAGCAGGAGAGGCCAATGCTCCA
GCAGGGGAGAGCAATTTAG
AA sequence
>Potri.001G171600.1 pacid=42788944 polypeptide=Potri.001G171600.1.p locus=Potri.001G171600 ID=Potri.001G171600.1.v4.1 annot-version=v4.1
MKDHMGNSLHLKSLNHISLLCRSVVESIDFYQDVLGFVPIRRPGSFNFDGAWLFGFGIGIHLLQSENPEKMPKKSEINPKDNHISFQCESMGAVEKKLKE
LGIQHVRALVEEGGIQVEQLFFHDPDGFMIEICNCDNLPVIPLAGEVARSCSCLNLQTMQQERPMLQQGRAI
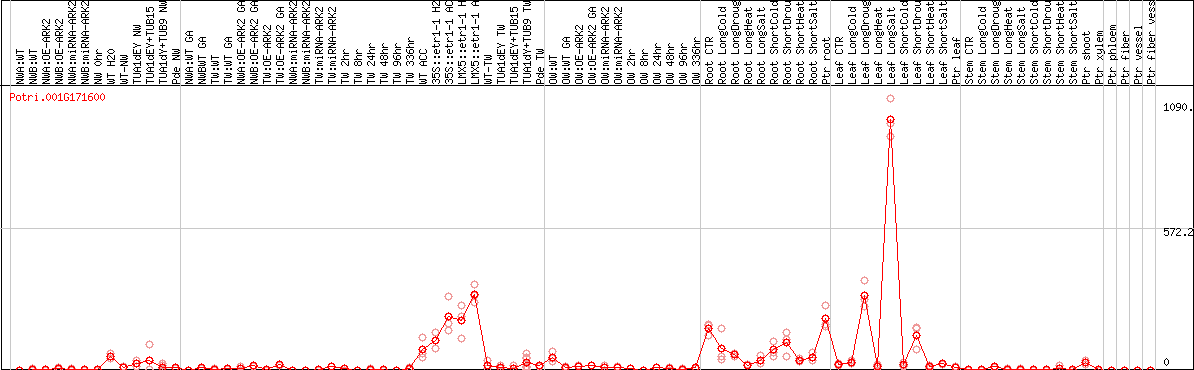
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G171600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.