PtrPDR12 (Potri.001G175700) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PtrPDR12 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G175700.1 pacid=42791332 polypeptide=Potri.001G175700.1.p locus=Potri.001G175700 ID=Potri.001G175700.1.v4.1 annot-version=v4.1
ATGGAAAGTGGTTATCTTTACAGAGCCGGTAGCAGTGTAAGAAGAGGAAACTCTTCAGGAACATTTAGTAACAATGCAGCTGCAGATCATCAAGTTTTTT
CACTTTCTTCTCACGGACAAGACGACGATGAAGAGGCTTTAAAATGGGCGGCATTAGAGAAACTACCAACCTATGATCGTTTGAGGAAAGGTATATTAAC
CACGTCCACAGGTGCTGCTAGTGAAGTGGAGGTGCAGAATCTTGGGTTCCAAGAAAGGAAGAATTTGGTGGAGAGGTTGGTCAATGTAGCAGAAGAGGAC
AATGAGAAGTTCTTGTTAAAGCTCAAGAACCGTATTGATAGAGTTGGAATTCATGTTCCCACAATTGAAGTTCGATTCGAGCATCTAAATGTTGAAGCAG
AAGCCTATGTAGGAAGTAGAGCTTTGCCTACATTCTTTAACTACTCTGTTAATATGCTGGAGGGTGTCTTGAACTATCTCCATATTCTTTCTAGTAGAAA
GAAACATATGTGGATCCTTAAGGATGTTAGCGGTATCATCAAGCCATCGAGAATGACATTGCTTTTGGGCCCTCCAAGTTCCGGAAAGACAACGCTTTTG
TTAGCTTTGGCTGGAAAGCTTGATCATGCTCTAAAGTTTTCAGGGAGGGTGACTTATAATGGTCATGAAATGGATGAGTTTGTACCGCAAAGAACTGCTG
CCTATATCAGTCAACACGATCTGCATATAGGAGAAATGACTGTAAGGGAAACCTTGGCCTTTTCTGCAAGATGCCAAGGGGTTGGAAGCCGTTATGATAT
GTTAGCAGAATTATCGAGAAGGGAGAAAGAAGCGGGGATCAAACCTGATCCTGATATTGATGTCTTCATGAAGGCTGCAGCGACAGAAGGTCAGGAGGAC
AGTGTGGTGATAGATTACATTCTGAAGGTTTTAGGACTGGAAGTCTGCGCGGATACATTGGTCGGGGATGAAATGCTGCGGGGTATTTCTGGAGGACAAA
AGAAGCGTGTTACCACAGGTGAGATGCTGGTTGGACCAGCAAAGGCACTGTTTATGGATGAGATATCTACTGGATTGGATAGCTCAACAACGTACCAGAT
TGTGAACTCCATCAAGCAATATGTCCAAATTCTTGAGGGAACTGCTCTCATCTCTCTCCTCCAGCCAGCGCCAGAGACTTATGATCTCTTTGATGACATT
ATTCTTCTCTCAGATGGCGAGATAGTGTATCAGGGACCTCGTGAGCATGTCCTCAGGTTTTTTGAATATATGGGCTTCAAGTGTCCTGCAAGGAAAGGTG
TGGCAGACTTCTTGCAAGAAGTGACATCAAGGAAAGATCAGATGCAGTATTGGGCACGACGAGATGTGCCATACAGATTTGTCACAGTCAAAGAATTTGC
CGAGGCATTCTATTCGTTTCATGAAGGAAAGAGACTTGGAAATGAGCTTGCAGTTCCATTTGATAAGAGCAAGAACCACCCAGCTGCTTTGACTACCAAG
AAATACGGAGTTAATAAGAGAGAGCTATGCAAAGCTTCTTTCTCAAGAGAATTCTTACTCATGAAAAGGAACTCATTCGTCTACGCCTTCAAGTTCATTC
AACTGACAATAGTCGCAGTGATTGCGATGACACTCTTTCTACGGACTGAAATGCACCGAGACTCGGTAACCGATGGAGGAATTTATGTGGGCGCTATGTT
CTTCATTGTGGTTGTGATCATGTTTAATGGTATGGCTGAGATTTCCATGACCCTTGCAAAGCTTCCTGTGTTCTACAAGCAAAGGGACCTTCTATTCTTT
CCTGCTTGGATATATGCTCTTCCTACATGGATCCTTAAAATCCCTATCACTTTCATCGAAGTTGCTATTATGGTTTTCATTACGTATTTTGTCATTGGAT
TTGATCCAAATGTAGGAAGGTTGTTTAAGCATTACCTCGTGCTCTTATTAACAAATCAGATGGCTTCTGGATTATTCCGAACCATTGCTGCTGTTGGCAG
GAACATGGTTGTTGCTAACACCTTCGGGTCATTTGTACTTCTCTTGCTGTTTGTTTTGGGTGGCTTTGTCTTGTCACGAGATGACATAAAGAAATGGTGG
ATTTGGGGCTTCTGGACATCACCCATGATGTATGCGCAGAATGCAGTTGTCGTGAACGAGTTTCTTGGAAAGAGTTGGAACCATGTTCTTCCAAACTCAA
CGGAGCCATTGGGAATTGAAGTTCTGAAGTCTCGTGGGTTCTTCACAGAAGCATATTGGTATTGGTTAGCAGTAGCAGCATTGTTTGGATTCACTCTACT
ATACAACTTCTTGTACATTTTGGCTCTCGCTTTTCTCAACCCATTAGGAAAGCCACAACAGGCTGGTATATCAGAAGAACCTCAAAGCAACAACGTTGGT
AGAATAGGAGAAGCCATTCATTTAATGAACCCTGGAATTAACTCTAGCCTCCACACAAGTGCAGAGAGCATAGATGAAATTGGAAGAAGCAAGTCATCAA
GGTTCACATGTAACAAACAAAGGGGGGTCATTATTCCATTCGAACCACATTCTATAACGTTTGATAAAGTTATGTACTCAGTTGACATGCCACAGGAAAT
GAAAAGTCATGGTGTTCATGAAGATAAATTGGTACTGTTGAAGGGTGTGAGTGGCGCATTTAGGCCAGGTGTTCTCACAGCTTTAATGGGCATCAGTGGT
GCTGGTAAAACCACAATGATGGATGTTCTTGCTGGTAGGAAAACTGGTGGATATATTGAGGGGAACATCACAATTTCTGGGTATCCGAAGAAGCAAGAAA
CATTTGCTAGAATATCTGGATACTGTGAGCAAAATGACATCCACTCACCACATATTACTGTCTATGAGTCCTTGCTCTACTCAGCTTGGCTTCGTCTTCC
CACTGAAGTAGACATTGAAACTAGAAAAATGTTCGTTGAGGAGGTCATGGAGCTTGTAGAACTGAATCCATTGAGGCAAGCTTTAGTTGGATTGCCTGGT
GTAGATGGTCTGTCAACTGAGCAACGGAAGCGACTAACCATTGCAGTTGAGCTAGTGGCAAACCCCTCAATTATTTTTATGGATGAACCAACTTCAGGGC
TTGATGCAAGAGCAGCTGCTATTGTTATGAGAACAGTTAGGAACACAGTGGACACAGGAAGAACAGTTGTCTGCACCATCCATCAACCAAGCATTGACAT
ATTTGAAGCTTTTGATGAGTTGTTTCTACTGAAGAGAGGAGGACAAGAGATATATGTGGGGCCATTGGGTCGACTTTCTTGCCATCTGATCAAGTATTTT
GAGGGAATTGAAGGAGTCAACAAAATCAAAGATGGTTACAATCCGGCTACTTGGATGTTGGAAGTGACCAGTACAGCAGAAGAATTGGCTCTGGGGGTTG
ATTTTGCTGAAATATATAGAAGTTCAGAATTGTTTAGGAGAAACAGAGCTCTCATAAAGGATTTGAGCACTCCTGCTCCTGGTTCGAAGGACCTCTATTT
CTCTACTCAATATTCGCGATCATTTTTCACCCAGTGTTTGGCTTGCTTATGGAAACAGCACTGGTCGTACTGGCGCAACCCTCCATACACTGCTATCAGA
TTTCTCTCTACAACAGTCATAGGCTTGATCTTTGGAACAATGTTTTGGGATATTGGCTCCAAAATAACAAAGCGACAAGATCTTTTTAATGCAATGGGTT
CAATGTATACCGCTGTTCTCTTTCTTGGTGTCCAAAATGCCGCGTCTGTTCAGCCTGTGGTAGCAGTTGAGCGAACTGTCTTTTACAGAGAGAGAGCTGC
TGGAATGTACTCTGCCTTGCCGTATGCCTTTGCACAGGTTCTCATTGAGCTTCCATATATTTTTGTCCAAGCTGCAGTTTATGGTGTTATAGTCTATTCA
ATGATTGGATTCGGATGGACCATCTCAAAGTTCTTCTGGTATCTGTATTTCATGTACTTCACGCTACTGTATTTCACCTTCTATGGCATGATGGCCGTGG
CCGTGTCACCAAATCACCAAATTGCGTCTGTAATTTCTGCTGCATTTTATGGAATCTGGAATGTATTTTCAGGATTCGTAATCCCACGATCTCGGATGCC
TTTGTGGTGGAGATGGTACTCTTGGATATGTCCAGTCTTTTGGACCCTGTATGGACTGGTTGCCTCCCAGTTTGGGGATATGAAGGATAGACTTGAAACT
GGTGAAACAGTGGAGCAATTCGTGACAATTTATTTGGATTTCAAACATGATTTTCTTGGAGTTGTTGCAGCTGTGATTCTTGGGTTTACAGTCCTTTTTG
CAATAACTTTCGCCATTTCCATCAAGCTCTTCAATTTCCAAAGGCGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G175700.1 pacid=42791332 polypeptide=Potri.001G175700.1.p locus=Potri.001G175700 ID=Potri.001G175700.1.v4.1 annot-version=v4.1
MESGYLYRAGSSVRRGNSSGTFSNNAAADHQVFSLSSHGQDDDEEALKWAALEKLPTYDRLRKGILTTSTGAASEVEVQNLGFQERKNLVERLVNVAEED
NEKFLLKLKNRIDRVGIHVPTIEVRFEHLNVEAEAYVGSRALPTFFNYSVNMLEGVLNYLHILSSRKKHMWILKDVSGIIKPSRMTLLLGPPSSGKTTLL
LALAGKLDHALKFSGRVTYNGHEMDEFVPQRTAAYISQHDLHIGEMTVRETLAFSARCQGVGSRYDMLAELSRREKEAGIKPDPDIDVFMKAAATEGQED
SVVIDYILKVLGLEVCADTLVGDEMLRGISGGQKKRVTTGEMLVGPAKALFMDEISTGLDSSTTYQIVNSIKQYVQILEGTALISLLQPAPETYDLFDDI
ILLSDGEIVYQGPREHVLRFFEYMGFKCPARKGVADFLQEVTSRKDQMQYWARRDVPYRFVTVKEFAEAFYSFHEGKRLGNELAVPFDKSKNHPAALTTK
KYGVNKRELCKASFSREFLLMKRNSFVYAFKFIQLTIVAVIAMTLFLRTEMHRDSVTDGGIYVGAMFFIVVVIMFNGMAEISMTLAKLPVFYKQRDLLFF
PAWIYALPTWILKIPITFIEVAIMVFITYFVIGFDPNVGRLFKHYLVLLLTNQMASGLFRTIAAVGRNMVVANTFGSFVLLLLFVLGGFVLSRDDIKKWW
IWGFWTSPMMYAQNAVVVNEFLGKSWNHVLPNSTEPLGIEVLKSRGFFTEAYWYWLAVAALFGFTLLYNFLYILALAFLNPLGKPQQAGISEEPQSNNVG
RIGEAIHLMNPGINSSLHTSAESIDEIGRSKSSRFTCNKQRGVIIPFEPHSITFDKVMYSVDMPQEMKSHGVHEDKLVLLKGVSGAFRPGVLTALMGISG
AGKTTMMDVLAGRKTGGYIEGNITISGYPKKQETFARISGYCEQNDIHSPHITVYESLLYSAWLRLPTEVDIETRKMFVEEVMELVELNPLRQALVGLPG
VDGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELFLLKRGGQEIYVGPLGRLSCHLIKYF
EGIEGVNKIKDGYNPATWMLEVTSTAEELALGVDFAEIYRSSELFRRNRALIKDLSTPAPGSKDLYFSTQYSRSFFTQCLACLWKQHWSYWRNPPYTAIR
FLSTTVIGLIFGTMFWDIGSKITKRQDLFNAMGSMYTAVLFLGVQNAASVQPVVAVERTVFYRERAAGMYSALPYAFAQVLIELPYIFVQAAVYGVIVYS
MIGFGWTISKFFWYLYFMYFTLLYFTFYGMMAVAVSPNHQIASVISAAFYGIWNVFSGFVIPRSRMPLWWRWYSWICPVFWTLYGLVASQFGDMKDRLET
GETVEQFVTIYLDFKHDFLGVVAAVILGFTVLFAITFAISIKLFNFQRR
|
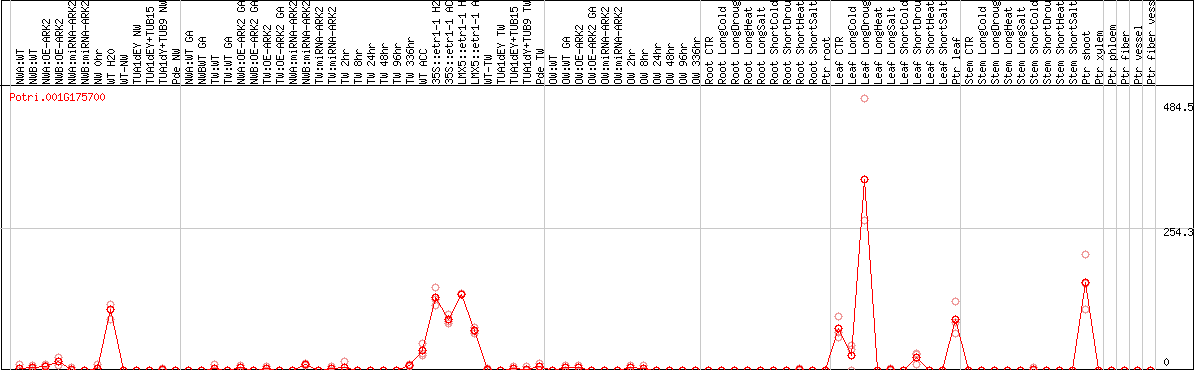
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G175700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
