External link
Symbol
2OGox10
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G52790 461 / 1e-164
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52800 346 / 2e-119
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52820 308 / 1e-104
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G80320 285 / 2e-95
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G15540 262 / 2e-86
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G03070 261 / 3e-86
AOP1.1, AOP1
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G28030 217 / 9e-69
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52810 204 / 2e-64
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G03050 129 / 2e-35
AOP3
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT1G14120 111 / 2e-28
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G176500
377 / 7e-132
AT1G52800 338 / 1e-116
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176200
329 / 9e-113
AT1G52820 496 / 8e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176100
308 / 2e-104
AT1G52800 286 / 5e-96
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.006G248000
266 / 1e-87
AT1G52820 280 / 6e-93
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.018G033400
263 / 6e-87
AT1G52820 269 / 4e-89
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G175800
262 / 1e-86
AT1G52820 338 / 2e-116
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G175900
261 / 3e-86
AT1G52820 328 / 2e-112
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.002G159500
134 / 5e-37
AT4G23340 402 / 3e-141
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.003G128100
119 / 5e-31
AT4G23340 458 / 2e-163
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041281
322 / 3e-111
AT1G52790 298 / 5e-102
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10016659
303 / 1e-102
AT1G52820 445 / 1e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10023024
269 / 5e-89
AT1G52820 371 / 2e-129
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005773
220 / 7e-70
AT1G52790 205 / 3e-64
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10021025
216 / 2e-68
AT1G52800 226 / 2e-72
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005776
213 / 3e-67
AT1G52790 198 / 2e-61
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10008097
201 / 1e-62
AT1G52800 221 / 2e-70
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005516
202 / 2e-62
AT1G52820 278 / 7e-92
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10013132
195 / 3e-60
AT1G52800 222 / 8e-71
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10023160
185 / 2e-57
AT1G52820 244 / 1e-80
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
CL0029
Cupin
PF14226
DIOX_N
non-haem dioxygenase in morphine synthesis N-terminal
Representative CDS sequence
>Potri.001G176000.1 pacid=42793183 polypeptide=Potri.001G176000.1.p locus=Potri.001G176000 ID=Potri.001G176000.1.v4.1 annot-version=v4.1
ATGGGTTCCAATGAACTGCCATCAAAGCTTCCTATTCTCGACTTCGCCAAGGAAGATTTGAAGCCTGGCACAAGTTCTTGGCTCAAAGCTTGCAGTGATG
TTAGGCTAGCGCTTGAAGAGTATGGCTGTTTTGTAGTAGAGTACAACAGACTACCTCTAGAACTTAGAGATAAAGTCTTTGGTGTTTTAAAGGAGTTGTT
CGATCTCCCGACTGAAACAAAAATGCAGAACAAATATGAGAAGCCCTTGAATGGTTATGTTGGACAGATTGCCAAACTCCCTCTCCATGAAAGCATGGGA
ATTGACAATGCAACATCTCTAGACGCAACTCAAAATTTCACAGATCTCATGTGGCCTAATGGGAATGATCGCTTCTGTGAATGTGTTCTCAAGTACGCTA
ATCTAGCTGCAGAGCTAGACCGAATGGTGACCAGAATGGTATTTGAAAGTTATGGAGTAGAAAAGTACCATGACTCTTATGTTGAGTCTACTACTTACCT
TCTTCGGCTTTTGAAAAATAGACCTCCCAAGGTGGACGAGCACAATTTAGGTTTTGTCACTCACACTGACAAGAGCTTCACAACCATACTTCATCAAAAT
GAAATTAATGGGTTGGAAGTGGACACCAAGGATGGCAAGAAGATCAATGTTGAGCTTTCACCTTCGTCTTTTGTTGTCATAGCTGGTGATGCATTAATGG
CATGGAGCAATGACAGAGTAATATCTCCTAGCCACCGTGTTATCATGAACGGAAAAATAGACAGATACTCCCTGGCGCTGTTTGCTTTCAACAAAGGAAT
ATTACAAGTTCCTGAAGAACTTGTTGACGAGGAGCACCCTTTAATGTACAAGCCACTGGATCATATTGGACTCCTTCATTTCTATCGGACAGAAGAGGGA
TACAAAAGCAAATGTCCTATCAAGGCTTATTGTGGCGTTTGA
AA sequence
>Potri.001G176000.1 pacid=42793183 polypeptide=Potri.001G176000.1.p locus=Potri.001G176000 ID=Potri.001G176000.1.v4.1 annot-version=v4.1
MGSNELPSKLPILDFAKEDLKPGTSSWLKACSDVRLALEEYGCFVVEYNRLPLELRDKVFGVLKELFDLPTETKMQNKYEKPLNGYVGQIAKLPLHESMG
IDNATSLDATQNFTDLMWPNGNDRFCECVLKYANLAAELDRMVTRMVFESYGVEKYHDSYVESTTYLLRLLKNRPPKVDEHNLGFVTHTDKSFTTILHQN
EINGLEVDTKDGKKINVELSPSSFVVIAGDALMAWSNDRVISPSHRVIMNGKIDRYSLALFAFNKGILQVPEELVDEEHPLMYKPLDHIGLLHFYRTEEG
YKSKCPIKAYCGV
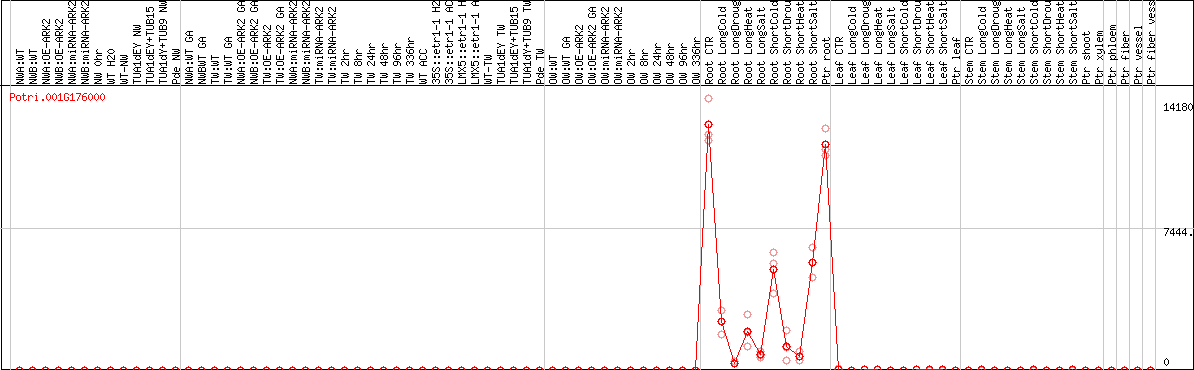
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G176000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.