External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G79900 425 / 5e-151
ATMBAC2, BAC2
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
AT5G46800 145 / 2e-41
BOU
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
AT2G33820 127 / 3e-34
ATMBAC1
Mitochondrial substrate carrier family protein (.1)
AT5G01340 116 / 4e-30
AtmSFC1
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
AT1G34065 101 / 2e-24
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT4G01100 92 / 6e-21
ADNT1
adenine nucleotide transporter 1 (.1.2)
AT4G39460 90 / 3e-20
SAMC1, SAMT1
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
AT2G30160 89 / 5e-20
Mitochondrial substrate carrier family protein (.1)
AT5G19760 88 / 7e-20
Mitochondrial substrate carrier family protein (.1)
AT1G07030 88 / 7e-20
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G053900
551 / 0
AT1G79900 419 / 2e-148
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Potri.001G142600
154 / 1e-44
AT5G46800 434 / 1e-154
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Potri.003G091500
149 / 1e-42
AT5G46800 443 / 4e-158
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Potri.004G046801
131 / 5e-36
AT2G33820 453 / 4e-162
Mitochondrial substrate carrier family protein (.1)
Potri.011G056000
129 / 5e-35
AT2G33820 460 / 9e-165
Mitochondrial substrate carrier family protein (.1)
Potri.016G119500
116 / 3e-30
AT5G01340 542 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Potri.012G110700
106 / 1e-25
AT5G51050 714 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Potri.005G197200
95 / 2e-22
AT4G39460 417 / 3e-147
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
Potri.006G252100
96 / 3e-22
AT4G32400 493 / 4e-175
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037544
460 / 1e-164
AT1G79900 408 / 2e-144
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Lus10011451
457 / 1e-163
AT1G79900 410 / 4e-145
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Lus10000906
157 / 1e-45
AT5G46800 482 / 1e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Lus10004565
155 / 5e-45
AT5G46800 481 / 2e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Lus10027817
110 / 8e-28
AT5G01340 526 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Lus10015420
106 / 1e-26
AT2G33820 415 / 7e-147
Mitochondrial substrate carrier family protein (.1)
Lus10005045
102 / 5e-25
AT5G01340 518 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Lus10043022
100 / 3e-23
AT5G51050 758 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Lus10032522
99 / 3e-23
AT5G51050 563 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Lus10005787
96 / 1e-22
AT4G39460 494 / 2e-177
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.001G182000.1 pacid=42788429 polypeptide=Potri.001G182000.1.p locus=Potri.001G182000 ID=Potri.001G182000.1.v4.1 annot-version=v4.1
ATGGATTTTTGGCCAGAGTTTCTTGCTAGCAGCTGGGGTAGGGAGTTTGTGGCTGGTGGATTTGGTGGAATTGCTGGTATAATTTCTGGTTATCCACTTG
ATACTCTACGTATTCGGCTACAGCAACCTAATTCTGGCTCCGCCTTCAGTATTCTTCGCCGAGTTATGGCCGGAGAAGGGCCTGCTGCTCTCTACAGAGG
CATGGGTGCACCCCTCGCATCCGTCACCTTCCAGAATGCCATGGTTTTCCAGACCTATGCTATCCTCTCGAGAGCATTTGACTCGTCCGTTTCAGCTAGT
GATCCTCCTTCTTACAAAGGTGTTGTTCTAGGAGGAGTTGGTACTGGGGCCATACAGAGCATTATGCTTTCTCCTGTAGAACTGGTAAAAATTCGCCTCC
AATTGCAGAACGTAAGTCATGCAAATCTCCATGGGGCTGCTAGTTATAAAGGTCCAGTAAGTGTTGCCAAGAGCATACTGAAAACAGAAGGTATAAAAGG
AATTTACCGGGGTTTTGTGATCACTGTCCTCAGAGATGCACCTGCTCATGGTGTATACTTTTGGACATATGAGTACATGAGAGAGCAGTTTCACCCTGGC
TGCAGAAAGAATGGTCACGAAAGCTTAAGAACCATGCTGACAGCAGGAGGGCTAGCAGGAGTTGCAAGCTGGCTCTGCTGCTATCCATTAGATGTTGTGA
AAACCAGATTACAAGCTCAAACACCATCATCATCATCCCCACTTAAGTATAAAGGCATTCTGGATTGCTTTCGCAGGAGTGTTAAAGAAGAAGGTTACTG
CGTGCTTTGGAGAGGTTTGGGGACTGCCGTAGCAAGAGCTTTTGTAGTCAATGGAGCAGTATTTGCTGCTTATGAGATTGCATTGCGGTGTCTATTCAAC
AATGGAAGCATTCAAACAGAGAACACTATCTAG
AA sequence
>Potri.001G182000.1 pacid=42788429 polypeptide=Potri.001G182000.1.p locus=Potri.001G182000 ID=Potri.001G182000.1.v4.1 annot-version=v4.1
MDFWPEFLASSWGREFVAGGFGGIAGIISGYPLDTLRIRLQQPNSGSAFSILRRVMAGEGPAALYRGMGAPLASVTFQNAMVFQTYAILSRAFDSSVSAS
DPPSYKGVVLGGVGTGAIQSIMLSPVELVKIRLQLQNVSHANLHGAASYKGPVSVAKSILKTEGIKGIYRGFVITVLRDAPAHGVYFWTYEYMREQFHPG
CRKNGHESLRTMLTAGGLAGVASWLCCYPLDVVKTRLQAQTPSSSSPLKYKGILDCFRRSVKEEGYCVLWRGLGTAVARAFVVNGAVFAAYEIALRCLFN
NGSIQTENTI
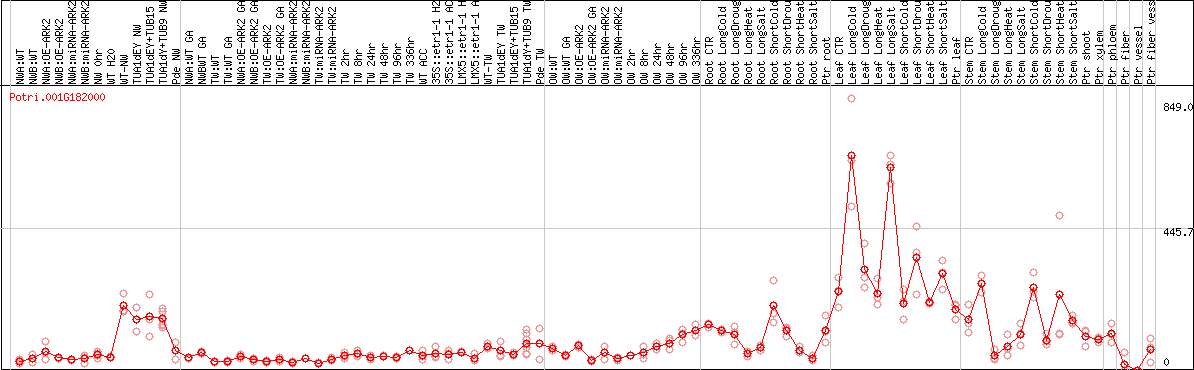
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G182000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.