Potri.001G185950 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
No hit found |
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G185950.1 pacid=42789577 polypeptide=Potri.001G185950.1.p locus=Potri.001G185950 ID=Potri.001G185950.1.v4.1 annot-version=v4.1
ATGATTCCTGACCAATATAATATCATCGGTCAATCTTCTTATCAAAAAAAAAGAAAACAGCGTTTGGCACGTTCATCAAAGAAAAACCCCTCCTTTACAT
CATGTTGTGCTAATGGCAAAATCCAATTGCCTGCTGCTCCTAACACACCTCAATTTCTAGATGATCTTTTAAATCCTGACAAAGGTTCATTGTCCATCAA
ATTCCGGCATAACATTCGTGCTTATAATTCAATGTTTGCTTTCACTTCAATGGGAGCGCAGATTGATCATACTGTCAATTCTCAACCAGGGCCATATATA
TTTAAAATAAATGGACAATGTCATCATCTTATGGGATCATTAGTACCAATTGATGCTGAATCACCAAGATTTGCACAACTATACATCTTTGACACAGACA
ATGAAATTGCTAATCGACTACACCCTTTTAATAATGATAATTGTCAATCATCTTTAGATGAAAATGTTGTCAACAAACTTATTGACATGCTTGATTCTTC
CAATGCATTGGTTAAATTGTTTCGTCAAGTTAGACATAGACTTAACAATGATGAATTCCCGAATTTCAAACTTCGTTTAATTGGGAAAAGAGATGGTGAC
TCAAAGCAATATGATGATCCTTCCTCTAATGATGTATGTGGTTTAATTGTTGGCGATATTGGAGAATCACAAACTGATAGAGATATTATTATTGAAGGTT
ATTCTAGAAATCTCCGCAGGATCTCTAAATTGCATCCAAAATTTATGTCTCTTCAGTATCCACTCTTATTTCCTTATGGTGAAGATGGCTATCATACTGA
CATTTTATTCACAAATCAAGAGCATTATACTCCTTCAAAACGTCAAAAAGTTACAATGCGAGCATATTATGCTTATGTCATTCAAGAAAGATTAGGAGAT
AGTAGTACACTTACCAAAGGCGGAAGACTTTATCAACAATTTTTAGTTGATGCATTTATGAATGTTGAACAAGAACGCCTTGATTTCATTCGCTCAAACC
AAGAAAATCTGAGAACTGAATCTTATAAAGGTGTCCAAGACGCAGTTTTAAGAGGTGATGTTAATGGTTCAAGTACTGGAAAAATCATCCTCCCCTCTTC
TTTAACTGGAAGCCCTAGATACATGATTAACAATTATCATGATGCAATGGCTATATGTCGTCATTATGGTAATCCTGACCTTTTTATTACATTCACTTGC
AATGTCAATTGGCCTGAAATTCAAAGAGAAATTAAAAAAAGCCGAAATTATAAAGCTGAAGACAAACCTGACATAATTGGAAGAGTATTTCGATACAAAC
TTAATGATATGATTTCATTCATTAAATCAGGACAGCCCTTCGGGAAAACAATTGCTGATGTTTGTGCTATTGAATTCCAAAAAAGAGGCTTGCCGCACAC
TCATTTATTGATTTGGCTAGCTTCTGAATACAAATTTCGATCACCTCAGGATGTCGATTCAGTTATTTCGGCTGAACTGCCAAACAAAGCAGATGATCCT
CATTGCTATGCAATTGTTTCAAAATTTATGCTGCATGGTCCATGCGGAATTGCTAGCCCAAAAGCCCAATGTATGAAGGGAAATCAATGTTCAAAAAAAT
TCCCTAAAAAGTTTAAGCAATCAACGGTTTTCGGTGAAAATGGCTTTGTGTTTTACAAGCGACGAAATTTCCCTGCGTCATTTGTTATGAAAAATGGAAT
TGCTCTCTCTAATTCCTATGTTGTCCCATACAACAAAGAGCTTCTAATCCGTTATAATGCTCATGTTAATGTTGAAATTTGTTGCCAATCAATGCTCATA
AAATATCTTTTCAAATATGTTAGCAAAGGATCTGATAGGTGTCGAGCTGTTATTCAAGGTCAAACAAACGATGAGATTCAGGCTTATCTTAATTGTAGAT
TTGTTTGTCCTTATGAAGCTGTATGGCGACTCTTACAATTTCCAATCCATTCAAGAAACCCAGCAGTTGAAAGACTTCAAATACATTTGCCAATGCAACA
TTCTGTTGTTTTCTTTGGAAATCAAAACTTATCTTCTGTTTTAAGAAAAAATGGTCTCAATAAGACAATGCTCACAGGATGGTTTGATCAGAACAAAGAA
GATGTTGAAGCAACACAGTTATATTACTCGCAATTTCCAAACAAATATGTTTGGGATGCTAGGCAGAAAGAATGGATTTACAGAACAAGAGGTTTTTCAC
TTGGACGTATAACATATGTGCATCCTGCTGCTGGCGAATTGTATTTCTTAAAGATGCTTTTAAATCATGTTAAAGGAGCTACAAGCTTTGAATATCTACG
TTGTGTCTCTGGTATTGTTTATCCAACCTTTCAACTCGCCTGCAAAGCACTTGGTCTCCTTGATGATGGCAAAGAATGGGCTGAAGCTTTTTCAGAAGCT
GTTTTAACTGCTTCCTCCTCACAACTTAGGCAACTATTTGTCAGTGTCACTCTATTTTGTCAAATTGCTAACCCCCAAGATTTGCTTGATCAATTTTGGC
ACACAATGCATGACGACATTAGAATCAAGCTATCATCTTTCTCCCCACATAATCTTCATTTCAGTGATAATGAACTTAAAAATTATGTTCTTTATGAACT
TGAACAACTTTTCAATGCCTTGGCAACATCCCTCAAAGACTATAATTTACCACTGCCAAATGATCGTTTGATGTCTGAAATTAGAAATAATCTCCTTAGA
GAAGAACTCAATTATGATATTTCTGAACTTCGAAGCAATAATGAAGCATCAATTTCATTACTTAACACATGCCAAAAGAAAATTTATGATCGAGTTATGG
AATCAATTTCAAAAAATCAACAATCTCTTATTTTTGTTTACGGGCATGGTGGCACAGGGAAAACTTTTTTATGGCATTCACTTATTAATAGTATTAGATC
AGAAGGGTTAATTGTTCTTGCTGTTGCATCATCAGGAATCGCATCTATTCTGCTCCCTGGTGGTCGCACAGCTCACTCAAGGTTTAAGATTCCTCTTGCT
ATAAATGAGAACTCAACATGTGAAATAAAAAAAAATACTCATCTTTCAAGGTTGATTGAAACGACAACACTTATTGTATGGGATGAAGCTCCTATGAATA
ATCGCTACTGTTTTGAAACATTGGATAGATCATTACGTGATATAATGGGCCAAACTGGTCATTCCAATCATAACCAACCTTTCGGTGGGAAATCTATTTT
ACTTGGTGGGGATTATCGTCAAATTCTCCCTGTTATACCTGGAGGAACAAAAGAAGATATTATTAATGCTTCCTTAAGTAGTTCACCACTCTGGCCAAAA
TTTGAAATCATGTTGCTAAAACAAAATATGAGATTGTCAATTGCTGGTCTTGGTTCAGATGAAATCAATGAAATTAAAACATTTGCTAAATGGATTCTTA
AAATAGGTGACGGAGACCTTTGTGATATACCTTTTTTTGATGAACTTGATGAATCTTTGATTAAAATTCCATGTGATTTGCAACTACATACCTCTGGTGA
TCCAATTAAAGCTATGGTTTCAGCTATTTATCCCAGCATTGAACAACCTGCTCTTGAACCATTTTACTTTAAAGAAAGAGCTATTATCACACCAAAAAAC
ATTACTGTTACTGAAATCAATAATTTCATACTTGGAGTTACACATGGTCCTCAACGCATTTATCTTAGCAATGATTCTGTTGATGCATCTTCCAGTGATA
ACGACAATATAAACTTATTATATCCATTGGAATTTATAAACCAACTTGAGTTCAGTGGTGTTCCTTCGCACATTCTTGCTTTGAAAATAGGAGCGCCAAT
TATGTTGCTGAGGAATCTGAGTCCAATGATAGGCTTATGTAATGGAACACGACTAATCATCACACAACTTGCTGATAGAGTTATTGAAGCTCAAATTATA
ACAGGCTCACACATTGGTGACCGAGTTTTTATACCAAGGATCATCTTCCCAATTAATGATGACAAATGCCCATTTACAATCAAACGAAGACAATTTCCAA
TACGATTATGCTATGCTATGACAATCAATAAAAGTCAAGGACAATCATTAAAATTTGTTGGTGTTTTCTTAAAGGAGCAAGTTTTTGCACATGGCCAACT
ATATGTCGCTCTTTCAAGAGTTACATCTAAAAAAGGTTTAAAAATCATTTCATGTGACCAAGAAGGAAAACAGCTTGATTATGCTAAAAATATTGTCTAC
AAAGATGTCCTCAATTTATTGCCCAAAGGTCTTTTTCCCATTCAAAATGATTTCTAA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G185950.1 pacid=42789577 polypeptide=Potri.001G185950.1.p locus=Potri.001G185950 ID=Potri.001G185950.1.v4.1 annot-version=v4.1
MIPDQYNIIGQSSYQKKRKQRLARSSKKNPSFTSCCANGKIQLPAAPNTPQFLDDLLNPDKGSLSIKFRHNIRAYNSMFAFTSMGAQIDHTVNSQPGPYI
FKINGQCHHLMGSLVPIDAESPRFAQLYIFDTDNEIANRLHPFNNDNCQSSLDENVVNKLIDMLDSSNALVKLFRQVRHRLNNDEFPNFKLRLIGKRDGD
SKQYDDPSSNDVCGLIVGDIGESQTDRDIIIEGYSRNLRRISKLHPKFMSLQYPLLFPYGEDGYHTDILFTNQEHYTPSKRQKVTMRAYYAYVIQERLGD
SSTLTKGGRLYQQFLVDAFMNVEQERLDFIRSNQENLRTESYKGVQDAVLRGDVNGSSTGKIILPSSLTGSPRYMINNYHDAMAICRHYGNPDLFITFTC
NVNWPEIQREIKKSRNYKAEDKPDIIGRVFRYKLNDMISFIKSGQPFGKTIADVCAIEFQKRGLPHTHLLIWLASEYKFRSPQDVDSVISAELPNKADDP
HCYAIVSKFMLHGPCGIASPKAQCMKGNQCSKKFPKKFKQSTVFGENGFVFYKRRNFPASFVMKNGIALSNSYVVPYNKELLIRYNAHVNVEICCQSMLI
KYLFKYVSKGSDRCRAVIQGQTNDEIQAYLNCRFVCPYEAVWRLLQFPIHSRNPAVERLQIHLPMQHSVVFFGNQNLSSVLRKNGLNKTMLTGWFDQNKE
DVEATQLYYSQFPNKYVWDARQKEWIYRTRGFSLGRITYVHPAAGELYFLKMLLNHVKGATSFEYLRCVSGIVYPTFQLACKALGLLDDGKEWAEAFSEA
VLTASSSQLRQLFVSVTLFCQIANPQDLLDQFWHTMHDDIRIKLSSFSPHNLHFSDNELKNYVLYELEQLFNALATSLKDYNLPLPNDRLMSEIRNNLLR
EELNYDISELRSNNEASISLLNTCQKKIYDRVMESISKNQQSLIFVYGHGGTGKTFLWHSLINSIRSEGLIVLAVASSGIASILLPGGRTAHSRFKIPLA
INENSTCEIKKNTHLSRLIETTTLIVWDEAPMNNRYCFETLDRSLRDIMGQTGHSNHNQPFGGKSILLGGDYRQILPVIPGGTKEDIINASLSSSPLWPK
FEIMLLKQNMRLSIAGLGSDEINEIKTFAKWILKIGDGDLCDIPFFDELDESLIKIPCDLQLHTSGDPIKAMVSAIYPSIEQPALEPFYFKERAIITPKN
ITVTEINNFILGVTHGPQRIYLSNDSVDASSSDNDNINLLYPLEFINQLEFSGVPSHILALKIGAPIMLLRNLSPMIGLCNGTRLIITQLADRVIEAQII
TGSHIGDRVFIPRIIFPINDDKCPFTIKRRQFPIRLCYAMTINKSQGQSLKFVGVFLKEQVFAHGQLYVALSRVTSKKGLKIISCDQEGKQLDYAKNIVY
KDVLNLLPKGLFPIQNDF
|
DESeq2's median of ratios [POPLAR]
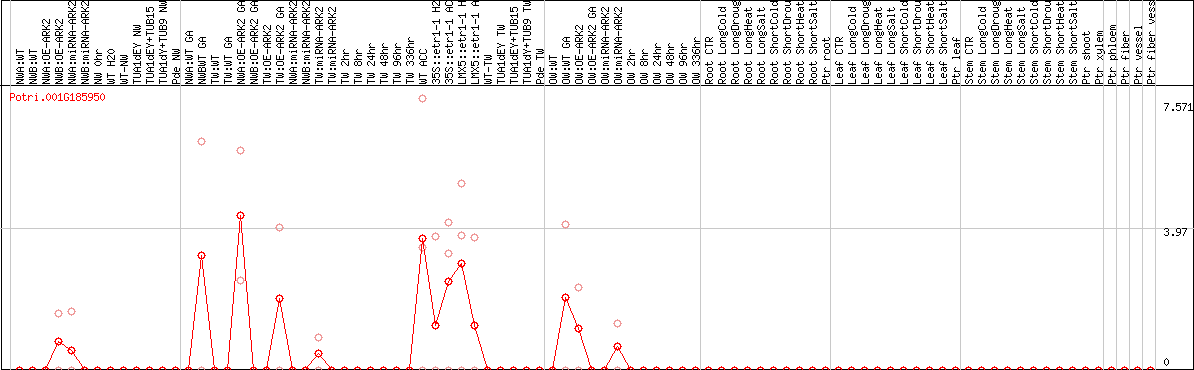
Coexpressed genes
Potri.001G185950 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
