Potri.001G189500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G189500.3 pacid=42788312 polypeptide=Potri.001G189500.3.p locus=Potri.001G189500 ID=Potri.001G189500.3.v4.1 annot-version=v4.1
ATGGACGGCGTAGAGAGAGCTCGAGCTTCCGGTAGACGATCAAGTCACAACAGCTTGAGCAGGAGCATAAGCAGGAGCTTGAGCAGGGCTAGTTGGAACA
TGGAAGACATGTTTTCAGTTGGCAGGCAATCTAGGAGAAGCAACCTTGTCGATGAAGATGAAGAAGCTCTAAAATGGGCTGCCATAGAGAAGTTACCGAC
ATACAATCGGTTAAGAACAAGTATCATCAAATCTTTCGTGGATACTGAGGACCAAGGCAACAAGATGCTGCAGCATAAAGAAGTTGATGTGAGAAAGCTT
GACATAAATGAGAGACAAAATTTTATCGACAAGCTCTTTAAGGTTGCAGAGGAAGATAATGAGAAATACTTGAAGAAATTCAGACAAAGAGTCGATAAGG
TTGGCATCCGACTCCCGACAATTGAGGTCAGGTTTGATCATTTAACAATTGAAGCTGACTGCCATTTTGGCACCAGAGCTCTCCCCACCCTTCCAAATGC
TGCAAGAAACATGTTTGAATCAGCTCTTGGTGTAGTTGGGATTAATTTGGCTCAGAGAACTAAGCTCACAATCCTCAAAGATGCATCTGGGGTTATAAAA
CCTTCACGGATGGCACTACTACTAGGACCACCATCCTCAGGAAAAACAACTCTTTTGTTGGCTCTAGCAGGAAAGTTGGACCCGAGTCTTAAGGTTACAG
GAGACCTAACATACAATGGATATGAATTCAAGGAATTTATGCCACGGAAATCATCAGCATATATCAGCCAAAACGATGTTCACATAGGAGAAATGACCGT
GAAAGAAACCTTGGATTTCTCAGCAAGGTGTCAAGGGGTCGGGACACGATATGATCTCCTAAGTGAGCTTGCAAGAAGAGAGAAGGATGCTGGAATATTT
CCGGAGGCAGAAGTGGATCTTTTCATGAAGGCAACTGCAATGGAAGGAGTTGAAAGCAGTCTCATTACTGACTACACACTCAAAATACTAGGACTTGATA
TATGCAAGGATACCATCGTCGGAGATGACATGATACGAGGGATATCGGGTGGACAGAAAAAGCGAGTGACTACAGGAGAGATGATTGTTGGGCCCACAAA
AACACTTTTCATGGATGAGATATCCACCGGTCTAGATAGCTCCACAACATATCAAATAGTGAAGTGCTTGCAGCACATTGTACACTACACCGAGGCTACA
ATCTTGGTGTCCCTGCTCCAACCTGCTCCTGAGACGTTTGATCTCTTTGATGATATCATCCTTTTATCAGAAGGCCAGATTGTTTACCAGGGCCCACGAG
AACACATTCTTGCTTTCTTTGAGAGCTGTGGGTTCCGCTGTCCTGAGAGGAAGGGCACAGCTGATTTCTTACAAGAGGTTACTTCGAAGAAAGACCAGGA
ACAGTATTGGGATGATAGAAACAAGCCATATAGATACGTAACAGTCCCAGAATTTGTTGAAAGGTTCAAGAGGTTCCATGTGGGGATGAGGCTAGAGAAT
GAGCTTTCCGTGCCATTTGACAAGACCCAAGGCCACAAAGCAGCTCTGTCATTCTCGAAGTATTCTGTTCCCAGAATGGAACTACTCAAGGCATGTTGGG
ATCGAGAATGGATACTGGTTAAGAGAAATGCATATGTTTATGTAGCCAAGACGGTTCAACTTATTATTATGGCAATTATAATGTCCACGGTGTTCATCAA
GTCTAAAATGCACACAAGGAATGAAGGAGATGGAGCGGTATACATTGGTGCCCTTTTGTTTACTATGATCATTAACATGTTCAATGGTTTTGCTGAGCTC
TCACTTGTAATCAAGAGGCTTCCAGTATTTTACAAGCAAAGAGATCTTCAATTCCATCCTGCCTGGACTTTCACTCTGCCAACTTTCTTGCTACAGCTGC
CAATGTCTATAATTGAGTCTGTTGTTTGGGTGTCAATTACCTATTACTCCGTTGGTTTTGCACCTGACGCTAGCAGGTTTTTCAAGCAACTGCTGCTGGT
ATTTTTCATCCAACAGATGGCTTCTGGGCTCTTTAGGCTTATTGCTGGGGTCTGCAGAACCATGATCATTGCCAACACCGGCGGGGCTCTCACTCTACTC
CTTGTTTTCTTGCTTGGAGGTTTCATCTTACCTAAAGGTGCCATTCCAGATTGGTGGGGATGGGGTTATTGGGTTTCACCTTTGTCTTATGGTTTCAATG
CCATAGCTGTGAACGAAATGTCTGCACCAAGGTGGATGAACAAAAACAGTTCAGACGCCTCTACCAGTTTAGGCACAGCAGTGCTCAAGAACTTTGATGT
TTACACAGATAAGAACTGGTATTGGATTGGCACAGCTGCTATTCTAGGCTTTGCTGTTCTATTCAATGTTCTCTTCACCTTTGCTCTCGCGTACTTTAGT
CCTGCTGGAAAATCACAGGCCATAATCTCTGAGGAAACAACAAAAGAAAGAACCAGGTCAACACAGTCGTTATCCCATTCTAATGGAAACAATACAAGTG
AAATGGCAATCTTGAGAACGAGGAGCCCATCCAATCCCAATGGACTAAGTGGAAATGCTGATTCAATTGAGGCAGCAAATGGAGTGGCTCCTAAGAGAGG
AATGGTTCTTCCTTTCTCTCCTCTAGCCATGTCCTTTGACAGTATGAATTATTTTGTGGACATGCCCCCGGAAATGAAGGAGCAAGGAGTTCCAGAGGAT
AGGCTGCAACTACTTCGAGAAGTAACAGGAGCATTTAGGCCTGGAGTACTCACGGCATTAATGGGAGTCAGTGGAGCTGGAAAGACCACACTGATGGATG
TTTTGGCAGGAAGAAAGACTGGTGGATACATTGAAGGTGAAATTAAAATTTCCGGGTTCCCCAAGAAACAGGAAACGTTTGCAAGAATTTCTGGATATTG
CGAACAGAATGATATCCACTCTCCTCAAGTCACTGTTAAAGAATCATTGATTTATTCAGCATTTCTTCGGCTCCCTAAAGAAGTCAGCAAACAAGAAAAG
ATGATTTTCGTGGATGAAGTGATGGAGTTGGTCGAGCTAAACAATCTCAAGGATGCTGTAGTTGGGCTTCCAGGAATCACAGGGTTGTCAACAGAACAGA
GAAAGAGGTTAACAATAGCAGTGGAACTGGTTGCTAATCCCTCCATCATTTTCATGGATGAACCAACTTCTGGTCTTGATGCAAGGGCAGCTGCCATTGT
TATGAGGACTGTGAGAAACACTGTGGATACTGGGAGAACAGTTGTCTGTACGATCCATCAACCTAGCATTGACATCTTTGAAGCCTTTGATGAATTGTTA
TTAATGAAGAGGGGAGGACAAGCAATCTACTCAGGACCATTGGGTCGAAACTCTCACAAGATCATTGAATACTTCGAGGCCATTCCTGGAGTCCCTAAAA
TCAAAGAGAAGTACAATCCAGCAACATGGATGCTAGAAGTGAGTTCAGTTGCAGCTGAAGTCCGGCTTGGAATGGACTTTGCGGAGCAGTACAGATCTTC
ATCCTTGCATCAGAGAAACAAGGCTTTGGTGAAGGAGTTGAGCACACCACCACCAGGAGCAACAAACCTTTATTTCGCCACTCAGTATTCAGAGTCTGCA
TGGGGGCAGTTCAAATCTTGCCTTTGGAAGCAGTGGTGGACATACTGGAGAAGTCCTGATTACAACCTTGTTAGATACTTCTTCACCTTGGTTTGTGCTC
TTATGGTTGGTAGCATATTCTGGAAGGTCGGGACAAAAAGGGATAGCTCAAGCGATCTGAATATGATTATCGGTGCCATGTATGCTTCTGTCTTGTTTGT
TGGTATTAACAACTGCTCAACTGTACAACCAGTTGTAGCAGTCGAAAGAACCGTGTTTTACCGAGAAAAAGCAGCTGGAATGTATTCTGCATTACCCTAT
GCCATTGCACAGGTTGTTTGTGAAATACCATACGTATTTGTTCAAACTACATACTATACACTTATTGTGTATGCCATGGTGTCCTTCGAATGGACTGCAG
CCAAATTCTTTTGGTTCTTCTTTGTCAATTTCTTTTCCTTCCTTTACTTCACATACTATGGAATGATGACTGTTTCCGTTACCCCAAACCACCAAGTAGC
AGCCATCTTTGCAGCAACATTCTATTCTCTCTTCAATCTTTTCTCTGGCTTCTTTATACCAAGACCGAAAATTCCCAAGTGGTGGGTCTGGTACTATTGG
ATTTGCCCTGTAGCGTGGACAGTGTATGGACTAATCGTGTCCCAATATGGTGATGTTATGGACACCATTAATGTTCCTGGTCGCGCTGGTGCTGATCCTA
CAATAAAGGTTTATATACAAGAAAATTTTGGATATGACCCAGATTTCATGGGGCAGGTTGCCGCAGTCTTGGTTGGCTTTACAGTCTTCTTCGCATTCCT
GTTTGCCTTCTGCATAAGGACACTGAACTTCCAGACAAGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G189500.3 pacid=42788312 polypeptide=Potri.001G189500.3.p locus=Potri.001G189500 ID=Potri.001G189500.3.v4.1 annot-version=v4.1
MDGVERARASGRRSSHNSLSRSISRSLSRASWNMEDMFSVGRQSRRSNLVDEDEEALKWAAIEKLPTYNRLRTSIIKSFVDTEDQGNKMLQHKEVDVRKL
DINERQNFIDKLFKVAEEDNEKYLKKFRQRVDKVGIRLPTIEVRFDHLTIEADCHFGTRALPTLPNAARNMFESALGVVGINLAQRTKLTILKDASGVIK
PSRMALLLGPPSSGKTTLLLALAGKLDPSLKVTGDLTYNGYEFKEFMPRKSSAYISQNDVHIGEMTVKETLDFSARCQGVGTRYDLLSELARREKDAGIF
PEAEVDLFMKATAMEGVESSLITDYTLKILGLDICKDTIVGDDMIRGISGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTYQIVKCLQHIVHYTEAT
ILVSLLQPAPETFDLFDDIILLSEGQIVYQGPREHILAFFESCGFRCPERKGTADFLQEVTSKKDQEQYWDDRNKPYRYVTVPEFVERFKRFHVGMRLEN
ELSVPFDKTQGHKAALSFSKYSVPRMELLKACWDREWILVKRNAYVYVAKTVQLIIMAIIMSTVFIKSKMHTRNEGDGAVYIGALLFTMIINMFNGFAEL
SLVIKRLPVFYKQRDLQFHPAWTFTLPTFLLQLPMSIIESVVWVSITYYSVGFAPDASRFFKQLLLVFFIQQMASGLFRLIAGVCRTMIIANTGGALTLL
LVFLLGGFILPKGAIPDWWGWGYWVSPLSYGFNAIAVNEMSAPRWMNKNSSDASTSLGTAVLKNFDVYTDKNWYWIGTAAILGFAVLFNVLFTFALAYFS
PAGKSQAIISEETTKERTRSTQSLSHSNGNNTSEMAILRTRSPSNPNGLSGNADSIEAANGVAPKRGMVLPFSPLAMSFDSMNYFVDMPPEMKEQGVPED
RLQLLREVTGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIEGEIKISGFPKKQETFARISGYCEQNDIHSPQVTVKESLIYSAFLRLPKEVSKQEK
MIFVDEVMELVELNNLKDAVVGLPGITGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELL
LMKRGGQAIYSGPLGRNSHKIIEYFEAIPGVPKIKEKYNPATWMLEVSSVAAEVRLGMDFAEQYRSSSLHQRNKALVKELSTPPPGATNLYFATQYSESA
WGQFKSCLWKQWWTYWRSPDYNLVRYFFTLVCALMVGSIFWKVGTKRDSSSDLNMIIGAMYASVLFVGINNCSTVQPVVAVERTVFYREKAAGMYSALPY
AIAQVVCEIPYVFVQTTYYTLIVYAMVSFEWTAAKFFWFFFVNFFSFLYFTYYGMMTVSVTPNHQVAAIFAATFYSLFNLFSGFFIPRPKIPKWWVWYYW
ICPVAWTVYGLIVSQYGDVMDTINVPGRAGADPTIKVYIQENFGYDPDFMGQVAAVLVGFTVFFAFLFAFCIRTLNFQTR
|
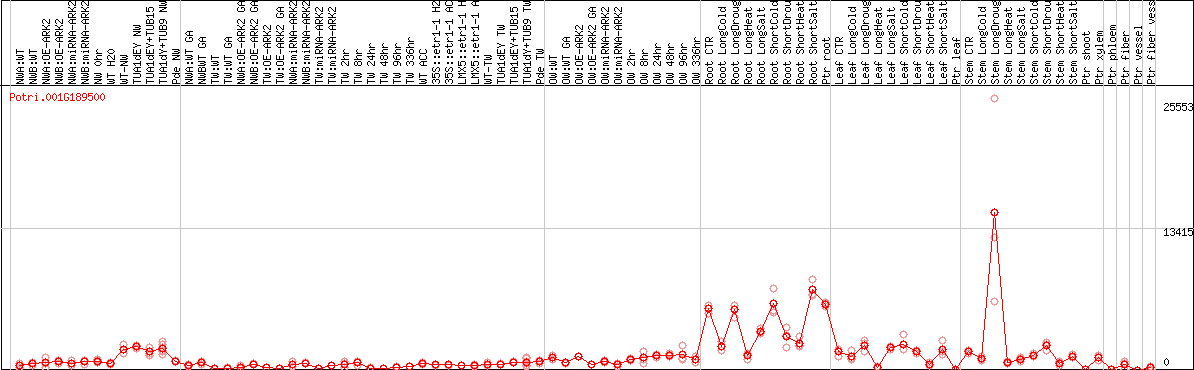
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G189500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
