External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G19210 116 / 4e-32
Leucine-rich repeat transmembrane protein kinase protein (.1)
AT1G51880 112 / 1e-30
RHS6
root hair specific 6 (.1)
AT1G51890 112 / 2e-30
Leucine-rich repeat protein kinase family protein (.1.2)
AT1G51870 108 / 3e-29
protein kinase family protein (.1)
AT1G51850 107 / 1e-28
Leucine-rich repeat protein kinase family protein (.1)
AT2G19230 107 / 1e-28
Leucine-rich repeat transmembrane protein kinase protein (.1)
AT1G51860 106 / 2e-28
Leucine-rich repeat protein kinase family protein (.1)
AT4G29990 106 / 2e-28
Leucine-rich repeat transmembrane protein kinase protein (.1)
AT5G16900 106 / 2e-28
Leucine-rich repeat protein kinase family protein (.1)
AT2G04300 105 / 3e-28
Leucine-rich repeat protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G094600
94 / 6e-24
AT4G29990 636 / 0.0
Leucine-rich repeat transmembrane protein kinase protein (.1)
Potri.019G094300
92 / 1e-23
AT4G29990 628 / 0.0
Leucine-rich repeat transmembrane protein kinase protein (.1)
Potri.019G094200
89 / 2e-22
AT4G29990 452 / 1e-150
Leucine-rich repeat transmembrane protein kinase protein (.1)
Potri.019G094700
89 / 4e-22
AT4G29990 652 / 0.0
Leucine-rich repeat transmembrane protein kinase protein (.1)
Potri.003G053301
74 / 3e-17
AT1G52290 531 / 0.0
proline-rich extensin-like receptor kinase 15, Protein kinase superfamily protein (.1)
Potri.004G153600
74 / 4e-17
AT4G34440 551 / 0.0
proline-rich extensin-like receptor kinase 5, Protein kinase superfamily protein (.1)
Potri.011G131200
74 / 6e-17
AT5G54590 631 / 0.0
calcium/calmodulin-regulated receptor-like kinase 1, Protein kinase superfamily protein (.1.2)
Potri.008G111600
73 / 8e-17
AT1G68690 622 / 0.0
proline-rich extensin-like receptor kinase 9, Protein kinase superfamily protein (.1)
Potri.009G115200
73 / 9e-17
AT4G34440 575 / 0.0
proline-rich extensin-like receptor kinase 5, Protein kinase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025849
95 / 3e-24
AT4G29990 803 / 0.0
Leucine-rich repeat transmembrane protein kinase protein (.1)
Lus10020452
75 / 4e-17
AT5G10530 510 / 2e-173
Concanavalin A-like lectin protein kinase family protein (.1)
Lus10011917
74 / 4e-17
AT2G20300 951 / 0.0
Abnormal Leaf Shape 2, Protein kinase superfamily protein (.1)
Lus10007077
74 / 5e-17
AT5G10530 511 / 4e-171
Concanavalin A-like lectin protein kinase family protein (.1)
Lus10022848
74 / 5e-17
AT2G20300 912 / 0.0
Abnormal Leaf Shape 2, Protein kinase superfamily protein (.1)
Lus10022129
73 / 1e-16
AT3G24550 474 / 3e-159
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Lus10027635
73 / 1e-16
AT5G56890 887 / 0.0
Protein kinase superfamily protein (.1)
Lus10011098
73 / 2e-16
AT3G24550 700 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Lus10023645
72 / 2e-16
AT3G24550 691 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Lus10017898
72 / 3e-16
AT1G68690 631 / 0.0
proline-rich extensin-like receptor kinase 9, Protein kinase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF07714
PK_Tyr_Ser-Thr
Protein tyrosine and serine/threonine kinase
Representative CDS sequence
>Potri.001G191701.1 pacid=42793567 polypeptide=Potri.001G191701.1.p locus=Potri.001G191701 ID=Potri.001G191701.1.v4.1 annot-version=v4.1
ATGACCAACTTCGGAAGAATTCTCGGTAAGGGAGGATTTGGGACAGTTAACCATGGCTACTTGAATGATACTCAAGCAGCTGTCAAGATGTTGACTCCAT
CATCGGTTCAAGGGTACAAGGAGTTTGAGGCAGAGGTTAAACTTTTTTTGAGAGTGCATCATAGAAACTTGACTAATCTTGTTGGGTACTGCCATGAAGG
CACCAAAATGGGTCTTGTGTATGTACGTACCTGGCTAATGAAAATTTGA
AA sequence
>Potri.001G191701.1 pacid=42793567 polypeptide=Potri.001G191701.1.p locus=Potri.001G191701 ID=Potri.001G191701.1.v4.1 annot-version=v4.1
MTNFGRILGKGGFGTVNHGYLNDTQAAVKMLTPSSVQGYKEFEAEVKLFLRVHHRNLTNLVGYCHEGTKMGLVYVRTWLMKI
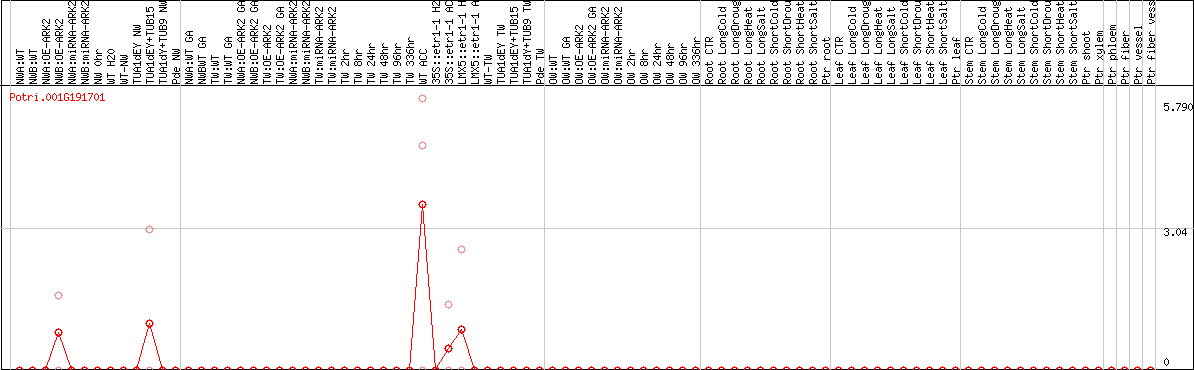
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G191701 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.