Potri.001G207700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G207700.4 pacid=42789855 polypeptide=Potri.001G207700.4.p locus=Potri.001G207700 ID=Potri.001G207700.4.v4.1 annot-version=v4.1
ATGGCGTCGAAAGGCCCAAGGTCTAAGCTTGATCACGAGACCAGAGCTCGGCGTCAAAAGGCACTGGAAGCTCCGAGGGAGCCGCGTCGCCCTAAGACTC
ACTGGGATCACGTCTTAGAGGAAATGGTTTGGTTGTCGAAGGACTTTGAGTCGGAGAGGAAATGGAAATTGGCACAAGCGAAGAAAGTGGCATTAAGAGC
CAGTAAGGGCATGTTGGATCAGGCCACCAGAGGAGAGAAGAAACTGAAGGAAGAGGAGCAGCGGCTGCGAAAAGTTGCTCTTAATATTTCAAAGGATGTA
AAGAAATTCTGGGTTAAAATAGAAAAGCTGGTACTTTACAAGCATCAGATGGAGCTTGATGAGAAGAAGAAAAAGGCACTTGATAAGCAGCTTGAGTTTC
TTCTAGGACAAACTGAGAGGTACTCTACAATGTTAGCAGAAAATCTAGTGGACAAACCATCAGAGCAATATGCTGCACAAGATAAACCGAGAATTGCATA
CAAGAAAGGAGATGATGCTAATATACCTGAGCAAGTCAATGATGAGCCTCAGTTGGACACTACAGACAATGACGATGAGTATGATGTGCAATCTGAAGAT
GAAGTGGAAGATGATGAGCATACTATTGAGGAAGATGAGGCTCTTATAACTGCAGAAGAAAGGCAAGAAGAATTGGAAGCTCTGCACAATGAAACAGATA
TTCCACTCGAGGAGCTACTTAACCGTTATCCTGTAGAGAAAGGTAGCGGAGAAAGTAGTGAAAATGGAGCTAAGCCATCTGCAAATGGGGAAGACCATTG
TGAGAGGAAAGGAAACGATATGTCTGCTGCTAGTGACATGGAGATAAGCTGCTCACCTGTTAATGCTAGTCGTCGTTGTGGTGAAAATAATGGTGCCTTG
CCCATTCCAGACAATGACTTATTGGAAATCAGGACCAATGAAACCAGAAATCAGTTAAGTATTTCTGATGATCCGGCCAAAGAACGTGTGCCATATGATT
TCAGTGATGAACAGGAAGATGGTGATTTTGATCTTGCTGCAGAAGAAGAAAAGGATGACGAGACAACCTTGTTGGAAGAGGAAGAGCTGGCAAAAGCAGA
TTCAAAGAATCCCATAGATGAGATCTCATTGCTGCAAAAGGAGAGTGAAATCCCTTTGGAAGAATTGCTTGCAAGGTATACCAAGGAGCCTAACAATGAA
GTTTCAGAGGATGAATCTGAATATGCATCTTTATTATCAGACAATGTGTCAAATTCCCCAGGCCATGAAGATGAACTGAAGCAGCTGGATAACTCAATGG
ATGGAATTGTTGAATGTGGAAACCATCCTCTTGTAGAAGAACAAGAAAAGGGAAATGAAAAAATTTCAGAAGATGGAAGGGAGAGTGAGAATAGGATTGC
TGATGCTGCTGCTGCTGCAAGATCTGCACAACCAACAGGCAATACATTCTCAACAACTAAAGTGCGCACGAAGTTCCCTTTTCTTCTTAAGTACCCTCTT
CGTGAATATCAACATATAGGCCTTGATTGGCTTGTTACAATGTATGAGAAAAGATTAAATGGGATTCTAGCTGATGAAATGGGGCTTGGAAAGACAATCA
TGACTATTGCTTTGCTTGCACACCTGGCATGTGAAAAGGGAATATGGGGTCCTCATCTGATCGTGGTTCCCACAAGTGTCATGCTTAACTGGGAAACAGA
GTTTTTTAAATGGTGTCCTGCCTTCAAGATTTTAACATATTTTGGCAGTGCAAAAGAGCGTAAATGCAAGAGGCAAGGTTGGCTGAAACCGAACTCCTTC
CATGTATGCATAACAACTTACAGACTGGTTATACAGGATTCCAAAGTCTTCAAAAGGAAGAAATGGAAGTATTTGATATTGGATGAAGCTCATCTGATTA
AAAATTGGAAGTCTCAGAGATGGCAAACTCTTTTAAACTTTAATTCAAAACGACGAATCTTATTAACTGGTACACCTCTGCAGAATGATCTCATGGAACT
CTGGTCACTGATGCATTTCTTGATGCCCCACATCTTTCAATCTCACCAGGAATTCAAGGACTGGTTTAGCAATCCAATAACTGGGATGGTGGAGGGACAA
GAAAGAGTGAACAAAGAAGTTGTTGATCGTTTGCATAATGTTCTCCGTCCATTCATCCTCCGTCGATTGAAAAGGGATGTGGAGAAGCAGCTGCCCATGA
AACATGAGCATGTCATCTATTGTAGACTTTCAAGGAGGCAGCGTAACTTGTACGAGGATTTCATTGCTAGCTCAGAGACACAAGCTACCCTTGCCACTGC
CAACTTTTTTGGGATGATCAGTATCATAATGCAACTTCGTAAAGTTTGTAACCATCCTGACTTATTTGAGGGTCGTCCAATTATTAGTTCTTTTGATATG
GCTGGTATAGACATGCAATTAAGTAGTTCTGTTTGTTCAATGCTTTCACCTGGGCCATTGTCATCAGTAGATCTTTGTGCTTTGGGGCTTATATTTACTC
ATCTTGATTTTAGCATGGCCTCTTGGGAGTACGATGAAGTAAAGTCTATTGCAACTCCATCAAGATTAATTAAGGAGCGTTCCAACCTGGATAACATAGA
AGAAGTTGGACCGGGATCTAAACATTGGAAGAAGTTGCCTGGAAAAAATATTTTTGAAGAGATTAGAAAGTCATTATTGGAGGAGAGATTAAGAGAAGTG
AAGCAACGGGCAGCATCTATTGCATGGTGGAATTCCTTGAGGTGTCAGAAAAAACCCATATACTCAACAACCCTACGAGAACTTCTCACAGTGAAGCATC
CCATCTATGATGTCCATCGCCATAAAACTGAGCGACTCTCTTACTTGTACTCTTCTAAGCTGGGTGATGTTATTCTTTCACCAATTGAACGGTTCCAGAA
GATGACTGATCTTGTGGAATCATTTATGTTTGCAATTCCAGCAGCACGTACCCCAGTACCTGTCTTCTGGTGTAGTCAGATTAGAACTCCAGTGTTTCTT
CATTCAACTTATGAGGAGAAATGCTCTGAAATGTTGTTACCTCTTCTTTCACCTATCAGACCTGCAATTGTTCGGAGACAACTTTATTTTCCAGACAGGC
GTCTAATACAGTTTGACTGTGGTAAATTGCAGGAGCTTGCAATTTTGCTCAGGAAGTTGAAGTCAGAAGGACACCGGGTATTAATATTCACACAGATGAC
TAAGATGCTTGATATTTTGGAGGTTTTCATGAATTTGTATGGCTACACTTACATGCGCTTAGATGGATCCACTCAACCAGAGGAGAGACAAACATTAATG
CAGCGTTTCAACACAAATCCCAAAATCTTTATCTTCATTTTATCAACCCGTAGTGGAGGTGTTGGCATCAATCTTGTAGGGGCAGATACTGTAATCTTCT
ATGATAGTGACTGGAATCCTGCTATGGATCAACAGGCCCAAGACCGCTGCCATCGGATAGGACAGACACGTGAAGTACATATCTACAGGTTAATCAGTGA
GAGCACCATTGAAGAAAACATTTTAAAGAAAGCCAATCAGAAGCGTGCTCTTGATGATCTAGTAATACAGAGTGGAGGGTACAATACGGAATTTTTCAAG
AAGCTCAATCCTATGGAATTATTCTCAGGGCATAAAACACTTCAGATTAAGAATATGCAGAGGGAGAAAAACCATAACAATGGAAACGAGGTGTCTTTGT
CTAATGCAGATGTGGATGCTGCTCTGAAATATGCAGAAGATGAAGCAGATTACATGGCATTGAAGAAAGTCGAACAGGAAGAGGCTGTAGACAACCAAGA
GTTTACTGAAGAGGCCATTGGGAGATTGGAAGATGATGAATTTGTGAATGATGATGATATGAAGGCTGATGAGCCTACAGACCATGAGATGACAACTTAT
AGTAAAGATGGTGCAGTGAATTTAAAAGAGAATGGTTGCATTGAAGAGAGAGCTGTAACTCTTACTGGCAATGAAGATGTTGATATGTTGGCTGATGTCA
AACAGATGGCGGCGGCGGCGGCTGCTGCTGGACAAGCTATCTCATCCTTTGAGAATCAGCTACGTCCAATTGACAGATATGCGGTTCGGTTTCTGGAATT
GTGGGATCCAATTATAGACAAAGCAGCCTTGGAATCTCAAGTTGGGTTTGAGGAGACCGAATGGGAGCTTGATCGTATTGAGAAGTACAAGGAGGAAATG
GAAGCTGAGATTGATGATGATGAGGAACCATTGGTTTATGAAAGATGGGATGCTGATTTTGCGACTGAGGCATACCGACAAGAAGTTGAGGCATTAACTC
AACATCAGCTGTTGGAAGAACAGGAAGCTGAAGCTAATGAGAAGGAAGGTGCAGATGATGGACATTTAGATGCTATGGTGTATAAAATGCCACGTAATCC
TAAATTGAAGTCTAAGAAGAAACCAAAGAAAGCCAAGTTCAAATCTCTGAAGAAAGAATCTTTGACTTCTGAATTGAAACACGTGAAAGAGGAGGTATCA
ATGGAAACTTTGTCTGTAGATGATGATGATGATGGTACATATTCGGATACGATGTCTCCTTGTTCCAGTATGTGGAGAAAGCGTAAGAAAGCAGAATCAG
CAATTTGTATTGACAAAACGAGGTCGAAGAAGACTAAGAAATTCAAAAAGGGTCCAGAAACGTGCACTTTTAGTGTAGATTCAGACCTGTCTGGAAAGCA
GCATGACAGATTTACGGAACTAAAACCATATGAGGTTGTGGTTTCCGATATTGAGCAGAAGCCAGCTAGCAGGAGCAAAATGGGAGGGAAAATATCCATC
AGTACCATGCCAGTGAAACGGGTTTTGATGATAAAGCCAGAGAAGTTGAAGAAGGGGAATGTTTGGTTGAAAGATTGTGTTCCACCACCTGCTTTATGGA
TGCCACAGGAGGATGCAGTATTATGTGCTGTTGTACATGAATATGGTCCACATTGGAGCTTGGTCAGTGAAATTCTGTATGGGATGACTGCTGGTGGGTT
TTATCGGGGAAGATACCGCCATCCAGTTCATTGTTGTGAGAGATTTAGGGAGCTAATCCACAGATATGTTTTATTTTCTCCGGAAAATCCTATTAATAAT
GAAAAGATGAGCAACATGGTCCCTGGGAAGGCCCTTCTGAAAGTAACCGAGGATAACATACGGATGTTGTTAAATGTTGTTGCTGAGCAGCCAGACCATG
AGTTACTTCTCCAGAAGCACTTCACAGCACTGCTTTCTTCTGTGTGGAGGGTGAAATCTCGTGTTGAAAATCAACAAAACATGCCATCATCTCGAAATGC
CCTTTATAACAGTGGAAGGGTTTTCAATTCTTCTGTTAATCCGTTACCTTGGAACTCCTTGAGGGAATCGGCAAAAAGAATGAAATTCACTAATTTGGGA
CAGAGTACCAAGTTGCTGGCAGCTGCACTCCATGATGCCAGCAGTAGAAGGCCAGGTGATAGAGTTTCCAATTCCAATGTGAATGAAGAAGCTCCTGCTG
TAGGCGAGAAGCTGGAAATAACATTGGAATTCCAAAAGGAGGAAAATGATTATTTGATTCCATTCCCACCTGTAATAAGTTTGTCAATACCTGGCTCAGC
ACCATGGATGTCTGTAAATAAGGATAGAGCGGCGGCTCACCATCTCAGGGCATCCACAAGCATAGCTGAAAATCGATTCAGGGATGCAGCAAGAGCTTCT
TCATCTGTGTTACCAGCAAATGATTTGAAGTTGTGGCTGGCTTCAAAGACTCAGTCCTTGGGAAAGCACAAGCTAACTGTCTCTGAGTCAACCAAGCCTC
CTAGGTCAAAGACGAGGAAAACTTTGCTGGAGCAAAATGAAGGACATGCCGAGCCAGTAATGCAACCACTGTCAGATAGGGATCCCAATTTAAGGTTTGA
CCTGCCACCAGAAGTCATTCAGGATGATAAGGATGGGTTTTCTATTTCATTCATGGAAAAAGAACTTTCAGTGGAAACGAAGATCTCCGAAGCTGTCCCG
CATATCTATGTTCCTGATTTGATATTGGGTCTTGATGATTATTCTTTGTTGCCGGAATACACTGACATTGGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G207700.4 pacid=42789855 polypeptide=Potri.001G207700.4.p locus=Potri.001G207700 ID=Potri.001G207700.4.v4.1 annot-version=v4.1
MASKGPRSKLDHETRARRQKALEAPREPRRPKTHWDHVLEEMVWLSKDFESERKWKLAQAKKVALRASKGMLDQATRGEKKLKEEEQRLRKVALNISKDV
KKFWVKIEKLVLYKHQMELDEKKKKALDKQLEFLLGQTERYSTMLAENLVDKPSEQYAAQDKPRIAYKKGDDANIPEQVNDEPQLDTTDNDDEYDVQSED
EVEDDEHTIEEDEALITAEERQEELEALHNETDIPLEELLNRYPVEKGSGESSENGAKPSANGEDHCERKGNDMSAASDMEISCSPVNASRRCGENNGAL
PIPDNDLLEIRTNETRNQLSISDDPAKERVPYDFSDEQEDGDFDLAAEEEKDDETTLLEEEELAKADSKNPIDEISLLQKESEIPLEELLARYTKEPNNE
VSEDESEYASLLSDNVSNSPGHEDELKQLDNSMDGIVECGNHPLVEEQEKGNEKISEDGRESENRIADAAAAARSAQPTGNTFSTTKVRTKFPFLLKYPL
REYQHIGLDWLVTMYEKRLNGILADEMGLGKTIMTIALLAHLACEKGIWGPHLIVVPTSVMLNWETEFFKWCPAFKILTYFGSAKERKCKRQGWLKPNSF
HVCITTYRLVIQDSKVFKRKKWKYLILDEAHLIKNWKSQRWQTLLNFNSKRRILLTGTPLQNDLMELWSLMHFLMPHIFQSHQEFKDWFSNPITGMVEGQ
ERVNKEVVDRLHNVLRPFILRRLKRDVEKQLPMKHEHVIYCRLSRRQRNLYEDFIASSETQATLATANFFGMISIIMQLRKVCNHPDLFEGRPIISSFDM
AGIDMQLSSSVCSMLSPGPLSSVDLCALGLIFTHLDFSMASWEYDEVKSIATPSRLIKERSNLDNIEEVGPGSKHWKKLPGKNIFEEIRKSLLEERLREV
KQRAASIAWWNSLRCQKKPIYSTTLRELLTVKHPIYDVHRHKTERLSYLYSSKLGDVILSPIERFQKMTDLVESFMFAIPAARTPVPVFWCSQIRTPVFL
HSTYEEKCSEMLLPLLSPIRPAIVRRQLYFPDRRLIQFDCGKLQELAILLRKLKSEGHRVLIFTQMTKMLDILEVFMNLYGYTYMRLDGSTQPEERQTLM
QRFNTNPKIFIFILSTRSGGVGINLVGADTVIFYDSDWNPAMDQQAQDRCHRIGQTREVHIYRLISESTIEENILKKANQKRALDDLVIQSGGYNTEFFK
KLNPMELFSGHKTLQIKNMQREKNHNNGNEVSLSNADVDAALKYAEDEADYMALKKVEQEEAVDNQEFTEEAIGRLEDDEFVNDDDMKADEPTDHEMTTY
SKDGAVNLKENGCIEERAVTLTGNEDVDMLADVKQMAAAAAAAGQAISSFENQLRPIDRYAVRFLELWDPIIDKAALESQVGFEETEWELDRIEKYKEEM
EAEIDDDEEPLVYERWDADFATEAYRQEVEALTQHQLLEEQEAEANEKEGADDGHLDAMVYKMPRNPKLKSKKKPKKAKFKSLKKESLTSELKHVKEEVS
METLSVDDDDDGTYSDTMSPCSSMWRKRKKAESAICIDKTRSKKTKKFKKGPETCTFSVDSDLSGKQHDRFTELKPYEVVVSDIEQKPASRSKMGGKISI
STMPVKRVLMIKPEKLKKGNVWLKDCVPPPALWMPQEDAVLCAVVHEYGPHWSLVSEILYGMTAGGFYRGRYRHPVHCCERFRELIHRYVLFSPENPINN
EKMSNMVPGKALLKVTEDNIRMLLNVVAEQPDHELLLQKHFTALLSSVWRVKSRVENQQNMPSSRNALYNSGRVFNSSVNPLPWNSLRESAKRMKFTNLG
QSTKLLAAALHDASSRRPGDRVSNSNVNEEAPAVGEKLEITLEFQKEENDYLIPFPPVISLSIPGSAPWMSVNKDRAAAHHLRASTSIAENRFRDAARAS
SSVLPANDLKLWLASKTQSLGKHKLTVSESTKPPRSKTRKTLLEQNEGHAEPVMQPLSDRDPNLRFDLPPEVIQDDKDGFSISFMEKELSVETKISEAVP
HIYVPDLILGLDDYSLLPEYTDIG
|
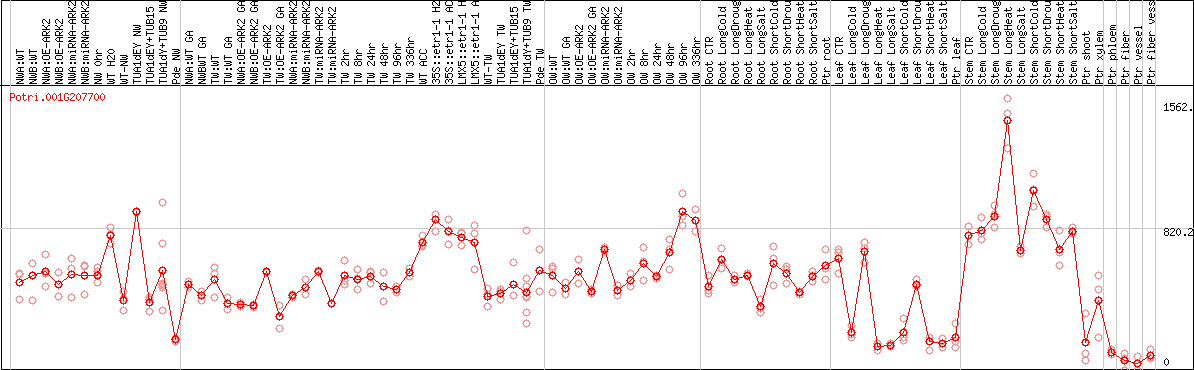
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G207700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
