External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G26070 253 / 5e-85
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
AT3G26080 229 / 1e-75
plastid-lipid associated protein PAP / fibrillin family protein (.1)
AT5G19940 55 / 8e-09
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
AT5G09820 54 / 2e-08
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
AT4G22240 44 / 7e-05
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
AT2G35490 42 / 0.0002
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
AT4G04020 42 / 0.0003
FIB
fibrillin (.1)
AT3G23400 41 / 0.0004
FIB4
fibrillin 4, Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G020700
268 / 5e-90
AT3G26070 244 / 6e-81
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Potri.004G003200
54 / 2e-08
AT4G22240 380 / 1e-132
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Potri.008G169100
50 / 5e-07
AT3G23400 267 / 2e-89
fibrillin 4, Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Potri.001G137900
49 / 1e-06
AT2G35490 301 / 3e-100
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Potri.003G095900
47 / 5e-06
AT2G35490 334 / 5e-113
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Potri.001G011700
43 / 0.0001
AT1G51110 532 / 0.0
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020982
293 / 2e-100
AT3G26070 265 / 1e-89
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10001099
56 / 6e-09
AT4G22240 362 / 5e-126
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10014790
55 / 1e-08
AT4G22240 363 / 2e-126
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10020860
54 / 1e-08
AT5G09820 269 / 5e-91
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
Lus10033514
54 / 2e-08
AT5G09820 270 / 3e-91
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
Lus10026221
50 / 4e-07
AT5G19940 276 / 4e-94
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
Lus10030987
48 / 3e-06
AT2G35490 354 / 5e-121
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10010444
47 / 9e-06
AT1G51110 490 / 2e-173
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10035384
45 / 3e-05
AT2G35490 357 / 5e-122
Plastid-lipid associated protein PAP / fibrillin family protein (.1)
Lus10042448
44 / 6e-05
AT5G19940 250 / 9e-84
Plastid-lipid associated protein PAP / fibrillin family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04755
PAP_fibrillin
PAP_fibrillin
Representative CDS sequence
>Potri.001G209600.4 pacid=42793482 polypeptide=Potri.001G209600.4.p locus=Potri.001G209600 ID=Potri.001G209600.4.v4.1 annot-version=v4.1
ATGACAATGGCCTTATCTTCATCTCCACACTCTCCAGCAGTCCTCACAGCCTCTCAATTCTCAACTCACTCACCATTTCCAAAACTCACCACCTCTCACT
TCTTCTTCTCTACTGGTAAACCCTCCAACCAAACCAGCTACTTTAACCTTTCTTCAAGCTACTCCACCATTGATAGATCATGGAGCGCTAAGGTTTCTTT
CTTTCCTGCTTTCTTGAAAAAGGGCAAGAGTGCTAAGGTCCTCAAGGAGGAACTTCTTGAGGCCATTGATTCCCTTGATCGTGGAGCAGACGCCATTCCT
GAAGACCAACAAAGAGTTGATGAGATTGCTCGGAAGCTTGAAGCAGTGAATCCGACAAAGGAGCCATTGAAATCTGGTTTACTAAACGGGAAATGGGAGC
TTCTATACACCACTTCACAATCTATTTTGCAAACACAAAGGCCAAAGCTCTTAAGATCCAGGACAAACTACCAAGCAATCAATGCTGATATACTTAGGGC
CCAGAACATGGAATCTTGGCCATTCTTCAACCAGGTAACTGCAGATTTAACACCATTGAGTGCAAAAAAAGTTGCTGTAAAGTTCGATGTCTTCAAGATT
CTTGGTCTGATACCAGTTAAGGCACCAGGAAGAGCTCGTGGTGAGCTGGAAATCACTTATTTGGATGAAGAATTACGAGTATCCAGGGGTGACAAAGGAA
ACCTGTTTGTCTTGAAAATGGTTGATCCATCCTACCGTGTTCCTGTCTGA
AA sequence
>Potri.001G209600.4 pacid=42793482 polypeptide=Potri.001G209600.4.p locus=Potri.001G209600 ID=Potri.001G209600.4.v4.1 annot-version=v4.1
MTMALSSSPHSPAVLTASQFSTHSPFPKLTTSHFFFSTGKPSNQTSYFNLSSSYSTIDRSWSAKVSFFPAFLKKGKSAKVLKEELLEAIDSLDRGADAIP
EDQQRVDEIARKLEAVNPTKEPLKSGLLNGKWELLYTTSQSILQTQRPKLLRSRTNYQAINADILRAQNMESWPFFNQVTADLTPLSAKKVAVKFDVFKI
LGLIPVKAPGRARGELEITYLDEELRVSRGDKGNLFVLKMVDPSYRVPV
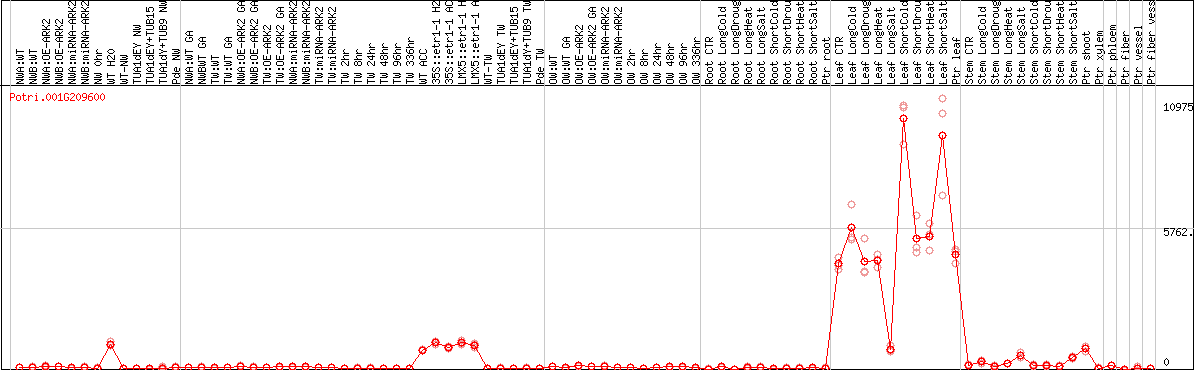
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G209600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.