External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G65430 66 / 1e-13
ATARI8, ARI8
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
AT1G05890 62 / 3e-12
ATARI5, ARI5
ARABIDOPSIS ARIADNE 5, ARIADNE 5, RING/U-box superfamily protein (.1.2)
AT2G31510 57 / 2e-10
ATARI7, ARI7
ARABIDOPSIS ARIADNE 7, ARIADNE 7, IBR domain-containing protein (.1)
AT2G31760 51 / 2e-08
ATARI10, ARI10
ARABIDOPSIS ARIADNE 10, ARIADNE 10, RING/U-box superfamily protein (.1)
AT2G31770 51 / 2e-08
ATARI9, ARI9
ARABIDOPSIS ARIADNE 9, ARIADNE 9, RING/U-box superfamily protein (.1)
AT2G31780 50 / 3e-08
ATARI11
ARABIDOPSIS ARIADNE 11, ARIADNE 11, RING/U-box superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G032000
71 / 2e-15
AT2G31510 834 / 0.0
ARABIDOPSIS ARIADNE 7, ARIADNE 7, IBR domain-containing protein (.1)
Potri.010G180600
71 / 3e-15
AT1G65430 860 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
Potri.008G077100
69 / 1e-14
AT1G65430 836 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
Potri.007G126900
66 / 2e-13
AT2G31510 836 / 0.0
ARABIDOPSIS ARIADNE 7, ARIADNE 7, IBR domain-containing protein (.1)
Potri.002G131200
62 / 2e-12
AT1G65430 571 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
Potri.002G131300
56 / 5e-10
AT1G65430 536 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025566
69 / 9e-15
AT2G31510 826 / 0.0
ARABIDOPSIS ARIADNE 7, ARIADNE 7, IBR domain-containing protein (.1)
Lus10026856
67 / 8e-14
AT1G65430 887 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
Lus10027027
66 / 2e-13
AT2G31510 807 / 0.0
ARABIDOPSIS ARIADNE 7, ARIADNE 7, IBR domain-containing protein (.1)
Lus10020229
64 / 7e-13
AT1G65430 833 / 0.0
ARABIDOPSIS ARIADNE 8, ARIADNE 8, IBR domain-containing protein (.1)
PFAM info
Representative CDS sequence
>Potri.001G214201.1 pacid=42789745 polypeptide=Potri.001G214201.1.p locus=Potri.001G214201 ID=Potri.001G214201.1.v4.1 annot-version=v4.1
ATGATACATCATATAATGGCTACGAGGAGGATGGATACTACGATGATGATGGTGATGGTGATTACGATGACTACAACAACTATATATGATGACAACAACA
ATGGTGATATGGATACTGGCTACGATGATGGTGGTGGTGAGGATTTCTTGGCATGTCGGAGCCAGCAAAGATGCAAGGAACCAGAGATCATTAGGCAACG
TCAGGAGGCTGATATCACAAGAATCTCAACTGTGCTCTCTATATCGAGAAACGAAGCAAGCCTCCTACTTCGTCTCTATGGTTGGAATGTTATCAAAGTA
GAGGATGAATGGTTTGGTAATGAAGAAGAGGTGCGTAACATTTCATATTAG
AA sequence
>Potri.001G214201.1 pacid=42789745 polypeptide=Potri.001G214201.1.p locus=Potri.001G214201 ID=Potri.001G214201.1.v4.1 annot-version=v4.1
MIHHIMATRRMDTTMMMVMVITMTTTTIYDDNNNGDMDTGYDDGGGEDFLACRSQQRCKEPEIIRQRQEADITRISTVLSISRNEASLLLRLYGWNVIKV
EDEWFGNEEEVRNISY
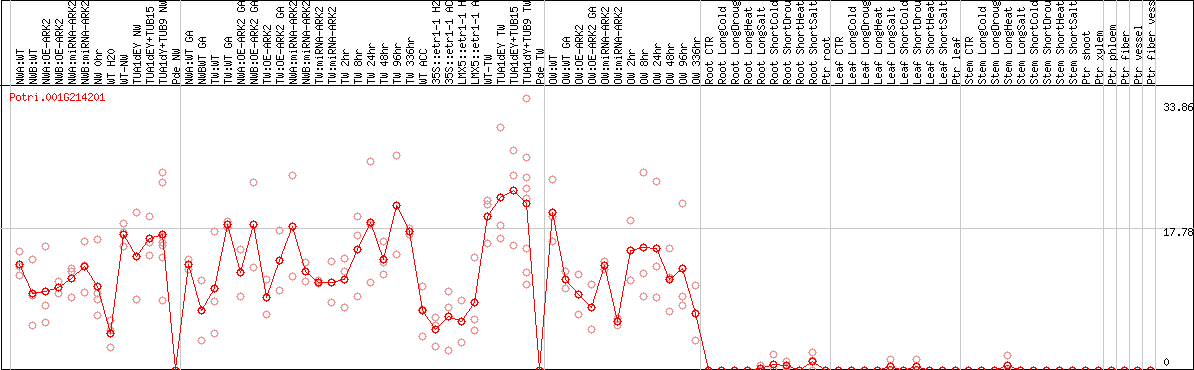
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G214201 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.