Potri.001G233800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G233800.8 pacid=42792956 polypeptide=Potri.001G233800.8.p locus=Potri.001G233800 ID=Potri.001G233800.8.v4.1 annot-version=v4.1
ATGGCGATTGAGAAGAACAATTTTAAGGTATCTAATAGGTTTGACGCGGAGCTTTCTCCCAATAGTAGGGATACTAGCATGTCCAGCGACGAGGATGAGG
ATGACCTGCTTCATCACCAGCGGATTAAGTCTGATGATGACGAGGAGGAGGTTGAGGATGCTGTTGATGTAGGTGTTGAAGAGGATGATGATGATGAATT
CGACGATGCTGATTCGGGGGCGGGATCTGATGATTTTGACTTGTTAGAATTAGGGGAAACCGGGGCGGAGTTTTGCCAATTTGGGAATTTAACCTGCAGT
GTTCCTTTTGAATTGTATGACCTTCCTGGCTTGGAGGATATTTTGTCTGTTGATGTGTGGAATGATGTTTTGACTGAAGATGATAAATTTAGCCTTACTA
AGTATTTACCTGATGTGGATCAGGACACCTTTATGCGAACCCTTAAAGAGCTTCTCGAGGGTGGGAATTTCCATTTTGGTAGTCCCCTCAACAAGTTGTT
TCAGATGCTTAAGGGAGGGTTGTGTGAGCCAAGGGTTGCCCTTTATAGGGATGGTTTGAATTCCTTTCAGCAACGGCAGCATTACCATATTTTGAGGAAG
CATCAGAACAGTATGGTTAGTCATTTGTGCCAGATAAGAGATGCTTGGCTTGATTGTAAAGGGTATAGCATCGATGAGAAACTTAGGGTTTGGAATATTA
TGAAGAGTCATAAGAGTTTGATGTATGAGAATGTGGAGGGAGAATTGGAGAGTGGCTCATCGGATAAAGGAGAATCAGGTGATGGGTTTTGGGGAAAAAG
GGTTAAAGATAAGAAGAGTGCATCCAAATTTGATCGTAATTCAGCATACCAGGTTGGTTCTAATTTGGAATTTTCTTCACCAGTCAGTTTGGAAGTGGTA
AAGTATGGAAAGCAAAATCCCAAGAGTATACTTAAGTCGGCCGGGTCAAAAGATCTTTCAACTAGAGATGTGCTGGGTCGAATTCCTTCTGATCATCATG
GTTTGGGGATGACTTCAAGGCCACGTCGATCAGCCTTGATGGTTTCTCGGCAGAATAAGCTAGCTGGGTATGACTCAGGAGATGCACTCAGGCTAAGGGA
TCAAACGAGAACTGATAATGATGATGCTGAATATGCAATGTATGGAATGGGTGTCCAAAGAGATCGAAATATGACACGTGGTGGTGATATGGTGAAATCC
AGGGTTCCAAAAGTGGGAAAGAAGCATGAGTTCTTGAGAAGTGATGGATTAGCTGCTGACAGTTTCATGGATTTGCCTTTCTCTTCAAACAATGAATTAC
TTGCCTATGGTAGGAACAAGAATGCAAATCAGCTGTCAGAAGCTAAAGTGTTCGCGTCAAATCGTAGCAATACTCGGACCAAGTCTGAATCTTCCAAAAA
GACCAAGTATGCTGAAAATTTTTCGCAATTCACAGTTCCTGATCAGATGAAGTATTTGAAAGGCCGGACTCTGCAGCTACCCCGTAAAGGTAATCGAGTT
GAATTGTCAGATCATGCCGAACCAGTTTGGCATAGCAAGAACCAAGGAGAAGTTTTCTCTATGGATTCTACATTTAAAATTAATGACTGGAATATGAGAG
GCAAGAAATGGAGGACAGAGAGAGAGTCTCCGGATCTTAATTTTAGAGCTTATAGAGCATCCTCGCCCCAGGTGAATGATAGAATGGTGCTGTCTGAGGT
GAAAGCAAAATCATCACGGGAGAAGATCAGAGGGAATGTTATACAGAATGGAGGACCAGATAAGGGAGCATTGAAAGGTAACAGAATTTATGTAAAAGGT
GAAGAAACAGAAACGGACTCTTCTGAACAATTTGAGGAGGAGGAGCAGGAGGACGAGGAGGAGGAGGAGGAGGAGGAGGAGGATAGCAATCCTTTGATGA
GAAGCAAGTCAGCTTACCCCATTGGTATCTCAGAAGGTTATCGATCATCCTTTTTGAAGTCTCGCTTAGATGCTAAAAAGGCCTCATCTATTAAGAAAGA
TACGCTGGAGAATGAACTAGCTTTCGACGGTGTCACTCAGTTCTCCAAAAAGGTGGGTGGCTTTACCGAGTCTGGACAAATGCCTGGATACTCATCAAAA
GCTAAACAGAAGGGTAAGATGCAAGAGACGCGTAGCTCTAGTGCTAGAGTTTTGGAAGACAGCTCTCCCATTGGCTTGGCTAAACTAAAAGATGATAATG
ACAGAAACCGGGTACACAGATTTGGCAAGATTGGTCAATTGCGTGTGGAATCTGGTGAAAGGTCACGCAGGACCTCGTCAAAAGCCCACCCTTCTGACAG
AAAGCATAAAGGTGAAGTTAGCCATGAATTTATTGTTGATGACGAGGATGAGTTACTTGAAACACAGTTAACGTCAGATGAAAATGCACTGGGTAGATTC
CGGAAGAAAGGACAGAGCATGGAAACATATGTTCATGGTCAAAGTGACCGATCTGAGGCTTCGTTGCTAGCGTGCAACTCGGTGACGAAGAAACGGAAAG
CAAAGTACAAAGTGATGGACATGGCTGGAAGAGATGAAGATAGTAATAGGCAGTCTAGTAGTGCTCAGCAGCAAATTGATGACTCCATTTCATTGAAGAA
AAAAGGGAAAAGGAAATTGGAGGCTGACGATGTTACTCCAGACAGGGAAACTCCTGAGGCACACATTCCAAAAACAGGAGTGGTGGATGTGGAGCTGGAA
GCCAAACCTCAGAAAAAACCGTATATTCCGATTACACCAACAGTTCATTCTGGCTTCTCCTTTTCCATTATACATCTCCTCTCAGCAGTTCGTGTGGCAA
TGATCACTCCTCTTTCAGAAGATTCCTTAGAGGTTGGCAAGGCTACAGCAGAGCTGAACAGAGCGCAAGAAGGTGACACCAATGGGGTTCTTTCTAATGA
GAATGTAGATGTTAATAAATCACACCCTGCTGTGCAAGTGAAAATGCCCTCTCTTACTGTACAGGAGATTGTTAACCGTGTGAGATCAAACCCTATGGAT
CCATGTATACTCGAGACACAGGAGCCACTCCAGGATTTGGTCAGAGGAGTTCTTAAGATATTTTCATCGAAAACAGCACCTTTAGGAATAAAGGGTTGGA
AAGCACTTGTTTTTTATGACAAATCTACAAAAAGTTGGTCCTGGATTGGCCCAATTTCTCATGCTTTGACTGATGAGGACACTATCGTTGAAGTGACATC
TCCTGAATATTGGGGCCTTCCTCACAAAAGTTGTGTCAAGTTGGTCGACTCATTTGCTAATTGGCTGAAAAGCGGGCAAGAGACTCTTCAACAAATTGGT
AGTCTTCCCGCACCGCCTGTGTCATTGATGCAATGTAACCTGGATGAGAAAGAAAGGTTCAGGGACTTGAGAGCTCAGAAAAGTCTCAACACTATTAGCC
CGAGTTCTGAAGAAGTTAGAGCTTATTTTCGCAGGGAAGAAGTGCTGAGGTATTCAATTCCTGACAGGGCCTTCTCATACACGGCTGCTGATGGTAAAAA
ATCTATTGTTGCTCCCTTGAGGAGGTGTGGTGGTAAGCCAACTTCAAAAGCTCGAGACCATTTCATGCTGAAGCGTGATCGACCACCTCATGTTACAATT
CTTTGCCTTGTGAGAGATGCAGCTGCTAGATTGCCAGGTAGTATTGGCACTAGAGCAGATGTTTGTACTTTGATACGGGACTCTCAATATATTGTTGAAG
ACGTGTCTGATGCCCAAGTGAATCAAGTTGTCAGTGGAGCCTTGGATCGTTTGCATTATGAGCGCGATCCTTGTGTGCAATTTGATGGAGAAAGGAAATT
GTGGGTTTACTTGCACAGAGATCGAGAAGAAGAAGATTTTGAAGATGATGGAACTTCATCTACTAAGAAATGGAAAAGACAGAAAAAAGATCCTGCTGAT
CAATCTGATCAAGGAACAGTTACTGTTGCTTTTCATGGGACAGGGGATCAATCCGGATTTGATTTGGGCTCTGACCTGAATGCTGAGCCACTAGCCGCTG
ATGATGATAAAAGAACAGATCTTGTATGTAGTGATGTGAGGCATAACGCAGAGGATAATATTGATACCTCTCATGGGCCCAAGCAAGGCAGCACATATGA
CGGTGATGCAATGGTTTGGGATGCCCTTAGCTTAAATCCTTTGCAAGAGAACAAAGTGATATGTCAAGAAAATTCCACAAATGAAGATTTTGATGATGAA
ACATTTGAGAGAGAAAGGCCGGCTGGACTTTTAAGTACAAGCTTATTATGA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G233800.8 pacid=42792956 polypeptide=Potri.001G233800.8.p locus=Potri.001G233800 ID=Potri.001G233800.8.v4.1 annot-version=v4.1
MAIEKNNFKVSNRFDAELSPNSRDTSMSSDEDEDDLLHHQRIKSDDDEEEVEDAVDVGVEEDDDDEFDDADSGAGSDDFDLLELGETGAEFCQFGNLTCS
VPFELYDLPGLEDILSVDVWNDVLTEDDKFSLTKYLPDVDQDTFMRTLKELLEGGNFHFGSPLNKLFQMLKGGLCEPRVALYRDGLNSFQQRQHYHILRK
HQNSMVSHLCQIRDAWLDCKGYSIDEKLRVWNIMKSHKSLMYENVEGELESGSSDKGESGDGFWGKRVKDKKSASKFDRNSAYQVGSNLEFSSPVSLEVV
KYGKQNPKSILKSAGSKDLSTRDVLGRIPSDHHGLGMTSRPRRSALMVSRQNKLAGYDSGDALRLRDQTRTDNDDAEYAMYGMGVQRDRNMTRGGDMVKS
RVPKVGKKHEFLRSDGLAADSFMDLPFSSNNELLAYGRNKNANQLSEAKVFASNRSNTRTKSESSKKTKYAENFSQFTVPDQMKYLKGRTLQLPRKGNRV
ELSDHAEPVWHSKNQGEVFSMDSTFKINDWNMRGKKWRTERESPDLNFRAYRASSPQVNDRMVLSEVKAKSSREKIRGNVIQNGGPDKGALKGNRIYVKG
EETETDSSEQFEEEEQEDEEEEEEEEEDSNPLMRSKSAYPIGISEGYRSSFLKSRLDAKKASSIKKDTLENELAFDGVTQFSKKVGGFTESGQMPGYSSK
AKQKGKMQETRSSSARVLEDSSPIGLAKLKDDNDRNRVHRFGKIGQLRVESGERSRRTSSKAHPSDRKHKGEVSHEFIVDDEDELLETQLTSDENALGRF
RKKGQSMETYVHGQSDRSEASLLACNSVTKKRKAKYKVMDMAGRDEDSNRQSSSAQQQIDDSISLKKKGKRKLEADDVTPDRETPEAHIPKTGVVDVELE
AKPQKKPYIPITPTVHSGFSFSIIHLLSAVRVAMITPLSEDSLEVGKATAELNRAQEGDTNGVLSNENVDVNKSHPAVQVKMPSLTVQEIVNRVRSNPMD
PCILETQEPLQDLVRGVLKIFSSKTAPLGIKGWKALVFYDKSTKSWSWIGPISHALTDEDTIVEVTSPEYWGLPHKSCVKLVDSFANWLKSGQETLQQIG
SLPAPPVSLMQCNLDEKERFRDLRAQKSLNTISPSSEEVRAYFRREEVLRYSIPDRAFSYTAADGKKSIVAPLRRCGGKPTSKARDHFMLKRDRPPHVTI
LCLVRDAAARLPGSIGTRADVCTLIRDSQYIVEDVSDAQVNQVVSGALDRLHYERDPCVQFDGERKLWVYLHRDREEEDFEDDGTSSTKKWKRQKKDPAD
QSDQGTVTVAFHGTGDQSGFDLGSDLNAEPLAADDDKRTDLVCSDVRHNAEDNIDTSHGPKQGSTYDGDAMVWDALSLNPLQENKVICQENSTNEDFDDE
TFERERPAGLLSTSLL
|
DESeq2's median of ratios [POPLAR]
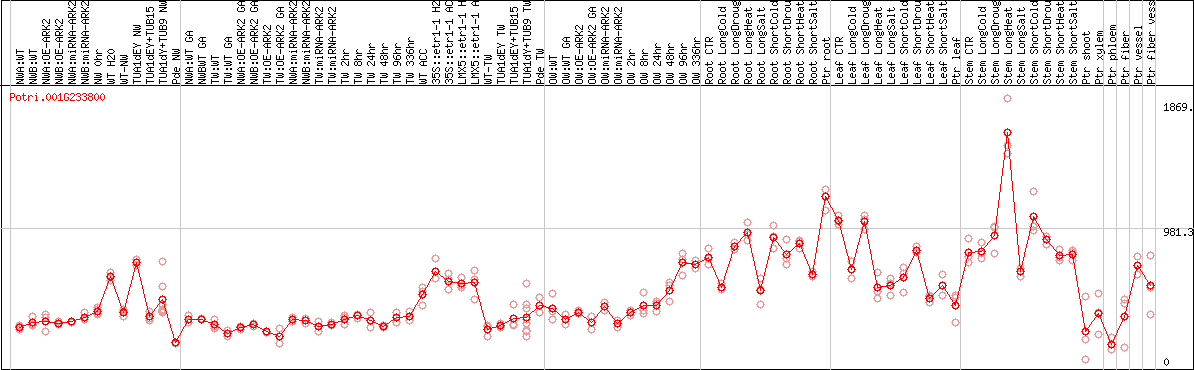
Coexpressed genes
Potri.001G233800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
