External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G13850 88 / 2e-23
ATGRP2, GR-RBP2
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
AT3G23830 86 / 6e-23
AtGRP4, GR-RBP4, GRP4
glycine-rich RNA-binding protein 4 (.1.2)
AT5G47320 79 / 4e-19
RPS19
ribosomal protein S19 (.1)
AT5G61030 80 / 5e-19
GR-RBP3
glycine-rich RNA-binding protein 3 (.1)
AT1G74230 79 / 5e-19
GR-RBP5
glycine-rich RNA-binding protein 5 (.1)
AT5G06210 73 / 1e-17
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G21660 69 / 4e-16
CCR2, ATGRP7, GR-RBP7
GLYCINE-RICH RNA-BINDING PROTEIN 7, "cold, circadian rhythm, and rna binding 2", GLYCINE RICH PROTEIN 7, cold, circadian rhythm, and rna binding 2 (.1.2)
AT1G18630 69 / 4e-16
GR-RBP6
glycine-rich RNA-binding protein 6 (.1)
AT3G08000 69 / 6e-16
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G39260 67 / 6e-16
ATGRP8, CCR1, GR-RBP8
glycine-rich RNA-binding protein 8, GLYCINE-RICH PROTEIN 8, cold, circadian rhythm, and RNA binding 1 (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G059000
85 / 1e-22
AT4G13850 134 / 9e-42
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Potri.012G061600
83 / 2e-20
AT5G61030 181 / 9e-56
glycine-rich RNA-binding protein 3 (.1)
Potri.015G057400
82 / 2e-20
AT5G61030 157 / 8e-47
glycine-rich RNA-binding protein 3 (.1)
Potri.001G319900
80 / 2e-20
AT4G13850 140 / 1e-43
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Potri.006G208500
77 / 3e-19
AT5G06210 160 / 1e-51
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.009G116400
76 / 2e-18
AT2G21660 145 / 5e-45
GLYCINE-RICH RNA-BINDING PROTEIN 7, "cold, circadian rhythm, and rna binding 2", GLYCINE RICH PROTEIN 7, cold, circadian rhythm, and rna binding 2 (.1.2)
Potri.001G319800
74 / 5e-18
AT4G13850 137 / 2e-42
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Potri.004G155300
74 / 6e-18
AT2G21660 145 / 3e-45
GLYCINE-RICH RNA-BINDING PROTEIN 7, "cold, circadian rhythm, and rna binding 2", GLYCINE RICH PROTEIN 7, cold, circadian rhythm, and rna binding 2 (.1.2)
Potri.008G022280
72 / 4e-17
AT3G20930 203 / 2e-65
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022551
90 / 4e-24
AT4G13850 167 / 7e-54
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Lus10016639
89 / 1e-23
AT4G13850 166 / 1e-53
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Lus10017852
91 / 2e-23
AT5G61030 178 / 7e-55
glycine-rich RNA-binding protein 3 (.1)
Lus10034685
90 / 4e-23
AT5G61030 177 / 7e-54
glycine-rich RNA-binding protein 3 (.1)
Lus10043158
87 / 8e-23
AT4G13850 154 / 1e-48
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Lus10032591
86 / 1e-22
AT4G13850 157 / 5e-50
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Lus10021154
86 / 2e-21
AT5G61030 188 / 5e-58
glycine-rich RNA-binding protein 3 (.1)
Lus10040519
83 / 8e-20
AT5G61030 189 / 2e-53
glycine-rich RNA-binding protein 3 (.1)
Lus10023569
75 / 3e-18
AT2G21660 147 / 8e-46
GLYCINE-RICH RNA-BINDING PROTEIN 7, "cold, circadian rhythm, and rna binding 2", GLYCINE RICH PROTEIN 7, cold, circadian rhythm, and rna binding 2 (.1.2)
Lus10025573
74 / 1e-16
AT3G20930 336 / 3e-109
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.001G236300.1 pacid=42793365 polypeptide=Potri.001G236300.1.p locus=Potri.001G236300 ID=Potri.001G236300.1.v4.1 annot-version=v4.1
ATGCAGTGTCTGAATCGAAGTGTCAGTCTGCCCTTAGCCAGACGGTTATTGGCCCGCCACTTATCCACCAAATTATTTGTTGGAGGGCTTTCTTATGACA
CCAATGAAACTGTTTTAAAGGATGCTTTTGGAAAATATGGTGAAATAATTGAAGCAGTGAAAGTGATATGTCACCGAGTGTATGGTGAATCCAAAGGCTA
CGGCTTTGTGCAGTTCTCTTCTGAAGCTGCAGCCAGCACAGCTTTGAAAGAAATGGATGGCCAGTTGCTGGATGACCGAAATATCCGTGTGCATTATGCT
CATAAGTGGTCATGA
AA sequence
>Potri.001G236300.1 pacid=42793365 polypeptide=Potri.001G236300.1.p locus=Potri.001G236300 ID=Potri.001G236300.1.v4.1 annot-version=v4.1
MQCLNRSVSLPLARRLLARHLSTKLFVGGLSYDTNETVLKDAFGKYGEIIEAVKVICHRVYGESKGYGFVQFSSEAAASTALKEMDGQLLDDRNIRVHYA
HKWS
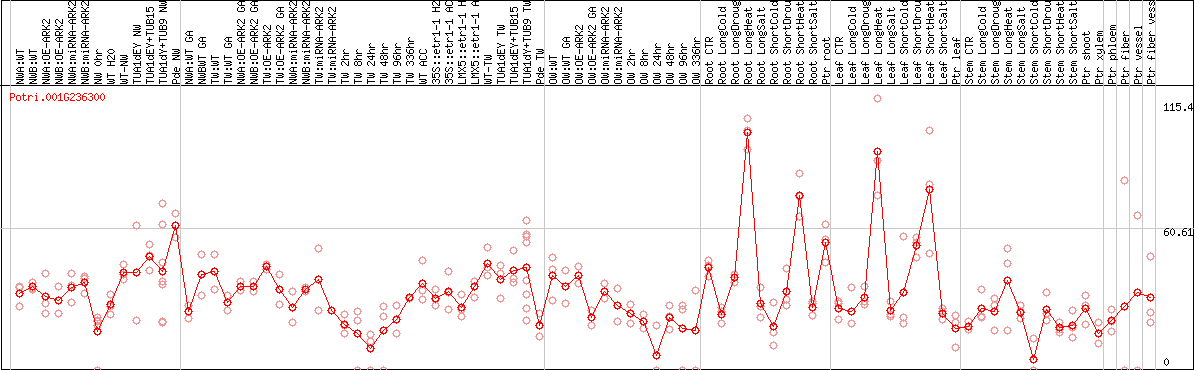
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G236300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.