External link
Symbol
Pt-UCP2.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G58970 456 / 3e-163
ATUCP2
uncoupling protein 2 (.1.2)
AT3G54110 413 / 3e-146
ATUCP1, UCP1, UCP2, UCP, ATPUMP1
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
AT1G14140 184 / 3e-56
Mitochondrial substrate carrier family protein (.1)
AT2G22500 183 / 5e-56
UCP5, ATPUMP5, DIC1
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
AT4G24570 178 / 6e-54
DIC2
dicarboxylate carrier 2 (.1)
AT5G09470 161 / 4e-47
DIC3
dicarboxylate carrier 3 (.1)
AT4G03115 150 / 2e-43
Mitochondrial substrate carrier family protein (.1)
AT5G19760 129 / 2e-35
Mitochondrial substrate carrier family protein (.1)
AT5G61810 102 / 7e-25
APC1
ATP/phosphate carrier 1, Mitochondrial substrate carrier family protein (.1.2)
AT3G21390 101 / 1e-24
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G110100
451 / 6e-161
AT3G54110 483 / 5e-174
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.006G095500
444 / 3e-158
AT3G54110 513 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.017G045100
190 / 3e-58
AT2G22500 451 / 6e-161
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045150
188 / 9e-58
AT2G22500 457 / 3e-163
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G047800
187 / 2e-57
AT4G24570 435 / 2e-154
dicarboxylate carrier 2 (.1)
Potri.007G113000
185 / 2e-56
AT4G24570 404 / 4e-142
dicarboxylate carrier 2 (.1)
Potri.005G157300
177 / 2e-53
AT2G22500 412 / 2e-145
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.002G104400
177 / 2e-53
AT2G22500 427 / 1e-151
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.010G165900
173 / 3e-52
AT1G14140 447 / 1e-159
Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040709
471 / 5e-169
AT5G58970 512 / 0.0
uncoupling protein 2 (.1.2)
Lus10016445
452 / 2e-161
AT5G58970 493 / 1e-177
uncoupling protein 2 (.1.2)
Lus10027755
422 / 1e-149
AT3G54110 515 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10035538
419 / 1e-148
AT3G54110 512 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10021123
418 / 4e-148
AT3G54110 521 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10017187
396 / 2e-139
AT3G54110 501 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10030443
181 / 3e-55
AT1G14140 421 / 3e-149
Mitochondrial substrate carrier family protein (.1)
Lus10024769
145 / 4e-41
AT4G03115 401 / 3e-141
Mitochondrial substrate carrier family protein (.1)
Lus10030361
129 / 2e-35
AT5G19760 489 / 9e-177
Mitochondrial substrate carrier family protein (.1)
Lus10007883
132 / 4e-35
AT5G19760 536 / 0.0
Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.001G247800.1 pacid=42788365 polypeptide=Potri.001G247800.1.p locus=Potri.001G247800 ID=Potri.001G247800.1.v4.1 annot-version=v4.1
ATGGCTGATCTCAAGCCCAGTTCTGATATTTCCTTCGTCGAAATCTTCCTGTGCAGTGCTTTCGCTGCTTGTTTTGCTGAGTTCTGCACCATCCCTTTGG
ACACTGCTAAAGTTAGGCTCCAGCTACAACGGAAAACATTTGCCTCGGAAGGAGTAAGTTTGCCTAAATATAGGGGTTTGTTGGGTACTGTGGCTACTAT
TGCCAGAGAAGAGGGTTTAGCAGCACTATGGAAAGGCATTACTGCAGGATTACATCGCCAGTTTATTTACGGTGGCTTAAGGATAGGATTATATGAACCG
GTTAAATCTTTTTTGGTTGGCAGTGATTTTGTTGGGGACATTCCTTTGTACCAGAAAATACTTGCAGCTTTGTTAACTGGTGCTATGGCAATTGTAATTG
CTAATCCAACTGATCTTGTTAAAGTTCGGCTTCAAGCTGAAGGGAAACTGCCAGCTGGGGTGCCTGGGCGATATGCTGGAGCTTTGGATGCTTATTTTAC
CATAGTGAGACAGGAAGGACTGGGGGCCTTGTGGACTGGACTTGGGCCAAATATAGCAAGGAATGCTATTATAAATGCTGCTGAACTTGCCAGTTATGAT
GAAGTGAAACAGACAATATTGCAAATTCCGGGGTTCACGGACAGCGCCTTTACTCATGTTCTAGCTGGTCTCGGTGCAGGTTTTTTTGCAGTGTGTATTG
GTTCTCCCATCGATGTAGTAAAATCTAGAATGATGGGAGATTCGTCCTACAAAAATACTGTTGACTGTTTTATCAAGACACTGAAGAATGAGGGAATTTT
GGCGTTTTACAAAGGTTTCCTCCCAAATTTTGGGCGGCTGGGATCTTGGAATGTTGTAATGTTTTTGACCCTTGAGCAAGTCAAGAAAATAGTCACAGGG
CAAGCATACTATGACTAA
AA sequence
>Potri.001G247800.1 pacid=42788365 polypeptide=Potri.001G247800.1.p locus=Potri.001G247800 ID=Potri.001G247800.1.v4.1 annot-version=v4.1
MADLKPSSDISFVEIFLCSAFAACFAEFCTIPLDTAKVRLQLQRKTFASEGVSLPKYRGLLGTVATIAREEGLAALWKGITAGLHRQFIYGGLRIGLYEP
VKSFLVGSDFVGDIPLYQKILAALLTGAMAIVIANPTDLVKVRLQAEGKLPAGVPGRYAGALDAYFTIVRQEGLGALWTGLGPNIARNAIINAAELASYD
EVKQTILQIPGFTDSAFTHVLAGLGAGFFAVCIGSPIDVVKSRMMGDSSYKNTVDCFIKTLKNEGILAFYKGFLPNFGRLGSWNVVMFLTLEQVKKIVTG
QAYYD
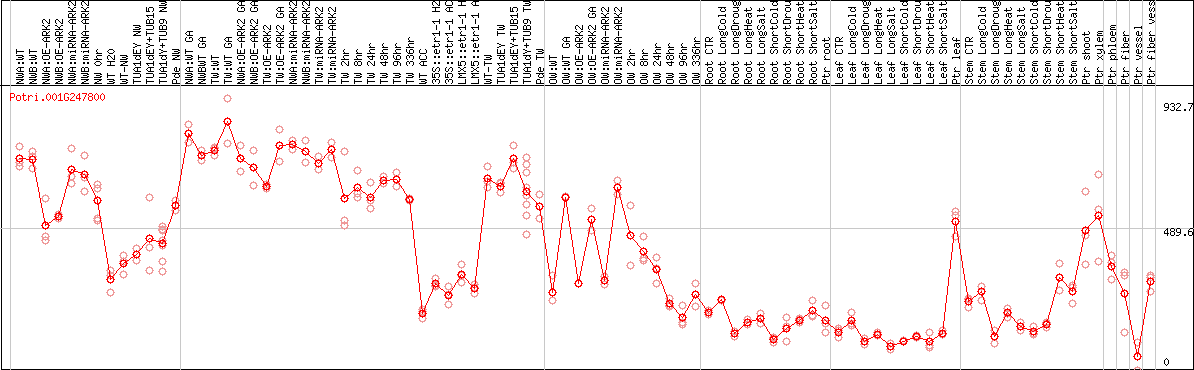
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G247800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.