Potri.001G249000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G249000.1 pacid=42791844 polypeptide=Potri.001G249000.1.p locus=Potri.001G249000 ID=Potri.001G249000.1.v4.1 annot-version=v4.1
ATGAAAATGGGCATTGATGCAAAGGAATTTCAAGCTTTCATGTGGAGAGTTGTTAAATTTTCCATAAATACTTGTACTGCATTGGCACAAAAATATCCCT
TTGCTTCAGGTGTTTTGTTATCCTTGCTGATTTTATACCTGTTTCTTCCTTCAGTATTCTTTTTCTTGATTTACTCTCTACCATTTCTTGGTTGCACTGC
TGTTTTTATCCATTATTATTTGAATACACAACGTCCCAAAATCCAACACGGGGATGAGAGGAAGGAACATGGGATCTCTTCTATTGAATCTCGGAGACTG
TTGCAGAGAAATGTCAACAAGAACAATATTGACGAATCGGATGCACATGCTGTTAAAGAGGAAAAGGATATGGTTTCCCCCATGATTTCAAATGATGAAT
TGATTGGTAGAACTGCCTTGGCCGAAGAAAAACCAAAAATAATCATGGAGGAGAAAGAAAGCCGTGCTTTGAATAGCGGAGAAAGTTCCTCTCACAATGT
GTCCATCGGTGAGAATATTTCAGAACTCGGCCAGGCGCCAAATCCTGATGCTGTCTCATGTGATGGCTTTAATGAGCAGCCTACAAAACTTCAAGTTGGT
GGTGAAGTGGAGTTGGAGAGCTCAAGCTCCGAAGCAGATGATGATGATGAAGAAGAAGAATCGGAAAAGGGTGGGGAAAACGCTGTACAATGGACCGAGT
ATGATCAAAAGAATGTCATGGATCTTGGGAATTCCGAGATAGAAAGAAATAGGAGGCTGGAGATGCTAATTGCAAGACGTAAAGCAAGAAAGTCGTTCAA
AATGAATAGTATTGCCGGTTCAGGTCCAAGGCATCCTGTTATGGTTGCCAGAAGCAACACTTTCCACGTTTCTAAGAGTTCAGATGACCAAATACCCGGT
TCAGCCCCTTCCATTCTGTTACCAACCAAAAATCCATTTGACCTTCCATACGACCCACATGAAGAAAAGCCTAATCTAATGGCTGACAGTTTCCACGAAG
AATTTATGGCAGATCACCAGAAAGAATTTCCCTTCTGCAGGCATGAGAGCTTCTCTTTGGGAAATTCTTTCCAGGATAACAGGCAAGGTCAACACGAGGG
CCGAGGATCTTCTAGACCTAAAATGCAATCAGGCAATAAGGGAAATCATGATCGGCTGGTTGATCATTTATTATTTCAGGGTGGTGAAACCTTGCGCCGT
AACTTAAGTGTTACTGATCTTGTGACAGAAGAACCCCAATCTTCCAACCAGGTAGCTAACAAACAGGAACGGGATAGAGAAGTTGGAACCACCAGGATCA
AACTCATAGGTGAGAAGATGGAGCAATCTCATTACAAGGATCCAAGTTTAGGTAATGGAAGTGATATTCAAATGAACAAAGATGCGAATGCGATCAAGCA
CAAGAAAATTTTGTCAAACCCATCTTCTTCTGCAGAAGACATTCTTAATGCTAAGACAGCTGAAAATTCAGAAAGTATACAACCGACAACGTTCAAATTT
CCCGAGGTGTTTTATGATCGGGCACCGAACTCTCTCCCTTGTCCAGTTCCTAAGGCTAGAGCTGTAGCTGAACCTTCGTATGATTCCAGCCCTTCTCCAA
TTGACAACACGAGAATGGAGGAACATTTCTTTTACAAGCTCAAGCCTGGGCATACCCCTAACCACTCCATAGCTTCTGACATGCAAGTGGAGGTCTCAGA
AATTGGTTCACCCCCTCTGATTGAAGATGGAACTGCTTCATCCAATGATGACGAGTCCTTGATTTATGACGGGGATAGCGAGAACGAATTCACTTCTGGG
AGTGAAGAACTGTGGGGTTCTTCGCCACTTGCACCAAAAGTTCAAGAACATGGAAAGGCACGTGGACAAATATACGAGGAAGGGGAAGAGGGTATGACAG
AAGTAGAGTTCTCAAGGGTCTGGGACGAACCCGAGAATCCAATTTCATCATCAATGTGGCCGTTATCATCTTCAAGAGCTGAAATATCTCAAGAGGATCA
AGCCCATTCAATGAAAATTGATCCTAAACTATCTAATCATGTCAAGGACGAAGTTGATGAAGTAAGGGAGCAGAGGCCTTCTAATGCTTCTGATGTGGAC
GAAGTTGAAGAAGTAAGGGAGGAGAGGCCTTCTAATTCTTCTGATGTAGTTCCACCTGAACATTCCCTGGAAGGAACGCGATTGATGGAGGGTTCAATGG
CTCACTCGCCTTCTGAAGTGTACTTTCCAGAACCGCAGGAATCACCGAGTGGTTCTGGGAATTCTGCAGAAGAGAAGAAAACGAATTGCGATGCAAATGA
AACAGTCACATATGATGATAGGGAATATTTGAAGTCAAATGAAAACAGACATGTTGGAGCTGAAAACTCCATTATGCAAGAAGTTTTAGGCAATCTATCA
GAACCAGCTGAGGGAAATAATTCAACATCTAATAGCAATATCAAGATCGAATCATTGTTTAACTCTGAGAAGTATGTAGAGGGCATAGGTGACGAAACTT
ACGATGTAAATGATTCAACGTTCCTCATAAGTGATCGCTTAGAAGACTCGAAATCAAATGAAGAAAGAGATAGTGGAGCTGAAAGAGTCATGCAAGCAGT
CATAGGTGATTTGTCACAACCAGCTGTGGAAAGTAATTCAGAATCTAGCAATCATATAGAGAGTAAATCATTGAATTCTCCTGAGAAGTCTAGAGAGGAG
GCAAACATCAACAATGTGAATGACCCACCAGTTCAAATTAAGGATCGTGTTGAAGACTTCAAAGATGTCGACCGAGATTCTGAAAGGCTGACAGAAGATT
CAGACATTCAGAGCATTTTAATGCCAGTCGAAGCTGAAGACAACCCGACATCAACTCAAGGCAGTAAAGAAGATCAAAGTACAATAGAAGTTGGAGTTTC
TGTGGTAAACCAACCTTTTACTGATCCAACTACATCAGCCACCCTGCCAGAATTTGTGGCTGAACAAGTTTCTAACAATTCAAGTTTATCCTCGTCTCCA
AAGTCTGTTTTAGCATACAGGATCCCAGCAGATATAGGTTCTTCATCAGATTTCAGTCAACTGGTGGCCACTGATATGGAGGAAAATTTACTTATGACCG
CAACTCAGGATACCAGTCTTGCGGTAAATGATTCAATAGATCACCCATCTATTGATAGAAAATCTGAAAAATCTGAGGAACCATATGACACTCAAGGGAA
GTGTACTGAAGAAGCCATTGACATGGAGAACATGAATGGATCAGTGCTTGATGATGAGCAAATCAAGGAGAATTTAAAATGTAGGAAAAACATAGAATAC
GAATCAGAAACATTGATCAGTAACGAGGCTTCTGTAGAACTGTCAGAACCACTTGAAGAACCCCAAACTGCTGATCATCTTGAAGGTGCATCTGCAAGAC
TTGTGGACAATGAAGCAAGCATTAACCTATCAAAACCAGATGAGGAGCATGTCAGTTCAAAAGTTCCAGGAGTTTTAGTAGAAAAGGAGGAGTCAACAAA
TCCTCCAAGAGATGTTGCAGGAGAAGTTAATCACATCAGTGATGTTAGTGATCCATCAATAAATAAGAACGACGATTTGGAGAAGTTAAAGTCTTTTGAA
GGCAGTGAAGGTGAACCTGATCAATTTTCAACTGGACATGAGATATTTGTGGAACCACTAAAACCAGCCAATATTACTTCCCTAGAAGGCCATGAGTACA
GTCCTGGAGTATTAACTGAAAATGAAACAATTGTCGCATCATCTCAAGCAATCGAGGAAGTTGATAATTCAAGAACCAGCAAGGAAACTGATGAGTTTGG
AACACAAATAGCAGACGAGGAAATTGAAGACTTGTTAAAGCCAGGAGAGGCTGTTGTCTCCTCAGAAACCACTAAGAATGTCCAAGGTGACCCTAAAGAT
TTAATTGATCAAAAGGCTGTCCTTAACCCATCAACACCAGCTGTTGATGACGACAACATCTTAGTGACTCCTGAAGCAAAGGACTCTGCAGCTGACACTA
TTCACAATGTGAATGAATCAGAGATGAGTGAATTTATCAGCAACGAGAAGTTCAAGCATGTACAAGACAGTGAAGACGAATCTCAAAGATTAGATAGACA
AGAGGATATCATGGAACCACTTAAAGCAGTGGAGGTTACAAACTCAGAATCCATCAGAGATATTGAAGGTGAATCAAAAAAATTGGCCGACGATGAAGTT
AATGTCACCATCCCATCGCAGCCAGAAGGGGAAATTAATAGTTCAGATGACAGAGAAAAAACCGAGGATCCTGGAAAGTCAATTGTTCAGGAGAATGGTA
TGGACATATCCGAGGTAAGCAGGGGTAACGATATTGCAAGGGCCGTTGAAGATAACAAAGATAAATCTGAAGACAAAACTGAGATCAATGGATAG
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G249000.1 pacid=42791844 polypeptide=Potri.001G249000.1.p locus=Potri.001G249000 ID=Potri.001G249000.1.v4.1 annot-version=v4.1
MKMGIDAKEFQAFMWRVVKFSINTCTALAQKYPFASGVLLSLLILYLFLPSVFFFLIYSLPFLGCTAVFIHYYLNTQRPKIQHGDERKEHGISSIESRRL
LQRNVNKNNIDESDAHAVKEEKDMVSPMISNDELIGRTALAEEKPKIIMEEKESRALNSGESSSHNVSIGENISELGQAPNPDAVSCDGFNEQPTKLQVG
GEVELESSSSEADDDDEEEESEKGGENAVQWTEYDQKNVMDLGNSEIERNRRLEMLIARRKARKSFKMNSIAGSGPRHPVMVARSNTFHVSKSSDDQIPG
SAPSILLPTKNPFDLPYDPHEEKPNLMADSFHEEFMADHQKEFPFCRHESFSLGNSFQDNRQGQHEGRGSSRPKMQSGNKGNHDRLVDHLLFQGGETLRR
NLSVTDLVTEEPQSSNQVANKQERDREVGTTRIKLIGEKMEQSHYKDPSLGNGSDIQMNKDANAIKHKKILSNPSSSAEDILNAKTAENSESIQPTTFKF
PEVFYDRAPNSLPCPVPKARAVAEPSYDSSPSPIDNTRMEEHFFYKLKPGHTPNHSIASDMQVEVSEIGSPPLIEDGTASSNDDESLIYDGDSENEFTSG
SEELWGSSPLAPKVQEHGKARGQIYEEGEEGMTEVEFSRVWDEPENPISSSMWPLSSSRAEISQEDQAHSMKIDPKLSNHVKDEVDEVREQRPSNASDVD
EVEEVREERPSNSSDVVPPEHSLEGTRLMEGSMAHSPSEVYFPEPQESPSGSGNSAEEKKTNCDANETVTYDDREYLKSNENRHVGAENSIMQEVLGNLS
EPAEGNNSTSNSNIKIESLFNSEKYVEGIGDETYDVNDSTFLISDRLEDSKSNEERDSGAERVMQAVIGDLSQPAVESNSESSNHIESKSLNSPEKSREE
ANINNVNDPPVQIKDRVEDFKDVDRDSERLTEDSDIQSILMPVEAEDNPTSTQGSKEDQSTIEVGVSVVNQPFTDPTTSATLPEFVAEQVSNNSSLSSSP
KSVLAYRIPADIGSSSDFSQLVATDMEENLLMTATQDTSLAVNDSIDHPSIDRKSEKSEEPYDTQGKCTEEAIDMENMNGSVLDDEQIKENLKCRKNIEY
ESETLISNEASVELSEPLEEPQTADHLEGASARLVDNEASINLSKPDEEHVSSKVPGVLVEKEESTNPPRDVAGEVNHISDVSDPSINKNDDLEKLKSFE
GSEGEPDQFSTGHEIFVEPLKPANITSLEGHEYSPGVLTENETIVASSQAIEEVDNSRTSKETDEFGTQIADEEIEDLLKPGEAVVSSETTKNVQGDPKD
LIDQKAVLNPSTPAVDDDNILVTPEAKDSAADTIHNVNESEMSEFISNEKFKHVQDSEDESQRLDRQEDIMEPLKAVEVTNSESIRDIEGESKKLADDEV
NVTIPSQPEGEINSSDDREKTEDPGKSIVQENGMDISEVSRGNDIARAVEDNKDKSEDKTEING
|
DESeq2's median of ratios [POPLAR]
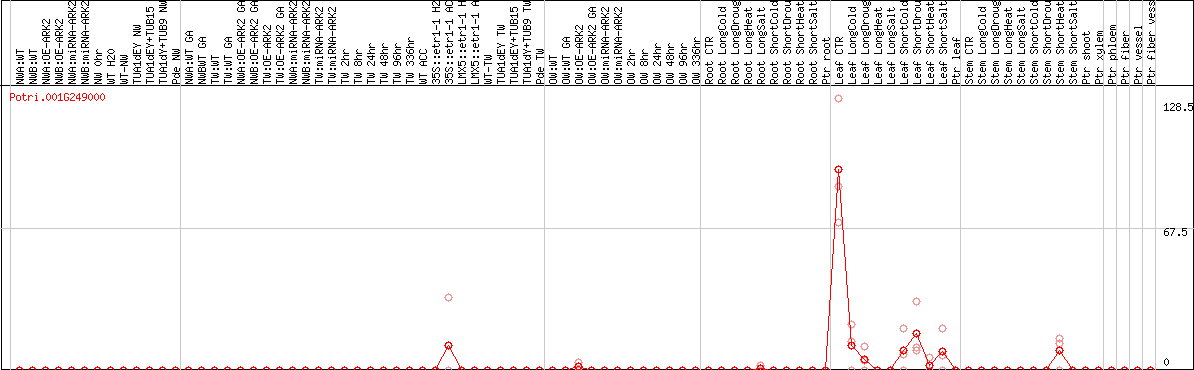
Coexpressed genes
Potri.001G249000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
