Potri.001G255700 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
No hit found | |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.001G255700.2 pacid=42792258 polypeptide=Potri.001G255700.2.p locus=Potri.001G255700 ID=Potri.001G255700.2.v4.1 annot-version=v4.1
ATGTATCGGAACCTTTTTATCACCAGATTCTGTCATGCCACTATGCCTTGTTTTCATTTAATATTTTTCATCTTGGTCCTCTTGTCATTGAGCCTTGGTT
CTGCTTCTTGGTGCCCACATAACTATGCTCAACAAAAGAATAGAGAATTTGAGCAGAAGACTGATCGTTTCTGGGAGTTCCAAGAGCAATCCAACACCTG
GGTTGAAGTGGAGCTGCCATATGAACTTGTCTCTTGTGTTAATGACAACTGTACTAAAGTGGGTAAAATTCATCCAGTAAAAAGAGATGTAGAAGAGAAC
TCAGAAAGAGAGAATGATGATTCCAAGAAAAATGAGAATTTGAAAAGGAAGGTTGAAGATGGAGGAACAGAAGCGAATTCTGAGATTGTTCTGCCTTTGA
GGAAGAGAATTTCTCTGACTAAAATGTCTGAGTCATCCATTTGGGTCACTGGTGAAAGTGGGTCTATCTATGAGAGGTTCTGGAATGGAATCCAGTGGGT
GATAGCACCCCATGATCTACCAGTATTGACTGGTCATGCAATCTGTGTCTTTATTGTCAATCAAACTATTCTTACTCTATCAGAAGCAGGAACCCTATAT
CAGATGATGCTCGGTGAGAGCTCACAGCCAATTTGGGTAGAGTTCACACCTACTCTCGATGAAAGCACAAATAGAGAAGCAGAAGAAAGTTCATTGATGC
TAATAAATTCCGGCGTCATTTCTCATGATGGGCTGAAAATTTATTTCTGCACAAAGAATGGCTCGTTATTAGAACTCAGTGAGGCAGAGCCTCCAAGATG
GGAAAATCACGGGCGACCACCTGGTGCAGATGTTGCAGCAATTGTTGATGCTGCTACTATTAGACCAGACGTAGTATATACCATAAGTTCTACAGGAGAT
CTTTATGAATATGATAGGAGCTCAAAGCCATCATGGAAGAAGCATATATGGGCAGAAGGGACAGTAGCAGATGCTTCTCTGATGCCATCAAGGGGATGCA
CACTACATGGCTTGAGTGGAGAATACTCCATCTCTCTTTTTCTTCTAACTAAGGGTGGTAAATTGGTAGAGAGACGATTAAACCAAAGAAAGTGGAAATG
GATAGTCCATGGAAGTCCCAAAGATCATAAGTTGACATCTATCACACCAGTTGTACAAGATGAGACAAATGAAAAATTCTTATCATTATTTTTCACCACA
TCGTCTGGATCGGTTTTTGAGTATCGAATATTGAAACAGTCAGGCACTGATCAGGAAAACCAAATTCCCGAAGCATGGTTGAGTCATATGCATCCTCCGC
ATGCGAAAGTTGCAAGTGGAATTGCTGGGATACCACTTCAAGCTGGCAGAATAGTTTTCCCACTGCATGATGGTAGACTTGCAGAATTGCATCTACCTGG
ACTGGGTGGAGAAAATACTGGGCCGAACCATCAAGTAAATCTCCGAAAAAGAGCATCGGTTAAATATGTATGGTCAATGATAGACGCCCCAGAGACTGAA
GGTTGGAATGCAGAGTATTGCAGAGAAGAACGTGGACCCATGAACTGCCTTGAAGGCATAAAAGATGAGCCAAATGAACAAGGAATTACAAGATCAATGG
CAAGAAGGCGAAAGGGAAGTAAAGCACAAGAAGATTACCTATTTGCAGGTGCCAATGGACCGAATAAAGTTTTGGAAGGATATAGTTTCCCAGATAATTG
GATCAACAATAACTTTCGTCTGCGAATGATACATGGAGGCAAATCATTTTTCCTTGTTACTGATGATGGTTTGACTTATGAACATCTTTATGCCGAGAAT
TTATGGCTGTGGCTGAGGCATGATCACTCAACACCAATGAAAGGTGCCCTTGGAAACTACAATGGAAGTCTCTTTTTGGTAGACATTTACGGGAGTCTGC
TTATGAGAGAAAGGAGCGATGAGGGGCTAACATGGGTAAATTGCACTGCCATGAGGAATCTCGGGCGTGTTATTGGAGGCCCTCCATGGGATGGAATTCC
AGGTAAAGATCCGAAAGTTACACCAGAAGATGCAATCTTCTTTGTGAGCAAAAATGGAAGGCTGCTGCAGTTCACAGTTGCCCTGAGGAAATTCAAATGG
AAAGATTGTCGAAACCCTCCAGATACCAAAGTTGCAAGTATTGTTGACCAGGAGCTGTTCCGGGATAACGTAGTGTTTGTCACTGGAAGGAATGGGAGGC
TATACCAGTACAACAAAGTGACTGAATTGTGGCATGAGCATTATCAGTCTCAACATTTGGTTCTATCAAGATCGCCAGGAACTGCTATGAGACCATCCTC
ACTATCACTCACAGGCTCACTATTTATGCTTTCAGAAGATGGTGGACTAGTTGAATATCATTGGAATACTGGTGATGGCTGGAATTGGATTGAACATGGA
ACACCTAATAAAGGTGTAACCTTGATCACTTCCCCTAGCCCATGTTTTGAAGGTAATCAACTTTTTCTTATAGGTTCAGATGGAAAAGTATACGTTAGGT
ATATGGACCAAATGACATGGAGATGGAAAAACTGTGGCTTCCCTCATGTGGGACAATTAATGAATGAAGATCAAACACAAGAGAGGGGAAATGATAATAA
TGAAGAAGTTTGCATAGATGAAGATTTTGCAGCCAGCTTGGAGAACGTTGCACGGAAGTACAGTGATTTTAATAGAAACTGTGACCCAAAGGTGGCACCA
ACAAGGCCGATCCCATTTTCTGATGATTCAGTTATCTTCGAGCTCAGGGATGGAAGATTGGCAGAGATGCGAAGAGTGGAGGGCACTCATTGGGTATGGT
CACGCACTATTGCCACTCCAACAACCTCGTGCATGGCTAATTATTGGACCGCTGTGGCTTCATGA
|
|||||||||||||||
|
AA sequence
|
>Potri.001G255700.2 pacid=42792258 polypeptide=Potri.001G255700.2.p locus=Potri.001G255700 ID=Potri.001G255700.2.v4.1 annot-version=v4.1
MYRNLFITRFCHATMPCFHLIFFILVLLSLSLGSASWCPHNYAQQKNREFEQKTDRFWEFQEQSNTWVEVELPYELVSCVNDNCTKVGKIHPVKRDVEEN
SERENDDSKKNENLKRKVEDGGTEANSEIVLPLRKRISLTKMSESSIWVTGESGSIYERFWNGIQWVIAPHDLPVLTGHAICVFIVNQTILTLSEAGTLY
QMMLGESSQPIWVEFTPTLDESTNREAEESSLMLINSGVISHDGLKIYFCTKNGSLLELSEAEPPRWENHGRPPGADVAAIVDAATIRPDVVYTISSTGD
LYEYDRSSKPSWKKHIWAEGTVADASLMPSRGCTLHGLSGEYSISLFLLTKGGKLVERRLNQRKWKWIVHGSPKDHKLTSITPVVQDETNEKFLSLFFTT
SSGSVFEYRILKQSGTDQENQIPEAWLSHMHPPHAKVASGIAGIPLQAGRIVFPLHDGRLAELHLPGLGGENTGPNHQVNLRKRASVKYVWSMIDAPETE
GWNAEYCREERGPMNCLEGIKDEPNEQGITRSMARRRKGSKAQEDYLFAGANGPNKVLEGYSFPDNWINNNFRLRMIHGGKSFFLVTDDGLTYEHLYAEN
LWLWLRHDHSTPMKGALGNYNGSLFLVDIYGSLLMRERSDEGLTWVNCTAMRNLGRVIGGPPWDGIPGKDPKVTPEDAIFFVSKNGRLLQFTVALRKFKW
KDCRNPPDTKVASIVDQELFRDNVVFVTGRNGRLYQYNKVTELWHEHYQSQHLVLSRSPGTAMRPSSLSLTGSLFMLSEDGGLVEYHWNTGDGWNWIEHG
TPNKGVTLITSPSPCFEGNQLFLIGSDGKVYVRYMDQMTWRWKNCGFPHVGQLMNEDQTQERGNDNNEEVCIDEDFAASLENVARKYSDFNRNCDPKVAP
TRPIPFSDDSVIFELRDGRLAEMRRVEGTHWVWSRTIATPTTSCMANYWTAVAS
|
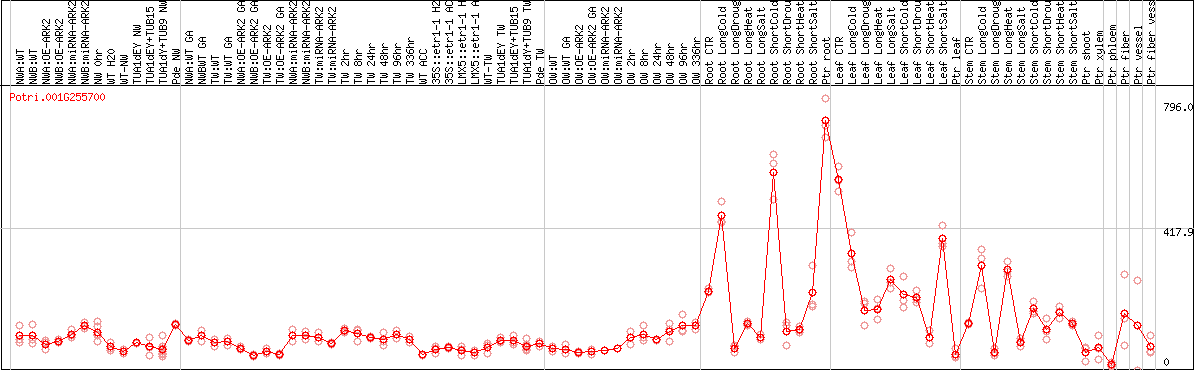
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G255700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
