External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G64330 94 / 8e-24
JK218, RPT3, NPH3
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
AT3G50840 81 / 3e-19
Phototropic-responsive NPH3 family protein (.1)
AT5G10250 80 / 6e-19
DOT3
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
AT5G66560 79 / 1e-18
Phototropic-responsive NPH3 family protein (.1)
AT5G03250 77 / 7e-18
Phototropic-responsive NPH3 family protein (.1)
AT2G30520 75 / 3e-17
RPT2
ROOT PHOTOTROPISM 2, Phototropic-responsive NPH3 family protein (.1.2.3)
AT1G30440 74 / 5e-17
Phototropic-responsive NPH3 family protein (.1)
AT5G48800 73 / 1e-16
Phototropic-responsive NPH3 family protein (.1)
AT4G31820 71 / 1e-15
MAB4, ENP, NPY1
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
AT3G44820 71 / 1e-15
Phototropic-responsive NPH3 family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G024400
160 / 3e-48
AT5G10250 331 / 5e-106
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.005G075400
100 / 6e-26
AT5G10250 695 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.017G048200
97 / 4e-25
AT5G64330 993 / 0.0
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Potri.007G112600
97 / 6e-25
AT5G64330 989 / 0.0
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Potri.007G093000
94 / 9e-24
AT5G10250 706 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Potri.005G130700
81 / 2e-19
AT5G66560 733 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.007G033900
81 / 3e-19
AT5G66560 789 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.007G053200
74 / 7e-17
AT5G67385 807 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.008G038600
73 / 1e-16
AT5G03250 723 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015169
100 / 3e-26
AT5G64330 264 / 7e-79
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Lus10040484
94 / 9e-24
AT5G64330 440 / 5e-150
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Lus10011290
92 / 3e-23
AT5G64330 438 / 3e-149
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
Lus10014337
91 / 1e-22
AT5G10250 620 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Lus10026046
90 / 2e-22
AT5G10250 639 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 3, Phototropic-responsive NPH3 family protein (.1)
Lus10041276
78 / 3e-18
AT5G66560 750 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10019522
77 / 1e-17
AT3G44820 781 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10039099
75 / 3e-17
AT5G47800 655 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10037432
75 / 5e-17
AT5G66560 747 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10019290
73 / 1e-16
AT5G67385 402 / 6e-135
Phototropic-responsive NPH3 family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0033
POZ
PF00651
BTB
BTB/POZ domain
Representative CDS sequence
>Potri.001G255950.1 pacid=42788940 polypeptide=Potri.001G255950.1.p locus=Potri.001G255950 ID=Potri.001G255950.1.v4.1 annot-version=v4.1
ATGATCTCAAGAAGTGAATATTTGAATAGATTAGTATTCCAGAGAAGAAGTAATGGAGGAAAGGGTACCTTCCCCAAGATTCAATTACACAATTTTCCAG
TTGGATCTGAAATCTTTGAGATAGCAGTGAAGTTCTGCTATGGATGGAAGGTTGATCTAACAGCATCCAACATTGCTCCTGTACACTGTGCTGCACGTTT
CTTGGAAATGAGCAATTATCTCGAACAAGGAAATCTAATTTCCAAAACAGAGGCTTTTATAAGTTTTGTATTGTTATGA
AA sequence
>Potri.001G255950.1 pacid=42788940 polypeptide=Potri.001G255950.1.p locus=Potri.001G255950 ID=Potri.001G255950.1.v4.1 annot-version=v4.1
MISRSEYLNRLVFQRRSNGGKGTFPKIQLHNFPVGSEIFEIAVKFCYGWKVDLTASNIAPVHCAARFLEMSNYLEQGNLISKTEAFISFVLL
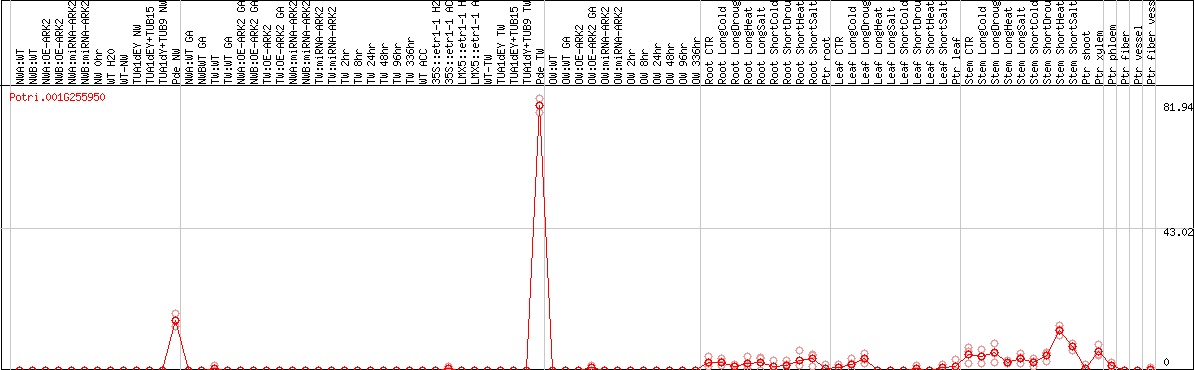
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G255950 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.