Potri.001G264100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G264100.1 pacid=42793480 polypeptide=Potri.001G264100.1.p locus=Potri.001G264100 ID=Potri.001G264100.1.v4.1 annot-version=v4.1
ATGAAGAGATTGAGATCAAGTGATGATCTTGATTCTTATAACGAGAAAAGCTCAGTCAAGGATTCAAATCCAAGTAGATCATCTAGAAGTTTCTATCACA
AATCTGATAATGTGCGCAAAGGATTGGTTTCCTCGTCATCATCCTCTAGCCGTTACGATCGAGACCGATCTACTGATGAGGATAATAGAGAGAGCTCCAG
AATGGTTAGGAAGCGATCAGATCATGAATTTGATAGCTTTGATAGGAGAAAAGGGTTAGGGTTAGGGTTTGATAGGTATGGCAGTGGAGGAGGTAGTAGT
AATAGCAGAGAAGGTTATTGTGGAAGTGGTGGTGGTGGTGGTAATGATAGGGTTATTCATCGACCGGAGAGTTTAGCTGGGTCGAGGAGGGAGTTTCCTA
AAGGGTTTAGGTCGGAGAGGGAAAGGTCAAGGAGAGAAGTGAGTGTTTCGTCGTGGAGGAGGTTTGGGAGTAAGGAATTTGAGGAGAGTAGAGGTGGTAG
TGGTAGAGGTGGAAATGAGGAGAGGATGGGGAGTGCTAGGTCTTCTCCTAAGGGGTTGAGAGATGTTGTTAGGTCACCTTCTTGGTCCAGGGATTCTGGG
AGTGAGCAAACTAGAGTGGCGAGAGGAAGTGGCAGTGGGAGGGATGAGGCGAAAGTGAAGTCTTCGAATTCCAAGTCTAGGTCTTCACCCACGTGGTCGA
AGGATTCGGGGAGTGAGCAATCTAAGAGTGTGGAGGTGGGGAAGAAAAGCGAAGCAGAGACAAAGAGTGTTGAGGTAGAGGCTAAAAGTGTGGAGATGGA
GGTGAAGGTTGTTCAGAGTGGGAATTGTAGCGAAATAGAAGAAGGGGAGCTTGAGCCAGAGCCTGATTCAGTTCCTAAAGTTGCCAAGGAAGATGAGAAT
GATAATGTAAATGAAGAGCTAGAGAATGTTAAAGTAGATATTGATCAGAGGAAGGTTGAGATTGAGGCTGAAGTTAAGGAGCTAGTAAATGAAGAGACAG
GGTCCCATAAAGAAAATGTTAATGAGGGGAAGGATGTGGTGAAGGAAGCTGGTGAGATGCCAAATGTTGAGGAGAATTCAAATGATAGTGTTAGTGAGGA
TGAAGTTGGAAATATGGATGGCGATGGAGATACCAAGGACAACAAGAGTTTGATGGAAAGAGTTGAGTGCAGGGGGGAGGTAAGTAAGAATATGATTGTT
GAGGAGTCTTTGAATTTGGAGGAGAATAATAAACAAGATAAGGGTATTGATCTCGAGGTGAAGGCAGATGATGTTGAAGTTACAGAGTCAAACAAGGAAA
CTGTGAAGGAAAATGGAGGAACTGAAGTGAATATAAATATGGTGACAGAGATTTCGAGTCAGAATGTGAAGGACAAAGGCAAAAGTGTGGCTGTTTCACC
GATCAATGCTCCTGATTCCGCAGAAGATGGTACATGGGCTGAAAGAGAATCGAGAAATGTTGCAACTTTCAGGAATGGGGAAGATGATATGGAAGGGCCA
AGTACTAGGGGATTTGAGTTGTTCTCTACCTCTCCTGTTAGAAGAGTCGAGAAAGCAGAGGAATCGAGTGGTATCAAATCGAAAGATGAAAAGCTGCTGT
TGGAACCGCTTGATCTTTCTCTTAGCTTACCGGATGTTTTGTTGCCTGTTGGTGCAACCGGAGACACAGGCCAGGCTCCTGGTTCTCCAAGTCATGGGAG
AAGTGTTCAGTCTTTTAGTTCATTTCGAACAAATTCAGATGGGTTTACTGCATCAATGTCCTTTTCAGGTTCTCAGTCATTTTATCACAATCCAAGCTGT
TCTCTAACTCAGAATTCCTTGGACATGGACAACTATGAACAATCTGTTCACAGCCGGCCCATATTTCAGGGTATTGATCAAACACATTGGCAGGGTCAGA
CTCAAAATGATTCCAAGTATAAAGATGTTCCCTTGTATCAAAAAATTTTGATGAATGGAAATGGCTCTCTTCATCAGCCTCAAGCAGTGCCAGGTTTGTC
AAATGGTCAAGCTTTGCAAGGAACCTCTAAAATGCACAATGAACTTGAAAGACAATTGAGTTTTCAAAGACAGTTGCCGGGGGGGCAGGCAAGGAACCAT
GATGACACTAGATCCCCTTCTCAAAGTGTGGGATCTCATGATATTGGTTCATCTTATAGCTTTGAGAAGAAGCGTGCCATGAAAGAGAAACATGGCAGTA
GCTTATATAGAAGTAATAGCCAGAAGGAACTAGAGCAATTCTCGATAGGTGGAGCTGATTTTGTTGAGACCATCATTGGCAGAATAGTTTCTGAGCCCAT
TCATGTGATGGCTAAGAAATTCCATGAAATGACAGCACAATCTGCATCGTGTTTGAAGGAGAGTATTCGAGAGATTTTGTTAAATGCCAATAAGCAAGGA
CAAGCATGTGCATTCCAAAGCATGCTGCAGAACAGGTCTGAGTTAACCTTGGATATGCTGTTGAAATCCCATCGTGTTCAACTGGAAGTCTTGGTTGCTT
TGCGAACGGGGTTGCCGGAGTATCTTCAAGTAGACAGTGGTATTTCATCTTCTGATTTGGCTGAGGTTTTCCTAAACTTGAGGTGCAGAAATCTGACATG
TCAAAGTCATTTGCCTGTGGATGAATGTGATTGCAAAGTTTGTGTGAAGAAGAATGGTTTTTGCAGTTCATGTATGTGTCTTGTGTGCTCAAAGTTTGAC
ATGGCATCTAATACATGCAGCTGGGTTGGGTGTGATGTTTGCCTTCATTGGTGTCATGCTGATTGTGCACTGCGAGAAGCTTGCATTAGGAATGGAAGGA
GTGTGAGTGGTGCTCAAGGGACTACTGAGATGCAGTTCCATTGTGTTGCCTGCGATCATCCTTCTGAGATGTTTGGCTTTGTGAAGGAGGTTTTCCAGAA
CTTCGCAAAAGATTGGACAGCAGAGACATTTTGCAGAGAACTTGAATATGTCAAGAGAATTTTCTGTGCCAGCAAGGATTTGAGAGGGAGACGGCTGCAT
GAAATAGCCGATCAGATGCTGGCAAAATTGGCAAATAAGTCCATTCTTCCCGAGGTGTATAATTATATAATGGGTTTCCTGACTGAAAGTGATCCATCCA
AGTTTGGCAACGCATCTGGTTTCTCTGGAAAGGAACAAGGAAATGGAAGCAATGGCATCATTGGTGGACCTAGCCAGGATACTGCATGGTTCAAATCTGT
CTATGCTGAAAAAACTCCTCAATTGGAAAGATCAACTAGCTTCCATTCTGATTTGAATGATAAACGCCCTGTTGAGTCCGAGTTGTTGAGAAGTGCCCAA
AAAGAGCCACTTTTTGATGAACTGGAGAGCATTGTCAGAATCAAACAGGCTGAAGCTAAGATGTTCCAAGCACGTGCTGATGATGCTAGAAGAGAAGCTG
AGGGCCTGAAACGAATTGTGATTGCAAAGAGTGAAAAAATTGATGAAGAGCATGCAGGTAGACTCTCAAAACTGCACATAGTTGAGGCTGAGGAAATGCG
GAGACAAAGATTTGAAGAATTTCAGTCCCTGGAAAGAGCTCATCGGGAATACTACAGTATGAAGATGAGGATGGAAGCAGATATTAAGGACCTCTTGTTA
AAAATGGAAGCTACAAAACGAAACCTCACCATGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G264100.1 pacid=42793480 polypeptide=Potri.001G264100.1.p locus=Potri.001G264100 ID=Potri.001G264100.1.v4.1 annot-version=v4.1
MKRLRSSDDLDSYNEKSSVKDSNPSRSSRSFYHKSDNVRKGLVSSSSSSSRYDRDRSTDEDNRESSRMVRKRSDHEFDSFDRRKGLGLGFDRYGSGGGSS
NSREGYCGSGGGGGNDRVIHRPESLAGSRREFPKGFRSERERSRREVSVSSWRRFGSKEFEESRGGSGRGGNEERMGSARSSPKGLRDVVRSPSWSRDSG
SEQTRVARGSGSGRDEAKVKSSNSKSRSSPTWSKDSGSEQSKSVEVGKKSEAETKSVEVEAKSVEMEVKVVQSGNCSEIEEGELEPEPDSVPKVAKEDEN
DNVNEELENVKVDIDQRKVEIEAEVKELVNEETGSHKENVNEGKDVVKEAGEMPNVEENSNDSVSEDEVGNMDGDGDTKDNKSLMERVECRGEVSKNMIV
EESLNLEENNKQDKGIDLEVKADDVEVTESNKETVKENGGTEVNINMVTEISSQNVKDKGKSVAVSPINAPDSAEDGTWAERESRNVATFRNGEDDMEGP
STRGFELFSTSPVRRVEKAEESSGIKSKDEKLLLEPLDLSLSLPDVLLPVGATGDTGQAPGSPSHGRSVQSFSSFRTNSDGFTASMSFSGSQSFYHNPSC
SLTQNSLDMDNYEQSVHSRPIFQGIDQTHWQGQTQNDSKYKDVPLYQKILMNGNGSLHQPQAVPGLSNGQALQGTSKMHNELERQLSFQRQLPGGQARNH
DDTRSPSQSVGSHDIGSSYSFEKKRAMKEKHGSSLYRSNSQKELEQFSIGGADFVETIIGRIVSEPIHVMAKKFHEMTAQSASCLKESIREILLNANKQG
QACAFQSMLQNRSELTLDMLLKSHRVQLEVLVALRTGLPEYLQVDSGISSSDLAEVFLNLRCRNLTCQSHLPVDECDCKVCVKKNGFCSSCMCLVCSKFD
MASNTCSWVGCDVCLHWCHADCALREACIRNGRSVSGAQGTTEMQFHCVACDHPSEMFGFVKEVFQNFAKDWTAETFCRELEYVKRIFCASKDLRGRRLH
EIADQMLAKLANKSILPEVYNYIMGFLTESDPSKFGNASGFSGKEQGNGSNGIIGGPSQDTAWFKSVYAEKTPQLERSTSFHSDLNDKRPVESELLRSAQ
KEPLFDELESIVRIKQAEAKMFQARADDARREAEGLKRIVIAKSEKIDEEHAGRLSKLHIVEAEEMRRQRFEEFQSLERAHREYYSMKMRMEADIKDLLL
KMEATKRNLTM
|
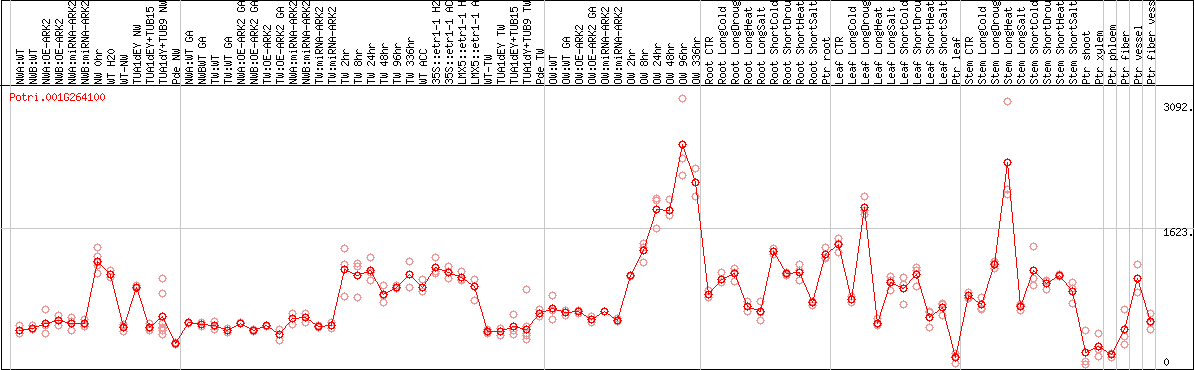
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G264100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
