Potri.001G274900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G274900.2 pacid=42790700 polypeptide=Potri.001G274900.2.p locus=Potri.001G274900 ID=Potri.001G274900.2.v4.1 annot-version=v4.1
ATGGCCGATACCAGAAGCCGAAACGGCAAAAAGATTCCGATACCGAAGTGCCCAATCTGCCAAACAGTTTACAACGATCACTCTCACACGCCTCTGCTTC
TCCAGTGCGGCCACTCTTTCTGTCTCAATTGTCTCTCTCGCCTCCTCCTCTCTTCCTCACCTTGTAAACCATCACAAAAATCCTCACTTACCTGTCCTAA
ATGCCGCCACGTCTCCCCTCTCGGCAACTCCCTCCTCTTCCTCCCTAAAAACTTCTCTCTCCTCTCTTCCCTCCTTCCCTCCTCCTCCGCCTCCGACTCC
GACTCCGACTCCTCCGACGACGAGAGCGATGTCAATCCTCGTCGATTTCATTGTTCTACAACAAGCGGCGTCGGATTGGCTGGGAATTTGTGTAGAGATC
ATGAATTGAGGATTGTGAAGGGGATTAGTGAGGTGAAGTGGAGCGGTTTGTTGGTGAAGGGGAAGGAGTGTAAGCATAAGGTAGTTGTGAGGAGAGTTAG
GGTTGATGACGTGGCCGATGGTAATTGGGTGGAGAAGGAGCTGGATAATTTGAGGAGGAAAGTTATGTGGTGTAGAAATGTGTGTGGATTTTATGGTGTA
GTGAAGAGTGGAGAGGATGATTTGTGTGTTGTGATGGAAAAATGTTTTGGTTCTGTTAAGGAACAGATGGAAGGGAATGGCGGCCGCCTCTCTCTCGATC
AAATCTTAAGGTATGGAGCTGACATTGCACGGGGGGTAATTGAAATTCATGCTGCGGGTATTGTGTGCCTGAACTTAAAGCCGTCCAATTTTCTTCTTGA
TGCAAGTGGTCATGTGGTGGTTTCTGATTATGGACTTCCGATGATTCTTAAGAAAACTTCTTGCTCAAAGGGTCTGCTTGTGCCTGAGGCTGATCCATTA
AAAACACATTGGTGTATTGATTGTTTGTCGCTAAGTCCATATTATTCAGCTCCAGAGGCATGGGAGCCATTAAAGAAATCTTTGCACTTATTCAGGGATG
GTAACATTGGCATATCTGCAAAATCAGATTCCTGGAGTTTTGGTTGTGCTCTCGTCGAAATGTGCACTGGCTTCACTCCGTGGGCTGGTCTAAGTGCAGA
AGAGATCTACCATGCTGTTGTTAAGGAAGGAAGATCACCTCCACAATATGAAAAAGTTGTAGGTCATGGGATTCCTACAGAACTATGGAAGATGATTGGC
GAGTGCCTACAATTTAAGGCATCGAACAGACCCACATTTCATGCAATGCTCGCCGTATTTCTGCGTCATTTGCAAGGAATACCTCGAAGTCCTGCAATTC
CTAATACAGAACCTGCAAACTGTTCCAGAATTGATATGCTGGAACAATCTCCGACATCCGTCCTGGACATTTTCCCAGTCAAAAGCAACCATCTGCATCA
ACTTTTATCTGACGGGGACGTGGATGGTGTCAGAGATCTTCTTGCCAAGTCTGCATCGGGTAACGATGGCAACTCAGTTATTTCTCTGCTTGAAGCACAT
AATGCTGATGGTCAAACTGCATTGCACTTGGCTTGTAGACGGGGTTGCCTTAAACTTGTTGGTGCCATTTTGGAGTACAGCGATGTGGATGTGGATATTC
ATGACAAAGATGGTAATCCACCAATAGTGTTTGCTTTAGCAGCTGGATCCCCAGAATGTGTAAGATCTCTTATCAGAAGATCAGATTATGCAACTTGTAG
GATGAGCGAAAGCATTGGCCGGTCAGTAGCTCATGTCTGTGCATATTATGGGCAACCAGACTGCATGTTGGAATTACTTTTGGCTGGTGCTGATGCCAAT
GCAGTGGATGATGATGGTGAATCTGTACTGCATGTAGCCATCACCAACAAACATACAGAATGTGCTATTGTCATATTGGAAAATAGTGGATGCAGATCAA
TGAGTTTTCTGAACTCAAAAAATATGACACCCCTACATTTGTGCATAGAAGCTCTAAATGTGACTGTTGTCAAGAGGTGGCTAGAAGTTGCATCAGAAGA
AGAAATCGCTGGAGCCATTGACCTGCCCAGTTCAGTTGGAACTGCATTATGTATGGCAGCTGCTTTAAGAAAAGATCACGAGACAGAGGGCAGAGAGCTG
GTGAGATTACTTCTTGCTGCTGGAGCAAACCCTGCTGCCCAAGATGCTGAAAATCATCAAACAGCTTTGCATACAGCTTCGGCAGCAAATGATGTTGAGT
TGGTCAAGATCATTCTTGATGCTGGAGTGAATGCTAACCTACGAAATGTGCATGGTACAATACCTCTTCATTTAGCATTGGCTAAAGGTGCAAAGCCTTG
CGTCGAATTGCTCTTGGCTGCTGGTGCAGATTGTAATTTGCAGGATGAGGATGGTGACAATGCCTTCCATATAGCTGCTGCGGCTGCAAGGTTGATACGT
GAGAATCTTGACTGTATCATCCTTATGCTGCAGTGCCCAAATGCAAGCATTAAAATTAGAAACAACAGTGGTAAGACGTTGTGTGACCTCCTGGAGAGCC
TGCCCCGAGAATGGATTTCAGAAGAGCTTTTGGAAGCACTAGCTAATAAGGGAGTTTCTTTATACCCAACAGTGTTTGAAGTTGGAGATTGGGTAAAATT
CAAGAGGGGCATGAAGAATCCAACATATGGGTGGCAAGGTGCAACGTGTGGCAGTGTTGGTTTTGTACAGAGTATTCCAGAGAATGGCAGTCTTACTGTA
TCATTTTGCACAGGAGTTGCTCATGTCTTGGCAAATGAAGTTGTAAAAGTGATTCCTTTTGACAGTGGCCAACTTGTGCAACTTAAACCTGATGTTATAG
AGCCCAGATATGCGTTGCATGAACAATTACGTCACAGTCAGGGAACTGTTCTGTGCATTGAGGATGAGGGGTTTATACGAATTGGTTTTCGCGGTGCCTC
TAAGGGGTGGCAAGCTGATCCTGCAGATTTTCAAAGATTTGAAGAATTCAAGGTTGGTGATTGGGTTTGTGTTCGCAAAACACCTGAGGCTGTAAAGCAC
GATTTTGGAATTGTGACTGCTGGAAGCATAGGGATCGTGCATGGTATCAGATCCGATAGTAGTTTGTTGATAGAGTTCAGTTGTGTACCTGCTCCATGCA
TATTTGAACCAGAGGAGGTTGAACCAGTTGTGCCTTTTAAGATAGGTGAGCTAGTATGTGTGAAACGATCTATTGCAGAGCCCAGCTTTGCTTGGGGTGG
TGAGACACATCATAGTGTTGGAAGAATATTTGACATAAAGAGTAATGGTCTTCTGATAATCGAAATACAAAATCGGTCCATACCATGGACGGCTGATCCT
TCAGATATGGAAAAGGTGGAGGACTTCAAGGTTGGTGACTGGGTTAGAGTAAAAGCTTCGGTACCCTCTCCAAAATATGGGTGGGATGATGTCACCAGGA
CTAGCATTGGTATTGTCCATAACTTAGAGGATGATGGTGACATGGGTGTTGCTTTCTCCTTTAGAAGCAGGCCCTTTCTATGTTCAATGACAGACATGGA
GAAGGTGTCACCTTTCAAAGTGGGACAAGAGATCCGTGTCATGCCGTCTATCACTCAGCCATTGCTTGGATGGTCAAATGAAAGTCCAGCTACTTTTGGG
ACTGTTGCGAGGATTGACATGGATGGAACCCTCAATGTGAGAGTGGCTCGTAGAGCAAGCTTATGGAAAGTTGCTCCAGGTGATGCTGAAAGGCTCTCAG
GACTCGCTGTTGGTGACTGGGTTAGGGTGAAGCAATGTCTGGGGTCTAGACCAAGTTATGAGTGGAACAGTTTTAGGAAGGAGAATATCGCTGTAGTGCA
CAGTATACAGGATTCTTTTCATCTAGATTTGGTTTGTTGTTTTCGTAAAGGAAAGTTGCCTGCTCACTGCACAGAGGTGGAAAAGGTTTCACACATAAAA
ATAGGGCAGCATGTTCGATTTCGTACTGGACTGGTTGAACCTAGGTGGGGATGGAGAGGCGCATGCCACAATTCAAGAGGTGTTGTAACTGCTGTAAATA
TTGATGGAGAAATAAGAGTTTCATTCTCTGGGTTGCACAATTTGTGGCGAGGAGATCCCGCGGATTTTGAGATAGATCAAATGTTTGAGGTGGGAGAATG
GGTAAAAATAAAAGACAGTGCCACCGGGTGGAAAAGTTTGGGGCCTGGAAGCCTTGGAGTTGTACAAGGAGTGAGGCATCAAGATGATGGTTGGGATGGT
ACTTTCCTTGTTGGGTTCTGTGGCGAACCTGAATTATGGGCAGGACCAGCATGTGAGCTTGAAACTGTGGATAAACTTGCAGTAGGCCAAAAGGTGAAGG
TGAAGCCCCACGTAAAGCAGCCACGATTTGGTTGGTCAGGCCATACACATGAGAGCATTGGTTCTATATCAGCGATAGATCTAGATGGAAAACTGAGGAT
ATTTACACCAGCTGGCTCCAAGGTCTGGATGCTTGATCCATCAGAAGTTGACGTGGTGGAGGAAGAGATAATACAAATTGGAGATTGGGTAAGAGTCAAA
TCATCTATAGCTACCCCAGTATACCAATGGGGAGAGGTCACTCGTGATAGCATTGGAGTGGTACACAAGATGGAGGAAGGAGAGCTTTTGGTGGCATTTT
GCTTCTTGGATCAGCTTTGGGTTTGCAAAGAATGGGAAATGGAAAAGGTAAGGGCATTTAAGGTAGGTGACAGTGTGAGATTTAGGGAACGACTGGTGAA
GCCCAGGTGGGGATGGGGAATGGAAACCTGTGCTAGCAAGGGCCAAGTAGTTGGGGTGGATGCTAATGGAAAGCTACGAATTAAATTCAAATGGAGAGAA
GGACGACCTTGGATTGGAGACCCAGCAGATATAATTCTCGATGAAAGCTCTAGTTTTGCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G274900.2 pacid=42790700 polypeptide=Potri.001G274900.2.p locus=Potri.001G274900 ID=Potri.001G274900.2.v4.1 annot-version=v4.1
MADTRSRNGKKIPIPKCPICQTVYNDHSHTPLLLQCGHSFCLNCLSRLLLSSSPCKPSQKSSLTCPKCRHVSPLGNSLLFLPKNFSLLSSLLPSSSASDS
DSDSSDDESDVNPRRFHCSTTSGVGLAGNLCRDHELRIVKGISEVKWSGLLVKGKECKHKVVVRRVRVDDVADGNWVEKELDNLRRKVMWCRNVCGFYGV
VKSGEDDLCVVMEKCFGSVKEQMEGNGGRLSLDQILRYGADIARGVIEIHAAGIVCLNLKPSNFLLDASGHVVVSDYGLPMILKKTSCSKGLLVPEADPL
KTHWCIDCLSLSPYYSAPEAWEPLKKSLHLFRDGNIGISAKSDSWSFGCALVEMCTGFTPWAGLSAEEIYHAVVKEGRSPPQYEKVVGHGIPTELWKMIG
ECLQFKASNRPTFHAMLAVFLRHLQGIPRSPAIPNTEPANCSRIDMLEQSPTSVLDIFPVKSNHLHQLLSDGDVDGVRDLLAKSASGNDGNSVISLLEAH
NADGQTALHLACRRGCLKLVGAILEYSDVDVDIHDKDGNPPIVFALAAGSPECVRSLIRRSDYATCRMSESIGRSVAHVCAYYGQPDCMLELLLAGADAN
AVDDDGESVLHVAITNKHTECAIVILENSGCRSMSFLNSKNMTPLHLCIEALNVTVVKRWLEVASEEEIAGAIDLPSSVGTALCMAAALRKDHETEGREL
VRLLLAAGANPAAQDAENHQTALHTASAANDVELVKIILDAGVNANLRNVHGTIPLHLALAKGAKPCVELLLAAGADCNLQDEDGDNAFHIAAAAARLIR
ENLDCIILMLQCPNASIKIRNNSGKTLCDLLESLPREWISEELLEALANKGVSLYPTVFEVGDWVKFKRGMKNPTYGWQGATCGSVGFVQSIPENGSLTV
SFCTGVAHVLANEVVKVIPFDSGQLVQLKPDVIEPRYALHEQLRHSQGTVLCIEDEGFIRIGFRGASKGWQADPADFQRFEEFKVGDWVCVRKTPEAVKH
DFGIVTAGSIGIVHGIRSDSSLLIEFSCVPAPCIFEPEEVEPVVPFKIGELVCVKRSIAEPSFAWGGETHHSVGRIFDIKSNGLLIIEIQNRSIPWTADP
SDMEKVEDFKVGDWVRVKASVPSPKYGWDDVTRTSIGIVHNLEDDGDMGVAFSFRSRPFLCSMTDMEKVSPFKVGQEIRVMPSITQPLLGWSNESPATFG
TVARIDMDGTLNVRVARRASLWKVAPGDAERLSGLAVGDWVRVKQCLGSRPSYEWNSFRKENIAVVHSIQDSFHLDLVCCFRKGKLPAHCTEVEKVSHIK
IGQHVRFRTGLVEPRWGWRGACHNSRGVVTAVNIDGEIRVSFSGLHNLWRGDPADFEIDQMFEVGEWVKIKDSATGWKSLGPGSLGVVQGVRHQDDGWDG
TFLVGFCGEPELWAGPACELETVDKLAVGQKVKVKPHVKQPRFGWSGHTHESIGSISAIDLDGKLRIFTPAGSKVWMLDPSEVDVVEEEIIQIGDWVRVK
SSIATPVYQWGEVTRDSIGVVHKMEEGELLVAFCFLDQLWVCKEWEMEKVRAFKVGDSVRFRERLVKPRWGWGMETCASKGQVVGVDANGKLRIKFKWRE
GRPWIGDPADIILDESSSFA
|
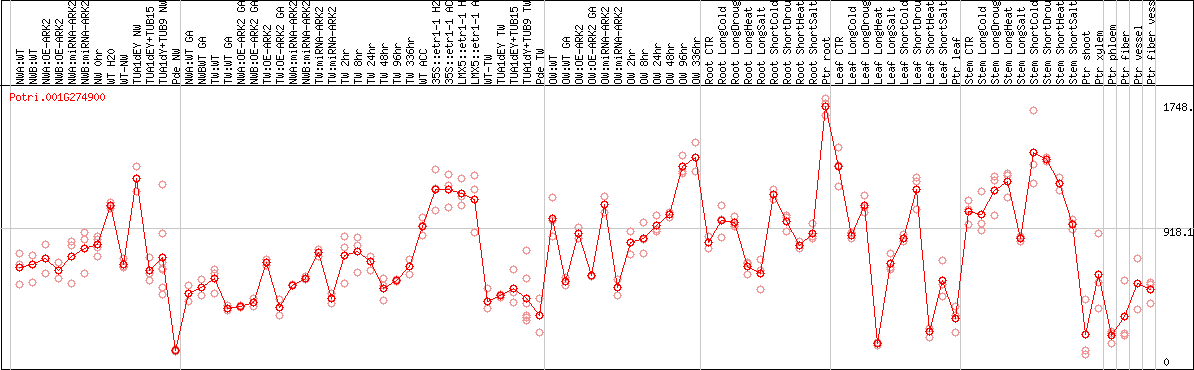
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G274900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
