Potri.001G278800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G278800.2 pacid=42792253 polypeptide=Potri.001G278800.2.p locus=Potri.001G278800 ID=Potri.001G278800.2.v4.1 annot-version=v4.1
ATGGCTGCTGCCAATGCCCCAATCACCATGAAAGAGGCTTTAACGTTGCCAAGCCTCGGGATCAATCCACAGTTCATAAATTTCACCCATGTGACGATGG
AATCAGAGAAGTACATCTGCATAAGAGAAACGGCGCCGCAGAACAGTGTGGTTATTGTCGATATGAGCATGCCTGCGCAGCCTCTGAGAAGGCCTATCAC
GGCCGATTCCGCCTTGATGAATCCTAATTCCCGAATTCTCGCTCTCAAAGCTCAACTCCCGGGAACTACTCAGGATCATTTACAGATATTTAATATCGAA
ATGAAAGCGAAAGTGAAATCACATCAAATGCCTGAGCAGGTTGTGTTTTGGAAGTGGAGTAGTGCTAACATGCTGGCCTTGGTAACACAGACTTCTGTTT
ATCACTGGTCAATTGAAGGTGATTCGGAGCCTGTGAAGATGTTTGATAGGACAGCTAATCTGCAAGGTAATCAGATAATCAACTATAGATGTGATCCTTC
AGAGAAGTGGTTGGTCCTCATTGGAATTGCTCAAGGTCCTCCAGAGAGGCCACAATTGGTCAAAGGGAATATGCAGCTTTTCTCTGTGGATCAGCAGCGT
AGTCAGGCTCTTGAAGCACATGCAGCGTCATTTGCGGCATTTAAAGTTGCAGGAAATGATAATGCTTCAATCCTTATCTCTTTTGCATCAAGGTCATTTA
ATGCTGGACAACTAACATCAAAGTTGCATGTTATTGAACTTGGTGCTGTGCCAGGCAAACCATCATTTACTAAGAAGCAAGCAGACCTGTTCTTTCCACC
AGACTTCGCTGATGATTTTCCTGTGTCAATGCAGATATCCCAAAAGTATGGCTTAATATATGTGATCACAAAGCAAGGACTTCTATTTGTTTATGACCTA
GAGACTGCTAGCGCGGTTTATAGAAACCGAATCAGTCCAGATCCTATATTTTTGACGACTGATGCTTCTTCAGTAGGTGGCTTCTATGCAGTCAATAGGA
GAGGACAGGTTTTGCTTGCCACTGTAAATGAAGCAACACTTGTGCCTTTCGTTAGTGGCCAGTTAAACAATCTTGAACTTGCTGTTAATCTTGCAAAAAG
AGGAAATCTTCCTGGTGCTGAAAATCTGGTTGTCCAGAGGTTTCAGGAGTTATTTTCTCAGGCAAAGTATAAGGAGGCTGCAGAACTTGCGGCAGAATCT
CCACAAGGAATCCTGCGCACTCCTGACACTGTTGCTAAATTTCAGAGTGTTCCGGTGCAAGCTGGGCAAACACCACCATTGCTGCAGTACTTTGGAACTC
TGTTGACTAGAGGAAAACTCAATGCTTTCGAGTCATTGGAATTATCACGCCTTGTTGTAAATCAGAATAAGAAAAATCTTCTGGAGAATTGGTTGGCAGA
GGACAAGCTTGAATGCACTGAAGAGCTTGGAGATCTTGTGAAGACAGTTGATAATGACCTTGCATTGAAGATATATATCAAAGCTAGGGCAACTCCAAAA
GTTGTTGCAGCATTTGCTGAGAGGAGGGAGTTTGACAAGATTCTTATATACTCAAAGCAGGTTGGCTATAATCCTGATTATTTGTTCCTTCTCCAAACAA
TATTGCGGACAGACCCTCAGGCTGCTGTTAATTTTGCTCTAATGATGTCTCAAATGGAGGGAGGTTGCCCAGTCGATTACAACACAATAACTGATCTATT
CCTTCAGAGGAATTTGATCCGTGAGGCAACAGCATTTTTGTTGGATGTTCTGAAGCCAAATTTGCCTGAGCATGGATTCCTTCAAACTAAGGTTTTGGAG
ATCAATCTTGTGACATTTCCAAATGTTGCAGATGCTATATTAGCCAATGGTATGTTTAGTCATTATGATCGCCCACGGATTGGCCAACTCTGTGAAAAAG
CTGGGTTGTATATTCGAGCGCTTCAGCACTATACCGAGTTGCCTGATATAAAACGTGTTATTGTGAACACACATGTTATTGAGCCTCAGGCTCTTGTTGA
GTTCTTTGGCACTCTTTCTCGAGAATGGGCTCTTGAATGTATGAAAGATCTTCTTCTAGTCAACCTGAGGGGCAACCTTCAGATTATTGTGCAGACTGCC
AAAGAATACTGTGAGCAATTGGGTGTGGATGCATGCATAAAGCTCTTTGAGCAGTTTAAGTCGTATGAAGGACTGTATTTCTTTTTGGGTTCATATTTGA
GCTCCAGTGAGGATCCTGAAATCCACTTCAAGTACATTGAAGCGGCTGCGAAAACTGGGCAGATTAAGGAGGTAGAACGTGTGACGCGTGAATCTAACTT
CTATGATCCTGAAAAGACAAAGAACTTTTTAATGGAAGCCAAACTTCCAGATGCACGTCCATTGATCAATGTTTGTGATCGTTTTGGATTTGTCCCGGAC
CTTACCCATTACCTGTACGTAAACAATATGCTTCGCTATATTGAGGGTTATGTTCAAAAGGTCAATCCAGGAAATGCTCCTTTAGTTGTAGGACAACTTC
TTGATGATGAATGTGCTGAAGATTTCATTAAAGGCCTTATTCTTTCAGTTCGTTCTCTTCTGCCAGTTGAGCCCTTAGTGGAGGAGTGTGAGAAAAGGAA
TCGTCTCCGCTTGCTCACTCAGTTCTTGGAGCATCTAGTCAGTGAGGGAAGCCAGGATGTGCATGTTCACAATGCTCTTGGTAAAATCATTATCGACAGT
GGCGACAATCCAGAGCATTTTCTTACCACCAACCCATATTACGATTCACGTGTTGTTGGTAAATATTGTGAGAAGCGGGATCCCACCCTTGCTGTTGTTG
CCTACCGAAGAGGACAGTGTGATGATGAACTAATCAATGTCACAAACAAGAATTCATTGTTCAAACTGCAGGCCAGATATGTTGTAGAACGGATGGATGG
TGATCTCTGGGAGAAAGTCCTTAGTCCTGATAATGAGTATAGACGGCAGCTGATTGATCAGGTTGTATCGACTGCTTTGCCTGAAAGCAAGAGTCCTGAT
CAAGTTTCAGCAACTGTTAAAGCTTTCATGACAGCTGATCTTCCACATGAACTGATCGAACTTCTTGAAAAGATTGTTCTCCAAAATTCTGCTTTCAGTG
GAAATTTCAATCTGCAAAACCTGCTCATCTTGACTGCCATCAAGGCTGATCCATCAAGAGTTATGGATTACATTAATAGGCTAGACAACTTTGATGGTCC
TGCAATTGGGGAGGTTGCTGTGGAAGCTCAATTGTATGAAGAGGCGTTTGCCATTTTCAAGAAGTTCAATCTGAATTTTTCAGCTGTCAATGTCCTGCTG
GATAATATTCGCAGCATTGATCGAGCTGTGGAGTTTGCATTCCGGGTGGAAGAAGAAGCTGTCTGGAGCCAAGTGGCCAAGGCCCAACTGCGAGAGGGAC
TAGTGAGTGAAGCAATTGAGTCATTTATCCGTGCAGACGATGCTACTCAATTTTTGGAAGTCATAAAGGCTGCCGAGGATGCAGATGTATATCATGACTT
GGTGAGGTACCTTCTAATGGTGAGGCAAAAGTCCAAGGAGCCCAAGGTGGATAGTGAACTCATTTATGCATATGCTAAAATTGATCAACTGGGTGAAATT
GAAGAGTTCATTCTCATGCCTAATGTCGCTAATCTCCAAAATGTTGGAGATCGCTTATATGATGAAGCCCTGTACGAGGCAGCAAAGATTATATTTAGAT
TTATTAGTAACTGGGCGAAGTTAGCTGTTACACATGTTAAGCTAAACGAGTTCCAAAGTGCTGTTGATGCAGCTCGAAAAGCAAACAGCTCGAAAACATG
GAAAGAAGTCTGCTTTGCTTGTGTTGATGCTGAGGAATTCCGCTTGGCTCAGATATGTGGTCTCAACATCATCATACAGGTTGATGATTTGGAAGAGGTC
AGTGAGTATTACCAGAACAGAGGATGCTTCAGTGAGCTCATCTCTCTAATGGAAAGTGGGCTTGGTTTGGAACGTGCGCATATGGGCATCTTCACTGAGT
TGGGAGTTTTATATGCGAGATACCGCCCTGAGAAGCTTATGGAGCACATCAAATTGTTCTCAACTCGTCTCAACATCCCCAAGGTTATACGGGCTTGTGA
TGAACAACAGCACTGGAAGGAACTGACTTATTTATACATCCAATATGATGAATTTGACAATGCTGCTACCACTATAATGAACCACTCTCCAGAAGCTTGG
GACCACATGCAATTCAAAGATGTTGTAGTAAAAGTTGCAAATGTTGAGCTCTACTACAAGGCTGTGCACTTTTACTTGCAAGAGCATCCAGACCTAATCA
ATGATCTTCTTAATGTGATTGCTCTTCGTGTTGACCATACTAGAGTGGTGGACATAATGCGAAAGGCTGGTCAGCTCCGCCTTGTTAAGCCATACATGGT
TGCAGTTCAGAGCAACAATGTCTCTGCTGTAAATGAAGCACTGAATGGAATTTACATTGAGGAGGAGGACTATGACAGACTTCGTGAATCTATTGAGTTG
CATGATAATTTTGACCAGATAGGTCTTGCACAGAAGATTGAGAAACACGAGCTTCTGGAGATGAGACGTGTTGCTGCTTACATATACAAGAAAGCTGGAA
GGTGGAAGCAGTCAATTGCTTTGTCAAAGAAAGACAATCTTTACAAAGATTGTATGGAAACTTGTTCACAGTCTGGTGAAAGGGAACTTTCTGAGGAGTT
GCTTGTCTATTTCATTGAGCAGGGAAAGAAAGAGTGCTTTGCAGCTTGCCTCTTTGTTTGTTATGATATGATCCGACCAGATGTTGCTCTTGAGCTGGCT
TGGATGAACAACATGATTGATTTTGCGTTCCCATATCTCTTGCAGTTCATCCGTGAGTACACAAGCAAAGTAGATGAACTCATCAAGGAAAAACTTGAGG
CTCTTAGTGAGGTGAAGGCTAAAGAGAAAGAGGAGAAGGACATGGTTGCTCAGCAGAATATGTATGCACAATTGCTGCCCCTTGCTTTGCCTGCGCCACC
TATGCCTGGAATGGGTGGAGGTTTTGCACCACCTCCAATGGGTGGAATGGGAATGCCTCCAATGCCACCTTATGGCATGCCGTCCATGGCCCCCTACTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G278800.2 pacid=42792253 polypeptide=Potri.001G278800.2.p locus=Potri.001G278800 ID=Potri.001G278800.2.v4.1 annot-version=v4.1
MAAANAPITMKEALTLPSLGINPQFINFTHVTMESEKYICIRETAPQNSVVIVDMSMPAQPLRRPITADSALMNPNSRILALKAQLPGTTQDHLQIFNIE
MKAKVKSHQMPEQVVFWKWSSANMLALVTQTSVYHWSIEGDSEPVKMFDRTANLQGNQIINYRCDPSEKWLVLIGIAQGPPERPQLVKGNMQLFSVDQQR
SQALEAHAASFAAFKVAGNDNASILISFASRSFNAGQLTSKLHVIELGAVPGKPSFTKKQADLFFPPDFADDFPVSMQISQKYGLIYVITKQGLLFVYDL
ETASAVYRNRISPDPIFLTTDASSVGGFYAVNRRGQVLLATVNEATLVPFVSGQLNNLELAVNLAKRGNLPGAENLVVQRFQELFSQAKYKEAAELAAES
PQGILRTPDTVAKFQSVPVQAGQTPPLLQYFGTLLTRGKLNAFESLELSRLVVNQNKKNLLENWLAEDKLECTEELGDLVKTVDNDLALKIYIKARATPK
VVAAFAERREFDKILIYSKQVGYNPDYLFLLQTILRTDPQAAVNFALMMSQMEGGCPVDYNTITDLFLQRNLIREATAFLLDVLKPNLPEHGFLQTKVLE
INLVTFPNVADAILANGMFSHYDRPRIGQLCEKAGLYIRALQHYTELPDIKRVIVNTHVIEPQALVEFFGTLSREWALECMKDLLLVNLRGNLQIIVQTA
KEYCEQLGVDACIKLFEQFKSYEGLYFFLGSYLSSSEDPEIHFKYIEAAAKTGQIKEVERVTRESNFYDPEKTKNFLMEAKLPDARPLINVCDRFGFVPD
LTHYLYVNNMLRYIEGYVQKVNPGNAPLVVGQLLDDECAEDFIKGLILSVRSLLPVEPLVEECEKRNRLRLLTQFLEHLVSEGSQDVHVHNALGKIIIDS
GDNPEHFLTTNPYYDSRVVGKYCEKRDPTLAVVAYRRGQCDDELINVTNKNSLFKLQARYVVERMDGDLWEKVLSPDNEYRRQLIDQVVSTALPESKSPD
QVSATVKAFMTADLPHELIELLEKIVLQNSAFSGNFNLQNLLILTAIKADPSRVMDYINRLDNFDGPAIGEVAVEAQLYEEAFAIFKKFNLNFSAVNVLL
DNIRSIDRAVEFAFRVEEEAVWSQVAKAQLREGLVSEAIESFIRADDATQFLEVIKAAEDADVYHDLVRYLLMVRQKSKEPKVDSELIYAYAKIDQLGEI
EEFILMPNVANLQNVGDRLYDEALYEAAKIIFRFISNWAKLAVTHVKLNEFQSAVDAARKANSSKTWKEVCFACVDAEEFRLAQICGLNIIIQVDDLEEV
SEYYQNRGCFSELISLMESGLGLERAHMGIFTELGVLYARYRPEKLMEHIKLFSTRLNIPKVIRACDEQQHWKELTYLYIQYDEFDNAATTIMNHSPEAW
DHMQFKDVVVKVANVELYYKAVHFYLQEHPDLINDLLNVIALRVDHTRVVDIMRKAGQLRLVKPYMVAVQSNNVSAVNEALNGIYIEEEDYDRLRESIEL
HDNFDQIGLAQKIEKHELLEMRRVAAYIYKKAGRWKQSIALSKKDNLYKDCMETCSQSGERELSEELLVYFIEQGKKECFAACLFVCYDMIRPDVALELA
WMNNMIDFAFPYLLQFIREYTSKVDELIKEKLEALSEVKAKEKEEKDMVAQQNMYAQLLPLALPAPPMPGMGGGFAPPPMGGMGMPPMPPYGMPSMAPY
|
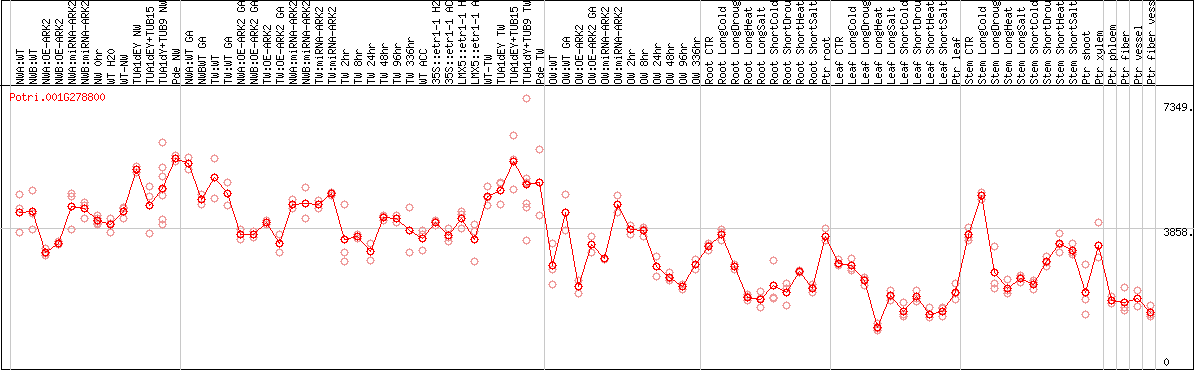
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G278800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
