External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G14610 178 / 4e-58
PR-1, PR1, ATPR1
pathogenesis-related gene 1 (.1)
AT2G14580 176 / 4e-57
ATPRB1
basic pathogenesis-related protein 1 (.1)
AT4G33720 159 / 2e-50
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT3G19690 149 / 2e-46
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT5G26130 145 / 5e-45
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G33710 143 / 4e-44
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT4G33730 142 / 1e-43
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT1G50060 133 / 3e-40
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
AT2G19990 122 / 1e-35
PR-1-LIKE
pathogenesis-related protein-1-like (.1)
AT4G31470 120 / 5e-35
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020493
162 / 2e-51
AT2G14610 206 / 9e-69
pathogenesis-related gene 1 (.1)
Lus10012479
161 / 2e-51
AT2G14610 208 / 7e-70
pathogenesis-related gene 1 (.1)
Lus10020491
159 / 2e-50
AT2G14610 207 / 2e-69
pathogenesis-related gene 1 (.1)
Lus10025697
148 / 6e-46
AT1G50060 213 / 1e-71
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10007102
147 / 3e-45
AT2G14610 182 / 3e-59
pathogenesis-related gene 1 (.1)
Lus10020480
142 / 2e-43
AT2G14610 182 / 3e-59
pathogenesis-related gene 1 (.1)
Lus10012478
140 / 8e-43
AT2G14610 170 / 1e-54
pathogenesis-related gene 1 (.1)
Lus10020481
140 / 1e-42
AT2G14610 176 / 6e-57
pathogenesis-related gene 1 (.1)
Lus10013693
120 / 9e-35
AT4G31470 196 / 1e-64
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
Lus10006980
118 / 3e-34
AT4G30320 203 / 1e-67
CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 protein) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0659
CAP
PF00188
CAP
Cysteine-rich secretory protein family
Representative CDS sequence
>Potri.001G288401.1 pacid=42793481 polypeptide=Potri.001G288401.1.p locus=Potri.001G288401 ID=Potri.001G288401.1.v4.1 annot-version=v4.1
ATGGAAATCTCCAAGAATTCACTAGCAATTGCTTTTCTTATTTCCTTAGCTATAATAATCCCTCTATCCCTTGCCCAAGACTCCCCACAAGACTACCTCA
ATGCCCACGCAAATGCTCGTGCACAGGTAGGTGTTGGAAATAATGTATGGGACAATAATGTGGCAGCTTATGCAAGTGACTATGTTAAACGGCTCACGGG
CGATTGCAGACTTGTGCATTCTGGTGGGCCTTATGGCGAGAACCTTGCATGGAGTAGTGGTGATCTTACAGGCAGCGATGCGGTGAAATTGTGGGTTGAT
GAGAAATCAAATTATGATTACAACTCCGATTCATGTGTTGGTGGAGAATGCAGGCATTATACTCAGGTGATTTGGCGCAACTCGTTTCGTTTAGGTTGTG
CTAAAGCAAGGTGCAGCAATGGGGGTACACTCATCAGTTGCAACTATGCTCCCTCTGGCAACTTCGTTAACGAGCGCCCTTACTAG
AA sequence
>Potri.001G288401.1 pacid=42793481 polypeptide=Potri.001G288401.1.p locus=Potri.001G288401 ID=Potri.001G288401.1.v4.1 annot-version=v4.1
MEISKNSLAIAFLISLAIIIPLSLAQDSPQDYLNAHANARAQVGVGNNVWDNNVAAYASDYVKRLTGDCRLVHSGGPYGENLAWSSGDLTGSDAVKLWVD
EKSNYDYNSDSCVGGECRHYTQVIWRNSFRLGCAKARCSNGGTLISCNYAPSGNFVNERPY
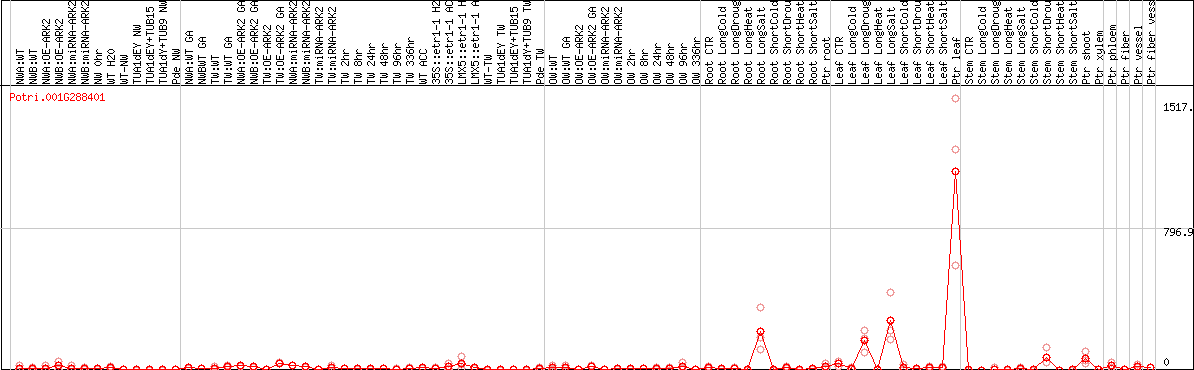
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G288401 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.