Potri.001G289800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G289800.1 pacid=42788715 polypeptide=Potri.001G289800.1.p locus=Potri.001G289800 ID=Potri.001G289800.1.v4.1 annot-version=v4.1
ATGGCCAACAACCCTCAATTCTCTGGAATGCAGCCTCTCCAGCCTCCTCTGGTTGGCCCCATGGATCATCCTCAAAATTTTGCTCCGCCACCACCAATGC
CCATTCAATTCCGACCGGTTGGTCCAGTGCAGCCATCACAGCAATTCATTCCTGTGAGTTCTCCACACTTTCAGCCTGTAGGCCGAGGTGTCACAGTGAT
GAATCCAGGATTGCCTCCCCAACCCCCACAACCTCAATTTCCTCACCCAATGCAGCAGTTACCAGCTAGACCTAACCAGCCAAGCCTTGGTCCACCACCA
CCTCAGGCAATTCCATTACCAAATGCTCAGCCAAACAGGCATGTCATGTCTGGATCTCCACTGCCTCCACCTAGTGTTCAAACTCCAAACAGCTACATGC
CTGGTTTAGGTGGCCCAGGAGTGCCTCTATCGTCATCGTATACTTTTGCACCGTCTTCATATGGTCAACCACCTGTAACCTTTAATGCTGTAACTCAATT
TCAGCCCATGCCTCAAATGCATGTACAGCCTATTCCCACTGGAGGACACCCTGCATCATCCATGAACCATAACACTGCACCAGTTACACCTATACAGCGC
AATGGCGAACAATCTTCAGTTACCACTACTAATGTTCGGGCAACAAGTATCCAGCCTAAGCCGACTGAAGAAGCCTTGACAGAGTGGAAAGAACATACAT
CTGCTAATGGAAGGAGATTTTATTACAATAAGAGGACAAGGCAATCTAGTTGGGAGAAGCCTTATGAGTTGCTGACGCCAATTGAGAGGGCAGATGCATC
AACTGACTGGAAGGAGTTTAAGAGTCCTGATGGACGAAAATATTACTACAATAAAGTTACCAAGCAGTCAAAATGGGAAATCCCAGAAGAATTGAAGTTG
GCCCGTGCACGAGTAGAAAATACATCTACCATGGAAAAACAATCAGAAGTCTTTACTAATTCTCATGCCTCTACTTCTGTTCCTCAATCCGCGGATAAGA
CACCATCAATTGTAGATGCTTCTACAGCTCAAGGGGCACCATCGAGTCCGGTTCTTGTTATACCTGTTGCTGCTGCTGGTAATTCACAGTCGCAGTTAGC
TTCTGAATCATCGACCTTACCTGTCATGTCATCTTCCATGACGACCAATGCAGATGAAGTTCAGACAATTGAGATTCCTGTTGCTGATGTTCCAAAAAGT
GCTGAAGTCACTGCCACAGCGGTGAACACCATAACTGCACCAATGAACAACTTTTCAGATCAAGATAAACCTAGTTCAGCAGATGAAGCTCCCGCACAAG
ACAAAGAGGAAGCTGAAAAGGAGGTGGTCATAGATGAAAAGGTTAACAATGTTCCATTGGAAGAGAAAGCAGTTAATCATGAGCCCTTACTTTATGCTGA
TAAGCTGGAAGCAAAAAATTTATTTAAAGCACTTCTGGAATCTGCAAATGTTGGGTCTGAATGGACATGGGACCAGGCCATGAGAGTAATAATTAATGAT
AAAAGATATGGTGCATTAAAAACACTTGGAGAGAGAAAGCAAGCATTTAATGAGTTCTTGGGTCAAAAGAGAAAACAGGAAGCTGAGGAAAGGAGGATTA
AACAGAAAAAAGCACGGGAAGAATTCAAGAACATGCTAGAAGAGTCCAAAGAGCTGACAGCATCAATTAGATTGAGCAAAGCAGTGACTTTGTTTGAAAA
CGATGAGCGTTTTAAAGCTGTTGAACGGGAAAGAGACCGTAAGGATCTGATTGAAACCTACTTGCAGGAACTTGAAGAAAAGGAACGAGCAAAGGCACAG
GAGCAGCGCAAGAGGAACATTATGGAATATAGGCAGTTTCTGGAATCATGCGAATTCATTAAGGCAAGCACCCAGTGGCGAAAAGTACAAGATCGGTTAG
AAGCTGATGAAAGATGTTCACGTCTTGAAAAAATTGATCGCATAGAAATTTTTCAGGATTATTTACATGATTTGGAGAAGGAAGAAGAGGAGCAAAGGAA
GATACATAAGGAAGAGCTGAGAAAGGCAGAGCGTAAAAATCGTGATGAGTTCCGCAAGCTACTTGAAGAACATGTTGCTGCAGGCACTCTTACTGCCAAA
ACTAACTGGCGTGATTATCACCTGAAGGTTAAAGATTTACCTGCATATGTGGCTGTTGCTTCAAACAACTCAGGTTCAACTCCAAAAGATTTGTTTGAAG
ATGTCGCTGAGGAGCTACAGAAACAGTATCATGAGGACAAAACTCGGATAAAGGATGTTGTAAAGTTGAAAAAGGTTCCGTTGGCATCAACCTGGACACT
TGAAGATTTGAAGGTTGCTATTATCGAGGATGTTGGCTCTCCACATATATCAGATGTGAACCTAAAGATGGTATTTGATGAATTGCTGGAAAGAGCCAGG
GAAAAGGAGGAGAAAGAAGCCAGGAAGCGTAAACGTCTTGAGGATGATTTTCTCATTCTACTACAGTCTATTAAGGATATTACTGCATCTTCTAAGTGGG
AGAGCTGCAAGGAAATCTTTGATGGTAGCCGAGAATACAGTTCCATTGGTGAAGAAGGCTTCTGTAGAGAGATATTTGAGGAATATGTATCTCAACTGAA
AGATCAGGAGAAAGAGAATGAGTGGAAACGAAAGGAGGAAAAGGCAAAAAAGGAAAAAGAGAGGGAGGAAAGAGAAAGGAGGAAAGCAAAACACAGAAGG
GAAAAGGAGAGGGGACATGATAGAGAAACAAGAAAGGAAGAAGAAGATGTTGAAATTGATGATACAATAGAAACTCAAGTTTGTAGTGATAAGAAAAGAT
CTGGGAGTGACAATAGCAGGAAACAGAGGAAGAGGCATCAAAATGCTGTGGATGATCTGGATGAAAGCGAGAAGGATAGGTCCAAGAGTTCACACAGACA
TAGCAGCAGCGACCTCAAAAAATCAAGAAGGCATGCTTCCACTCCTGAGTCAGATAGTGAAAGCAGGCACAAGAGACACAAGAGAGACCACCGAAATAGT
TCTCGTAGAACTGGTGATCTTGAGGATCTTGAAGATGGGGAATTTGGTGAGGATAGGGAAACTCGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G289800.1 pacid=42788715 polypeptide=Potri.001G289800.1.p locus=Potri.001G289800 ID=Potri.001G289800.1.v4.1 annot-version=v4.1
MANNPQFSGMQPLQPPLVGPMDHPQNFAPPPPMPIQFRPVGPVQPSQQFIPVSSPHFQPVGRGVTVMNPGLPPQPPQPQFPHPMQQLPARPNQPSLGPPP
PQAIPLPNAQPNRHVMSGSPLPPPSVQTPNSYMPGLGGPGVPLSSSYTFAPSSYGQPPVTFNAVTQFQPMPQMHVQPIPTGGHPASSMNHNTAPVTPIQR
NGEQSSVTTTNVRATSIQPKPTEEALTEWKEHTSANGRRFYYNKRTRQSSWEKPYELLTPIERADASTDWKEFKSPDGRKYYYNKVTKQSKWEIPEELKL
ARARVENTSTMEKQSEVFTNSHASTSVPQSADKTPSIVDASTAQGAPSSPVLVIPVAAAGNSQSQLASESSTLPVMSSSMTTNADEVQTIEIPVADVPKS
AEVTATAVNTITAPMNNFSDQDKPSSADEAPAQDKEEAEKEVVIDEKVNNVPLEEKAVNHEPLLYADKLEAKNLFKALLESANVGSEWTWDQAMRVIIND
KRYGALKTLGERKQAFNEFLGQKRKQEAEERRIKQKKAREEFKNMLEESKELTASIRLSKAVTLFENDERFKAVERERDRKDLIETYLQELEEKERAKAQ
EQRKRNIMEYRQFLESCEFIKASTQWRKVQDRLEADERCSRLEKIDRIEIFQDYLHDLEKEEEEQRKIHKEELRKAERKNRDEFRKLLEEHVAAGTLTAK
TNWRDYHLKVKDLPAYVAVASNNSGSTPKDLFEDVAEELQKQYHEDKTRIKDVVKLKKVPLASTWTLEDLKVAIIEDVGSPHISDVNLKMVFDELLERAR
EKEEKEARKRKRLEDDFLILLQSIKDITASSKWESCKEIFDGSREYSSIGEEGFCREIFEEYVSQLKDQEKENEWKRKEEKAKKEKEREERERRKAKHRR
EKERGHDRETRKEEEDVEIDDTIETQVCSDKKRSGSDNSRKQRKRHQNAVDDLDESEKDRSKSSHRHSSSDLKKSRRHASTPESDSESRHKRHKRDHRNS
SRRTGDLEDLEDGEFGEDRETR
|
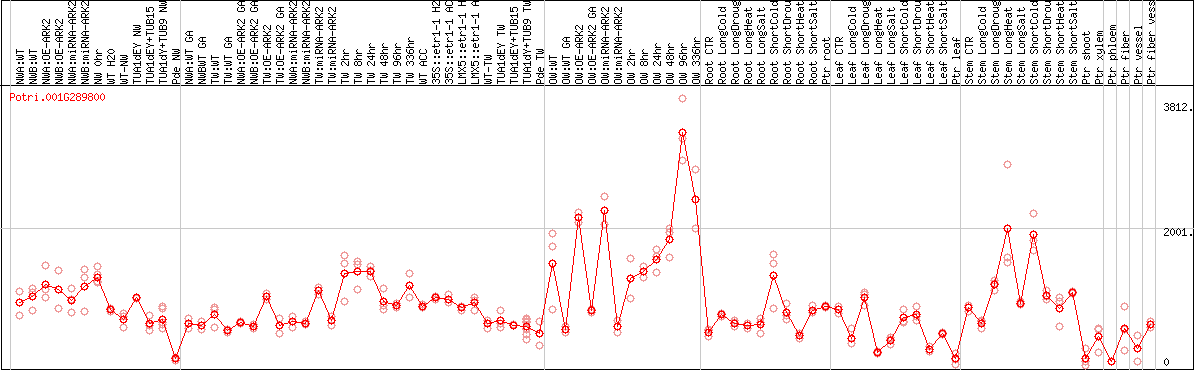
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G289800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
