Potri.001G310900 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G310900.1 pacid=42789466 polypeptide=Potri.001G310900.1.p locus=Potri.001G310900 ID=Potri.001G310900.1.v4.1 annot-version=v4.1
ATGGATCCTGAAATCGCCAAAACCCAAGAAGAACGCAAAAAAATGGAACAACAACTTGCCTCTTTAACCTCTTTAACCTTCGATCGTGACTTGTATGGTG
GAGTCGACCGTGAGGCTTATGAAACATCGATTCCAGCTACAGATGATGAAGAACCCGACATGGGTCTTAACGAGGTTGCTCAGAGACTCGCTTCGTATAC
TGCACCGAAATCAGTTCTTAAAGAAATGCCTCGTGGCGGTGATGATAGCGAAGAAACTGATGGGTTTAGGAAGCCATCGAGAATTATTGATCGCGAGGAT
GATTATAGGAGGAGGAGATTGGATAGAATTATTTCGCCGGAGAGGCATGATCCTTTTGCTGCCGGTGAGAAGACGCCGGATCCCAGTGTGAGGACTTATT
CGGATGTTATGATGGAAGAGTCTTTGAAAAAGCAAAAGGAGGAGTTGTTGAGGAAGATTGCCAACAAAAAGAAGGAAGAAGAGGAGGCGAGGGCTGGAAA
GGAGGAAAATGTGGCGAGTACAGTGCCTAAAAGGAGCAATAGGTGGGATCAATCGAAGGAGGATGATGGGAAAGTGGTTAAGAAGGCGAAGACAGGTTCG
GATTGGGATTTACCTGATGCTACACCGGGAATTGGGAGATGGGATGCGACACCTACTCCGGGGAGGATTGGAGATGCTACACCTGGTGCTGGGAGGAGGA
ATAGGTGGGATGAGACTCCTACACCGGGGAGAGTGGTTGATTCAGATGCTACTCCTGCAGGTGGGGTTACGCCAGGGGCTACTCCTGCTGGTGTTGCTTG
GGACGCTACTCCTAAAGGAATGGTTACCCCTACCCCGAAAAGGCAAAAGTCTCGTTGGGATGAGACTCCTGCTAGTATGGATAGTGCGACTCCTGCTCTT
GGGGCGGTTACTCCTAGTCTTGGTGGTGTGACTCCTGGACCTACTCCTCTTGGTGCAATTGATATGGCTACTCCAACTCCGAATGCACTTGCCATGCGCG
GGGCAATAACGCCTGAGCAGTATAATCTGCTGAGGTGGGAGAAGGATATTGAGGAGAGGAATAGGCCATTGACTGATGAGGAGTTGGATGCTATGTTTCC
TCAAGAAGGGTACAAGATTTTGGAGCCACCGGCTTCTTATGTGCCTATTAGGACACCAGCAAGGAAGTTGTTGGCTACCCCAACACCAATGGGCACTCCT
CTTTATTCTATTCCTGATGAGAACAGAGGGCAACAGTTTGATCTAGGGCAGGAGCCTCCTGCTGGGTTGCCATTTATGAAACCTGAGGATTATCAGTATT
TTGGTGCTTTGTTGAATGAGGAAGATGAGGAGGAGTTGTCCCCAGAGGAGCAGAAAGAGAGGAAGATTATGAAACTTTTACTGAAAGTGAAGAACGGGAC
ACCTCCACAGAGGAAGACTGCTTTGAGGCAGTTGACGGATAAGGCTAGAGAGTTTGGTGCTGGGCCGTTGTTCAACAGAATTTTGCCACTGCTTATGCAG
CCTACCTTGGAGGATCAGGAGAGGCATCTTCTGGTCAAGGTAATTGATAGGGTTTTGTATAAATTGGATGAGTTAGTTAGGCCTTATGTTCATAAGATTC
TTGTTGTTATTGAACCACTCTTGATTGATGAGGATTATTATGCCCGTGTGGAAGGGAGAGAAATTATTTCTAATTTGAGTAAAGCAGCTGGTTTGGCCAC
TATGATTGCTGCTATGCGTCCTGATATTGATAATATTGATGAGTATGTTAGGAACACAACTGCAAGAGCTTTCAGCGTTGTCGCTTCAGCTCTCGGGATT
CCTGCATTGTTGCCCTTCTTGAAAGCTGTGTGTCAGAGTAAGAAATCCTGGCAAGCTCGTCATACTGGAATCAAAATTGTTCAACAGATTGCTATTTTGA
TTGGCTGTGCTGTTCTTCCCCATTTGAGGTCTCTTGTGGAAATCATAGAACATGGTCTGAATGATGAGAATCAGAAGGTGAGGACAATCACTGCCTTGTC
CTTGGCTGCTCTTGCTGAGGCTGCTGCCCCATATGGCATTGAAAGCTTTGATTCTGTTCTGAAGCCATTGTGGAAGGGCATTAGGTCACACCGTGGCAAG
GTCTTGGCTGCCTTCTTGAAAGCAATTGGTTTTATTATTCCTCTTATGGATGCTATGTATGCCAACTACTACACCAAGGAAGTCATGTTTATTCTGATAC
GTGAGTTTCAGTCACCTGATGAAGAAATGAAGAAGATTGTGTTGAAAGTGGTTAAGCAATGTGTGAGTACAGAGGGTGTGGAGGCTGAATATATACGTTC
TGATATTCTTCCTGAGTTCTTTAGGAACTTCTGGGTTAGAAGGATGGCCTTGGATAGGAGAAATTATAGGCAACTTGTGGAAACAACTGTTGAGATTGCT
GACAAGGTTGGTGTTAAAGATATTGCTGGCAGAATTGTTGAGGATCTGAAAGATGAGAGTGAACCATATAGGCGTATGGTTATGGAAACAATTGAGAAGG
TTGTTGCGAACATGGGTGCATCTGATATTGATTCTAGGTTGGAAGAGCTTTTGATTGATGGCATTCTCTATGCTTTCCAAGAGCAGACCAGTGATGATGC
TAATGTGATGCTTAATGGGTTTGGTGCAGTTGTCAATTCTCTTGGACAGAGGGTGAAACCTTACCTTCCCCAGATTTGTGGTACCATTAAGTGGCGCTTG
AACAACAAGAGCGCTAAAGTGAGACAGCAAGCTGCTGATCTTATTTCTAGGATTGCTGTTGTCATGAAGCAATGTCAGGAGGAACAACTGATGGGTCATC
TTGGTGTTGTGTTGTATGAGTATTTGGGAGAAGAATACCCTGAAGTTCTTGGTTCAATTCTAGGAGCTCTCAAAGCAATTGTCAATGTCATTGGTATGAC
CAAAATGACTCCTCCTATTAAGGACTTGCTTCCAAGACTGACACCAATTCTGAAGAATAGGCATGAGAAAGTCCAGGAGAACTGTATTGACCTTGTTGGT
CGAATTGCTGATCGTGGAGCTGAGTTTGTTCCTGCCAGGGAGTGGATGAGGATTTGTTTTGAGCTTCTTGAGATGCTTAAGGCCCACAAGAAGGGCATTC
GGCGTGCTACTGTGAACACTTTTGGTTACATTGCTAAGGCCATTGGACCGCAGGATGTCTTGGCGACTCTGTTAAATAATCTCAAGGTCCAGGAGCGCCA
AAACCGGGTTTGCACAACTGTTGCAATTGCTATAGTTGCAGAAACCTGTTCACCTTTTACAGTCTTGCCTGCCTTGATGAATGAATATCGTGTCCCAGAG
CTTAATGTGCAGAATGGTGTCTTAAAATCCCTCTCTTTCCTGTTCGAGTACATTGGCGAAATGGGTAAAGACTATATCTATGCAGTGACTCCATTACTTG
AGGATGCACTCATGGACCGAGATTTAGTTCACAGGCAGACTGCAGCTTCCGCTGTTAAGCATATGGCTTTGGGTGTGGCTGGTTTGGGTTGTGAAGATGC
ATTGGTCCACTTACTTAACTATGTGTGGCCCAACATATTTGAGACTTCCCCTCATGTGATAAATGCTGTCACTGAAGCCATTGAAGGAATGAGGGTGGCC
TTAGGTGCTGCTGTTGTGTTGAACTACTGTCTTCAGGGGCTGTTCCATCCTGCCCGGAAGGTCAGGGAAGTGTACTGGAAGATATACAATTCACTGTATA
TTGGAGCTCAGGATGCTCTTGTGGCAGCTTATCCTATACTGGAGGATGAGCAGGATAACGTTTATAGTAGGCCCGAGTTGATGATGTTTGTATAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G310900.1 pacid=42789466 polypeptide=Potri.001G310900.1.p locus=Potri.001G310900 ID=Potri.001G310900.1.v4.1 annot-version=v4.1
MDPEIAKTQEERKKMEQQLASLTSLTFDRDLYGGVDREAYETSIPATDDEEPDMGLNEVAQRLASYTAPKSVLKEMPRGGDDSEETDGFRKPSRIIDRED
DYRRRRLDRIISPERHDPFAAGEKTPDPSVRTYSDVMMEESLKKQKEELLRKIANKKKEEEEARAGKEENVASTVPKRSNRWDQSKEDDGKVVKKAKTGS
DWDLPDATPGIGRWDATPTPGRIGDATPGAGRRNRWDETPTPGRVVDSDATPAGGVTPGATPAGVAWDATPKGMVTPTPKRQKSRWDETPASMDSATPAL
GAVTPSLGGVTPGPTPLGAIDMATPTPNALAMRGAITPEQYNLLRWEKDIEERNRPLTDEELDAMFPQEGYKILEPPASYVPIRTPARKLLATPTPMGTP
LYSIPDENRGQQFDLGQEPPAGLPFMKPEDYQYFGALLNEEDEEELSPEEQKERKIMKLLLKVKNGTPPQRKTALRQLTDKAREFGAGPLFNRILPLLMQ
PTLEDQERHLLVKVIDRVLYKLDELVRPYVHKILVVIEPLLIDEDYYARVEGREIISNLSKAAGLATMIAAMRPDIDNIDEYVRNTTARAFSVVASALGI
PALLPFLKAVCQSKKSWQARHTGIKIVQQIAILIGCAVLPHLRSLVEIIEHGLNDENQKVRTITALSLAALAEAAAPYGIESFDSVLKPLWKGIRSHRGK
VLAAFLKAIGFIIPLMDAMYANYYTKEVMFILIREFQSPDEEMKKIVLKVVKQCVSTEGVEAEYIRSDILPEFFRNFWVRRMALDRRNYRQLVETTVEIA
DKVGVKDIAGRIVEDLKDESEPYRRMVMETIEKVVANMGASDIDSRLEELLIDGILYAFQEQTSDDANVMLNGFGAVVNSLGQRVKPYLPQICGTIKWRL
NNKSAKVRQQAADLISRIAVVMKQCQEEQLMGHLGVVLYEYLGEEYPEVLGSILGALKAIVNVIGMTKMTPPIKDLLPRLTPILKNRHEKVQENCIDLVG
RIADRGAEFVPAREWMRICFELLEMLKAHKKGIRRATVNTFGYIAKAIGPQDVLATLLNNLKVQERQNRVCTTVAIAIVAETCSPFTVLPALMNEYRVPE
LNVQNGVLKSLSFLFEYIGEMGKDYIYAVTPLLEDALMDRDLVHRQTAASAVKHMALGVAGLGCEDALVHLLNYVWPNIFETSPHVINAVTEAIEGMRVA
LGAAVVLNYCLQGLFHPARKVREVYWKIYNSLYIGAQDALVAAYPILEDEQDNVYSRPELMMFV
|
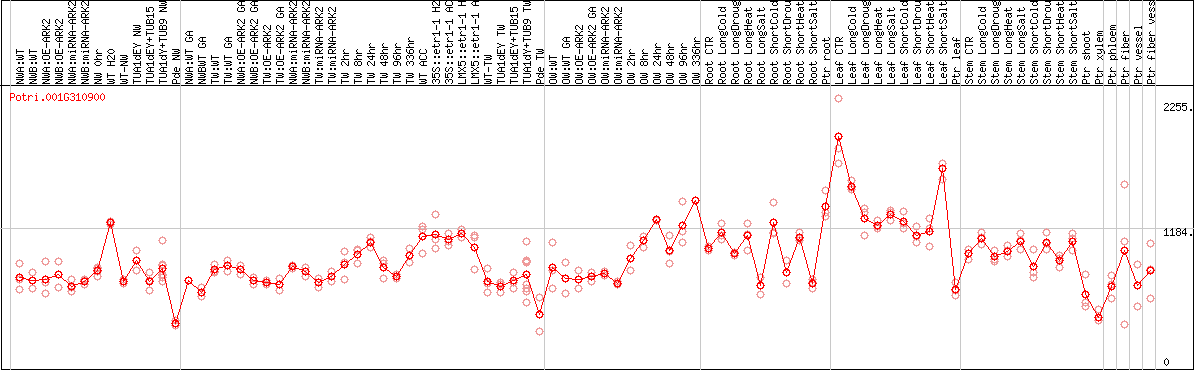
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G310900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
