External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G65260 238 / 3e-80
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G10350 235 / 6e-79
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT5G51120 212 / 6e-70
PABN1, ATPABN1
polyadenylate-binding protein 1 (.1.2.3)
AT3G12640 61 / 1e-10
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G36660 55 / 6e-09
PAB7
poly(A) binding protein 7 (.1)
AT3G18610 54 / 2e-08
ATNUC-L2, PARLL1, ATRANGAP1
PARALLEL1-LIKE 1, nucleolin like 2 (.1)
AT3G55460 51 / 8e-08
SCL30, At-SCL30
SC35-like splicing factor 30 (.1)
AT2G23350 49 / 1e-06
PABP4, PAB4
poly(A) binding protein 4 (.1)
AT2G35410 48 / 1e-06
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G16940 48 / 1e-06
Splicing factor, CC1-like (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G051600
341 / 3e-121
AT5G65260 288 / 4e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.005G073200
260 / 6e-89
AT5G10350 292 / 2e-101
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Potri.007G095800
249 / 2e-84
AT5G10350 285 / 3e-98
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Potri.012G115100
233 / 6e-78
AT5G65260 271 / 5e-93
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.015G110800
231 / 2e-77
AT5G51120 266 / 1e-90
polyadenylate-binding protein 1 (.1.2.3)
Potri.012G145050
65 / 6e-13
AT2G24350 107 / 4e-27
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.009G063800
67 / 8e-13
AT3G12640 418 / 6e-138
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G269200
66 / 1e-12
AT3G12640 432 / 3e-143
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.008G056600
53 / 2e-08
AT3G55460 134 / 6e-38
SC35-like splicing factor 30 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.001G311600.1 pacid=42788494 polypeptide=Potri.001G311600.1.p locus=Potri.001G311600 ID=Potri.001G311600.1.v4.1 annot-version=v4.1
ATGGACGGTGACGATGTGGACATGGCCGCTGCAGAGACCAACGAATCTGTACCAGAACTTGATGATATGAAGAAGAGACTAAAGGAAATGGAAGATGAAG
CTGCTGCTCTTCTTGAGATGCAGGCCAAGGTTGAGAAGGAAATGGGCTCTGTACAAGATCCTAGTGCTTCAGCTGCTGCATCCCAAGCAAATAGAGAGGA
AGTAGATTCTCGATCAGTGTTTGTTGGTAATGTGGACTATTCGTGCACCCCTGAAGAAGTGCAGCAGCATTTTCAGGCATGTGGGACAATAAATAGGATC
ACTATCCGTTCTGACAAGTATGGCCAACCAAAGGGTTATGCATATGTCGAATTCCTTGAACCAGAGACTGTGCAGGAGGCTCTCCTTCTGAATGAGTCTG
AGTTACATGGCCGCCAATTGAAGGTAACTGTTAAGCGGACCAATCTACCTGGGATGAAGCAGTTTCGCGCGCGTCGACCCAACCCATATATGGGGTTTCC
ACCCAGGGGCGCTCCTATGCCTCCTTATCTCTTTTCTCCTTATGGATATGGGAAGGTTCTGAGATACAGAATGCCAATGCGTTACAGCCCCTATGGCTGA
AA sequence
>Potri.001G311600.1 pacid=42788494 polypeptide=Potri.001G311600.1.p locus=Potri.001G311600 ID=Potri.001G311600.1.v4.1 annot-version=v4.1
MDGDDVDMAAAETNESVPELDDMKKRLKEMEDEAAALLEMQAKVEKEMGSVQDPSASAAASQANREEVDSRSVFVGNVDYSCTPEEVQQHFQACGTINRI
TIRSDKYGQPKGYAYVEFLEPETVQEALLLNESELHGRQLKVTVKRTNLPGMKQFRARRPNPYMGFPPRGAPMPPYLFSPYGYGKVLRYRMPMRYSPYG
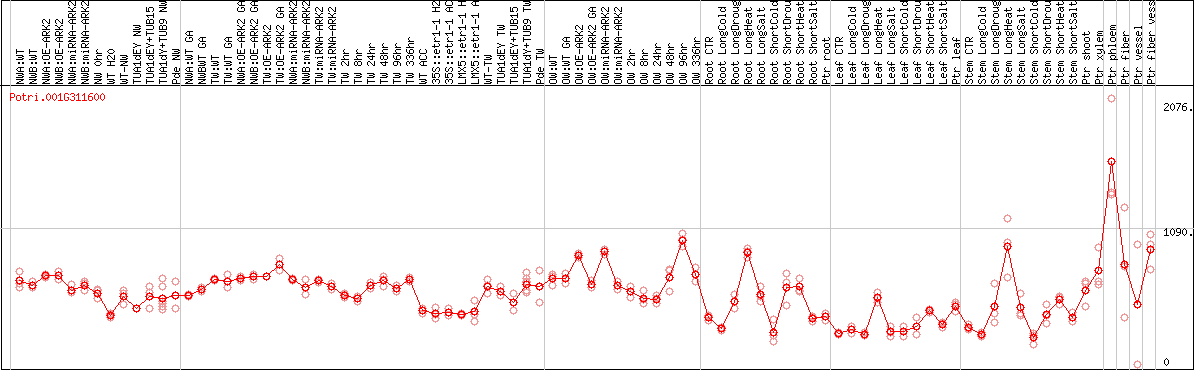
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G311600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.