External link
Symbol
NAC114
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13180 162 / 2e-49
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
AT2G33480 149 / 8e-44
NAC
ANAC041
NAC domain containing protein 41 (.1.2)
AT5G14000 141 / 9e-42
NAC
ANAC084
NAC domain containing protein 84 (.1)
AT3G15510 122 / 9e-33
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G77450 114 / 2e-30
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT1G01720 114 / 3e-30
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G61110 114 / 6e-30
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G04060 114 / 8e-30
NAC
ANAC046
NAC domain containing protein 46 (.1)
AT3G04070 112 / 3e-29
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT5G39610 111 / 3e-29
NAC
ORE1, AtNAC2, ANAC092, ATNAC6
ORESARA 1, NAC domain containing protein 2, Arabidopsis NAC domain containing protein 92, NAC domain containing protein 6 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G063300
398 / 3e-142
AT5G13180 150 / 9e-45
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.003G166500
194 / 1e-61
AT5G13180 310 / 4e-107
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.001G061200
187 / 1e-58
AT5G13180 310 / 6e-107
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.015G102100
153 / 9e-46
AT5G13180 232 / 8e-77
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.012G103500
149 / 4e-44
AT5G13180 229 / 2e-75
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.011G046700
125 / 9e-34
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Potri.013G054000
123 / 3e-33
AT3G04070 338 / 9e-115
NAC domain containing protein 47 (.1.2)
Potri.001G404400
123 / 3e-33
AT3G15510 345 / 1e-117
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Potri.002G081000
122 / 3e-33
AT1G01720 416 / 5e-148
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002581
188 / 3e-59
AT5G13180 273 / 6e-93
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10041534
186 / 2e-58
AT2G33480 125 / 1e-34
NAC domain containing protein 41 (.1.2)
Lus10001809
181 / 1e-56
AT5G13180 271 / 6e-92
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10012557
169 / 7e-52
AT5G14000 122 / 4e-34
NAC domain containing protein 84 (.1)
Lus10032004
159 / 9e-48
AT5G14000 113 / 2e-30
NAC domain containing protein 84 (.1)
Lus10035174
154 / 5e-46
AT5G14000 115 / 2e-31
NAC domain containing protein 84 (.1)
Lus10032657
124 / 4e-33
AT3G15510 353 / 1e-120
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10043095
122 / 1e-32
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10006547
121 / 3e-32
AT3G04070 337 / 4e-114
NAC domain containing protein 47 (.1.2)
Lus10011215
121 / 3e-32
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.001G325100.1 pacid=42793721 polypeptide=Potri.001G325100.1.p locus=Potri.001G325100 ID=Potri.001G325100.1.v4.1 annot-version=v4.1
ATGGAAAGGCCAAACTTTGTTGAGAATGGAGGGCTCAGATTGCCTATTGGATATAGGTTTCATCCAACGGATGAAGAGCTTGTTGTTCATTACCTGAAGC
GAAAGGTCCTTGGTCTTCCATTGCCAGCTGCTGTCATTCCTGAGTTTGATGTATTCCAAACTGATCCTTGGAGCTTGCCAGGTAATTTGAAGGAAAAGAG
GTACTTCTTTAGCCAAAAGAAGTTGAATGACTTTGGAACAAAATGCAAGAGATCCGCTGGTTATATACCTTCTGGTTACTGGAAACCCGTTGGGAAAGGC
AAGCAAATTGTAGCTTCTGGTAGCAACCAAGCAGTTGGAATAAGGAAAACGCTGGTTTTCAGAGAAAGAAATCATTCTAGTAAGACTAGAACTCAATGGG
TGATGCATGAATATTGCCTTGTAGGCTTGCCAACAGACCCCAAAACCACCCAGATGAAAGTGGGAGATTGGGTTGCTAACAGTATATTTCACAGGAAGAG
GAAACCTAAAAACCATGTGGTCATGATTTCCAACCCTTCAAACATTAACAAGAGGCAAAATGTCGAGATTATTAGTCCTAGTTTCATGGATTTTATGATG
GAACAAAGCTCTGATGAAGCTGGTGCTCCTTCACAATGTTCAAGTGGAGTCACCGAGGTGTCTTCTAATGAGTTTGATCAGGAAGAAATCAGTTCCTTCA
TTAGCCTGTTTACTTATCCTTGTAAGAGGAAGAGGACTTGA
AA sequence
>Potri.001G325100.1 pacid=42793721 polypeptide=Potri.001G325100.1.p locus=Potri.001G325100 ID=Potri.001G325100.1.v4.1 annot-version=v4.1
MERPNFVENGGLRLPIGYRFHPTDEELVVHYLKRKVLGLPLPAAVIPEFDVFQTDPWSLPGNLKEKRYFFSQKKLNDFGTKCKRSAGYIPSGYWKPVGKG
KQIVASGSNQAVGIRKTLVFRERNHSSKTRTQWVMHEYCLVGLPTDPKTTQMKVGDWVANSIFHRKRKPKNHVVMISNPSNINKRQNVEIISPSFMDFMM
EQSSDEAGAPSQCSSGVTEVSSNEFDQEEISSFISLFTYPCKRKRT
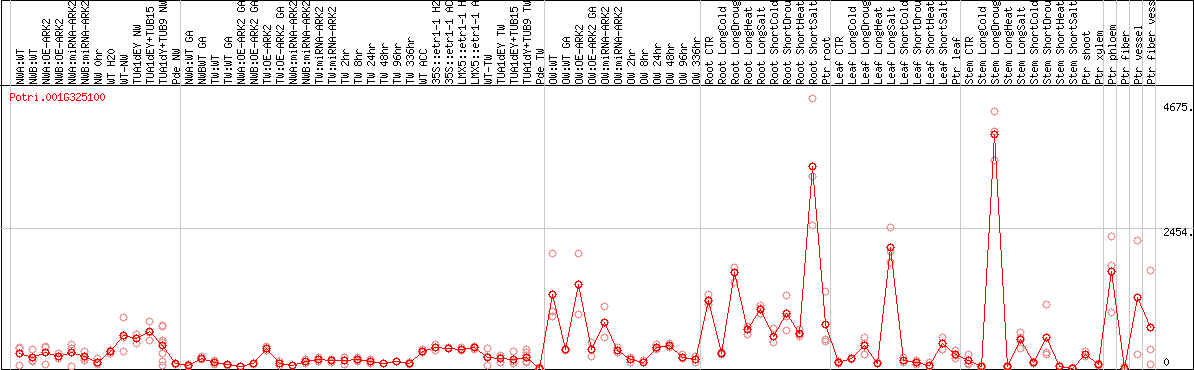
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G325100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.