External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13080 174 / 2e-56
WRKY
ATWRKY75, WRKY75
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 75, WRKY DNA-binding protein 75 (.1)
AT3G01970 157 / 1e-49
WRKY
ATWRKY45, WRKY45
WRKY DNA-binding protein 45 (.1)
AT1G64000 134 / 6e-40
WRKY
ATWRKY56, WRKY56
WRKY DNA-binding protein 56 (.1)
AT5G41570 132 / 4e-39
WRKY
ATWRKY24, WRKY24
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 24, WRKY DNA-binding protein 24 (.1)
AT2G46130 122 / 3e-36
WRKY
ATWRKY43, WRKY43
WRKY DNA-binding protein 43 (.1.2)
AT1G29860 113 / 6e-31
WRKY
ATWRKY71, WRKY71
WRKY DNA-binding protein 71 (.1)
AT5G49520 114 / 2e-30
WRKY
ATWRKY48, WRKY48
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 48, WRKY DNA-binding protein 48 (.1)
AT4G18170 112 / 3e-30
WRKY
ATWRKY28, WRKY28
WRKY DNA-binding protein 28 (.1)
AT2G47260 111 / 1e-29
WRKY
ATWRKY23, WRKY23
WRKY DNA-binding protein 23 (.1)
AT5G46350 110 / 4e-29
WRKY
ATWRKY8, WRKY8
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 8, WRKY DNA-binding protein 8 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012547
186 / 3e-61
AT5G13080 172 / 2e-56
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 75, WRKY DNA-binding protein 75 (.1)
Lus10041546
186 / 4e-60
AT5G13080 171 / 1e-54
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 75, WRKY DNA-binding protein 75 (.1)
Lus10011346
179 / 1e-57
AT5G13080 179 / 1e-58
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 75, WRKY DNA-binding protein 75 (.1)
Lus10003128
178 / 2e-57
AT5G13080 179 / 2e-58
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 75, WRKY DNA-binding protein 75 (.1)
Lus10007906
137 / 1e-40
AT1G64000 168 / 6e-54
WRKY DNA-binding protein 56 (.1)
Lus10036401
137 / 1e-40
AT1G64000 170 / 1e-54
WRKY DNA-binding protein 56 (.1)
Lus10017067
119 / 2e-32
AT5G49520 184 / 2e-54
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 48, WRKY DNA-binding protein 48 (.1)
Lus10037785
115 / 6e-31
AT5G49520 179 / 4e-53
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 48, WRKY DNA-binding protein 48 (.1)
Lus10004612
113 / 2e-30
AT1G29860 188 / 3e-58
WRKY DNA-binding protein 71 (.1)
Lus10004537
111 / 5e-30
AT1G29860 188 / 1e-58
WRKY DNA-binding protein 71 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0274
WRKY-GCM1
PF03106
WRKY
WRKY DNA -binding domain
Representative CDS sequence
>Potri.001G328000.2 pacid=42790488 polypeptide=Potri.001G328000.2.p locus=Potri.001G328000 ID=Potri.001G328000.2.v4.1 annot-version=v4.1
ATGGAAAAGTTCCAAATGTTATTTCCTTTTTCGTCGACAGGATCACCAAACTACCCTGTCTCTTCAAGCATGACAAATAGTTCTCATGTTTTTCATAATT
TTCAAGCAGATAGTTCAGGTGGGTTCTTGGGATTGAAGACAGAAACTCAACACCCAGTACCAAGAACATCACCCGAAATTAATAAAGACCTAAATTCTCA
AAGTGCAAGCTGTGCAATCGGGTCTGAAACTGAGCCTCAAGCAGCCAAGAAGAAAGGGGAGAAGAAGATTAGAAAGCCCAAGTATGCTTTTCAAACTAGG
AGTCAAGTTGATATACTTGATGATGGTTATAGATGGAGGAAGTATGGACAAAAGGCAGTGAAGAACAACAAATTTCCTAGGAGCTATTACCGATGCACGT
ATCAAGGGTGTAACGTAAAGAAGCAAGTCCAACGCCTAACCAAAGACGAAGGTGTTGTAGTGACGACTTACGAAGGAATGCACACCCATCCTATAGAGAA
GCCAAATGATAATTTTGAACATATCTTGAGCCAGATGCAAATTTACAGTACTCCCTTTTGA
AA sequence
>Potri.001G328000.2 pacid=42790488 polypeptide=Potri.001G328000.2.p locus=Potri.001G328000 ID=Potri.001G328000.2.v4.1 annot-version=v4.1
MEKFQMLFPFSSTGSPNYPVSSSMTNSSHVFHNFQADSSGGFLGLKTETQHPVPRTSPEINKDLNSQSASCAIGSETEPQAAKKKGEKKIRKPKYAFQTR
SQVDILDDGYRWRKYGQKAVKNNKFPRSYYRCTYQGCNVKKQVQRLTKDEGVVVTTYEGMHTHPIEKPNDNFEHILSQMQIYSTPF
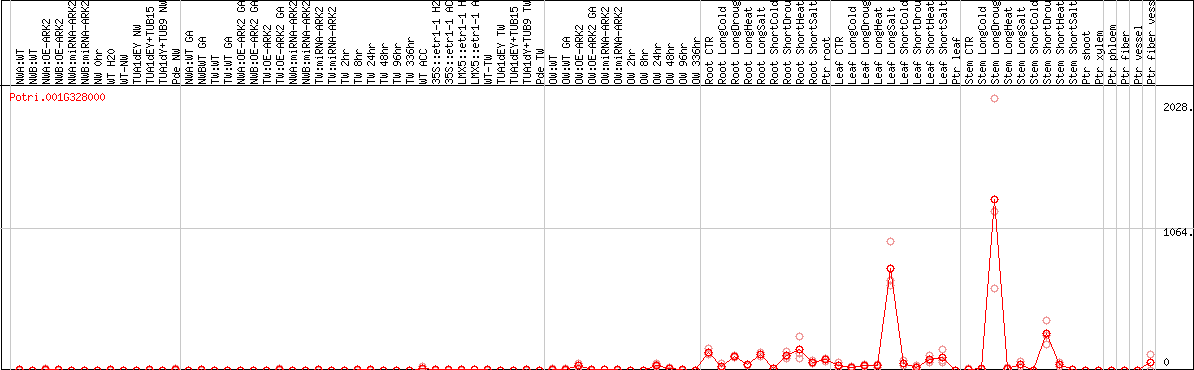
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G328000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.