External link
Symbol
GAPDH.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04120 585 / 0
GAPC1, GAPC-1, GAPC
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
AT1G13440 584 / 0
GAPC2, GAPC-2
GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE C-2, glyceraldehyde-3-phosphate dehydrogenase C2 (.1.2)
AT1G79530 484 / 4e-172
GAPCP-1
glyceraldehyde-3-phosphate dehydrogenase of plastid 1 (.1)
AT1G16300 484 / 4e-172
GAPCP-2
glyceraldehyde-3-phosphate dehydrogenase of plastid 2 (.1)
AT1G42970 270 / 1e-87
GAPB
glyceraldehyde-3-phosphate dehydrogenase B subunit (.1)
AT1G12900 249 / 1e-80
GAPA-2
glyceraldehyde 3-phosphate dehydrogenase A subunit 2 (.1.2.3.4)
AT3G26650 249 / 7e-80
GAPA-1, GAPA
GLYCERALDEHYDE 3-PHOSPHATE DEHYDROGENASE A SUBUNIT 1, glyceraldehyde 3-phosphate dehydrogenase A subunit (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G055400
602 / 0
AT3G04120 592 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Potri.012G094100
590 / 0
AT1G13440 580 / 0.0
GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE C-2, glyceraldehyde-3-phosphate dehydrogenase C2 (.1.2)
Potri.015G091400
587 / 0
AT1G13440 578 / 0.0
GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE C-2, glyceraldehyde-3-phosphate dehydrogenase C2 (.1.2)
Potri.008G179300
582 / 0
AT3G04120 573 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Potri.008G083900
483 / 2e-171
AT1G16300 634 / 0.0
glyceraldehyde-3-phosphate dehydrogenase of plastid 2 (.1)
Potri.010G172400
481 / 6e-171
AT1G16300 618 / 0.0
glyceraldehyde-3-phosphate dehydrogenase of plastid 2 (.1)
Potri.002G007100
256 / 2e-82
AT1G42970 690 / 0.0
glyceraldehyde-3-phosphate dehydrogenase B subunit (.1)
Potri.005G254100
255 / 9e-82
AT1G42970 732 / 0.0
glyceraldehyde-3-phosphate dehydrogenase B subunit (.1)
Potri.002G220566
243 / 1e-77
AT3G26650 614 / 0.0
GLYCERALDEHYDE 3-PHOSPHATE DEHYDROGENASE A SUBUNIT 1, glyceraldehyde 3-phosphate dehydrogenase A subunit (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014603
597 / 0
AT3G04120 634 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10032071
597 / 0
AT3G04120 634 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10022332
596 / 0
AT1G13440 632 / 0.0
GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE C-2, glyceraldehyde-3-phosphate dehydrogenase C2 (.1.2)
Lus10006435
582 / 0
AT3G04120 593 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10011375
582 / 0
AT3G04120 593 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10015826
573 / 0
AT3G04120 606 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10036976
556 / 0
AT3G04120 592 / 0.0
glyceraldehyde-3-phosphate dehydrogenase C subunit 1 (.1)
Lus10009602
482 / 4e-171
AT1G16300 694 / 0.0
glyceraldehyde-3-phosphate dehydrogenase of plastid 2 (.1)
Lus10000872
481 / 1e-170
AT1G16300 694 / 0.0
glyceraldehyde-3-phosphate dehydrogenase of plastid 2 (.1)
Lus10012243
258 / 8e-83
AT1G42970 759 / 0.0
glyceraldehyde-3-phosphate dehydrogenase B subunit (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00044
Gp_dh_N
Glyceraldehyde 3-phosphate dehydrogenase, NAD binding domain
CL0139
GADPH_aa-bio_dh
PF02800
Gp_dh_C
Glyceraldehyde 3-phosphate dehydrogenase, C-terminal domain
Representative CDS sequence
>Potri.001G335800.1 pacid=42791923 polypeptide=Potri.001G335800.1.p locus=Potri.001G335800 ID=Potri.001G335800.1.v4.1 annot-version=v4.1
ATGGCCAAGATTAAGATCGGTATCAACGGTTTTGGAAGGATCGGGCGTTTGGTGGCGAGAGTTGCTTTGCAGAGAGATGATGTTGAGCTTGTCGCTGTCA
ACGATCCTTTTATCACCACTGATTACATGACCTACATGTTCAAATATGACACTGTTCATGGGCAATGGAAGCACGGAGATATCAAGGTCAAGGATGAGAA
GACTCTTCTTTTTGGTGACAAGCCAGTCGCTGTTTTTGGTGTCAGAAACCCAGAGGAGATTCCATGGGGCGCTGCTGGTGCTGATTTTGTTGTTGAGTCC
ACTGGTGTTTTCACTGACAAGGACAAAGCTGCTGCTCATTTGAAGGGTGGTGCAAAGAAGGTTATTATTTCTGCTCCTAGTAAGGATGCTCCCATGTTTG
TTATGGGTGTCAATGAGAAGGAGTACAAGCCTGATCTCAACATTGTTTCCAATGCCAGCTGCACTACCAACTGCCTTGCTCCTTTGGCCAAGGTCATTCA
TGACAAGTTTGGTATTGTTGAGGGTCTTATGACTACTGTCCACTCCATTACTGCCACACAAAAGACTGTTGATGGACCATCAATGAAGGATTGGAGAGGT
GGTAGAGCTGCTTCCTTCAATATCATTCCCAGCAGCACTGGAGCTGCTAAGGCAGTAGGGAAAGTTTTGCCTGCACTCAATGGAAAGTTGACTGGAATGG
CTTTCCGTGTTCCCACAGTTGATGTCTCAGTGGTTGACCTCACTGTGAGAATTGAGAAGAAGGCTACATATGACCAGATCAAAGCAGCTATCAAGGAGGA
ATCTGAGACCAATCTGAAGGGAATCCTTGGTTACACTGAGGATGATGTTGTCTCAACCGACTTCATGGGTGACAGCAGGTCAAGCATATTTGATGCCAAG
GCTGGAATTGCATTGAATGATAACTTTGTGAAGCTTGTCTCCTGGTATGACAATGAATGGGGCTACAGCTCTCGCGTGATTGACTTGATCATTCATATCG
CCTCTGTGCAATAA
AA sequence
>Potri.001G335800.1 pacid=42791923 polypeptide=Potri.001G335800.1.p locus=Potri.001G335800 ID=Potri.001G335800.1.v4.1 annot-version=v4.1
MAKIKIGINGFGRIGRLVARVALQRDDVELVAVNDPFITTDYMTYMFKYDTVHGQWKHGDIKVKDEKTLLFGDKPVAVFGVRNPEEIPWGAAGADFVVES
TGVFTDKDKAAAHLKGGAKKVIISAPSKDAPMFVMGVNEKEYKPDLNIVSNASCTTNCLAPLAKVIHDKFGIVEGLMTTVHSITATQKTVDGPSMKDWRG
GRAASFNIIPSSTGAAKAVGKVLPALNGKLTGMAFRVPTVDVSVVDLTVRIEKKATYDQIKAAIKEESETNLKGILGYTEDDVVSTDFMGDSRSSIFDAK
AGIALNDNFVKLVSWYDNEWGYSSRVIDLIIHIASVQ
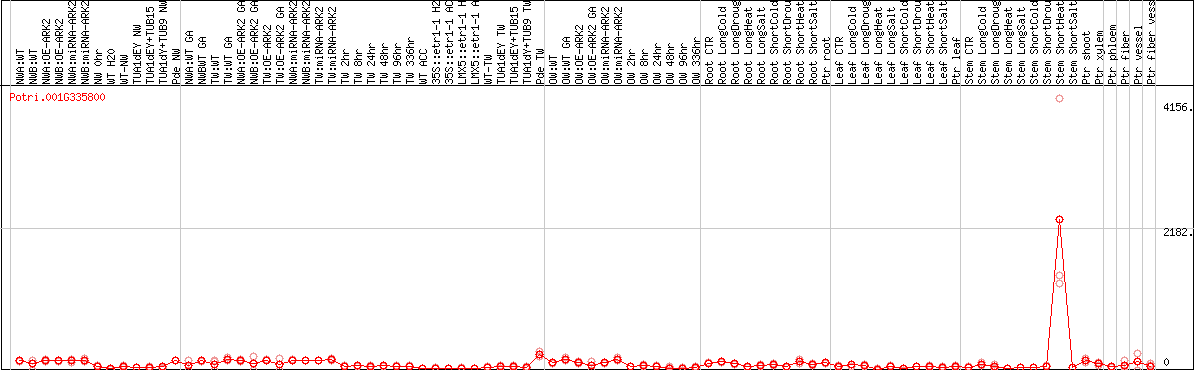
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G335800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.