Potri.001G345100 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G345100.1 pacid=42791539 polypeptide=Potri.001G345100.1.p locus=Potri.001G345100 ID=Potri.001G345100.1.v4.1 annot-version=v4.1
ATGAAGGTGCTGCGTTTTGCTGCGTTTTTGGTAGCTTTCACTACTCTCATTGTCGTCGGCAAAACGACGTCACATTTGAGCCCCGGACCTCACGTCACCG
ATGTCAATATTCTGTTGCCTCCTAAAATGACGCGTCCCGTCGAGTATCGCCTCCAAGGAAGTGATGGCTGCTTTAAATGGTCATGGGATCATCACGATAT
TCTATCAGTTATACCTGAGTATAATTTAAGCTACAGCCACTGTTCAACCAGTGCACGGTTGAGATCAATTGCTCCCTTTACCGGTCGGAAGGAGACAGCT
ATTTATGCTGCTGATGTACACACTGGCATTGTGATTCGCTGCAAGGTCTTTATTGACAATATTTCCAGAATCCAGATATTTCACAATTCTATTAAACTTG
ATTTAGATGGTCTTGCTACTCTCCGAGCTCGCGCCTTCGATCGTGAAGATAATGTGTTTTCGTCATTAGCTGGATTACAATTCATGTGGCAGCTGATGCC
TGAAACGGATGGATTGCCGCACCACCTTGTCCATGTTCCTCTCAGGGATTCTCCGCTCAGTGATTGTGGTGGTTTATGTGGTGATTTGAACATCCAAATA
GAACTTGAAGATAGTGGTGTATTCTCGGATTTGTATGTTGTGAAAGGAGTCGAAATTGGCCATGAGAATGTTTCGGTGCATTTGCTTGAGCCACAATTTA
AGCACTTGGCTGATAAGATTGTTCTAACTGTGGCAGAAGCCATGTCACTTGAACCTCCCTCACCTGTTTTGGTCCTCATTGGTGCTGCCTTTCGATATAC
GCTCAAAGTCATTCGTGGGAATATTCTGCAAGTGGTTGCTCTTCCATCTCCACATCATAGGTGGTCTGTTTTGAACTCTTCGGTTGCTGAAGTGGACCAC
TTGTCAGGCTTTGCTCAGGCACTGAGCTTAGGGGTAACATCTGTCATTGTCAAAGATACTAGAGTTGCTGGCCATATGCAAGTTTCATCACTTAATGTGG
TTCTACCTGACACTTTGTGCTTATTTATAATGCCCTTGCCTGTTTCTGGTGATCCTGTTGATGGATTGAAAGCCATTCCATCCTTGGCACGTTGGTTTGT
TGTTTCTGGTCGTCAATATATTATTCAAATGAAGGTTTTCTTGGGGGGACCTGATGCGCAAGAAATCTATATTACAGAGAGTGATGACCTTAAGTTGCAC
CATGAGCAGTCTGAATACTGGACAATTTTCATGTTATCAGAGGATATTGTGGTCAAACATGGCTGGCGAAATTCCAGAATTCTGAGAGCAATTTCCCTTG
GTCAAGGAAAACTAACAGCTTCTTTAACTTATTTTAGTGGGCACCGGGAGAGAAAAAAGGTTCTTAGCGCCGCGCAAGAAATTATGGTATGTGATCAAGT
GAAGTTCAGTTTGGATGGAGCAAGCGGTACACATCAGACTATTCTTCTTCCCTGGACTCCTACTATTTACCAGGAGGTGGAGCTAAAGGCAACAGGAGGC
TGTGCAAAAACATCCAGTGATTATAAATGGTTTTCTTCAGACACGTCTACTGTGTCAATATCTCCATTTGGGGTACTTCAGGCAAAAAAGCCTGGTAAAG
CGACTGTCAGAGTAGTGTCCATTTTTGATTCTTCTAATCATGACGAGGTGGTCATTGAAGTTTCTGTACCATCATCTATGATTATGCTTCAAAACTTCCC
TGTAGAGACTGTTGTAAGATCACATCTTCCAGCTGCCGTAACAATGAAAGCATCAGATGGTTCCTATTATTATACATGTGATGCTTTTCATCTATTCGTA
AAATGGAAAGTTGGAAGTGAGTCTTTCGTCATTGTCAATGCAACAGAGGAGGCACGTATTGCAGAAAAACTGGGGGCTTTTCACCTCTATGAATCAGTTT
ATGGTCCACCATGTTCATGGGCCTATATATATGCTTCCGGTCCTGGGCAGACCATGCTGCAGGCAACATTGTCGAAAGAACATCACTTGGATCAATCGTT
TCATGGGTCCATTATTTTGAAGGCATCATCACGTATTGCAGCATACCCACCACTTACAGTGCATCAGGTAGTAGATGGAAGCCAATTTGGTGGGTATTGG
CTTGATTTGGGTCATGTTGAAGCAAGCAATCACCTGGAGCCTTTGGAGAGCTTGTATCTTGTCCCTGGAACAAGTTTAGATATCATGCTTCTTGGTGGAC
CAGCACCATGGGACAAAGATGTTGACTATCTTGAAACTGTGGAAATATCAGGTGACAAGCATGCTTCCAGTAAAGATGGAATTCATGCACGTCGGATTTC
TGGCAGCTATCAGAGTATGTACAGAGTTTCATGCCAAACTCTGGGGATATTTGATTTGGTTTTCAAACGCGGGAATTTGGTTGGAGATGACCACTCTCTG
CCTGCAATAGCAGAAGTTGCTTTGTCTCTTACATGCAGCTTTCCATCTTCAATTGCACTTATCGTTGATGAACCCGTTAACAGACATGATGCCATAAGGT
TCGCATCTCTTGCTGATCGTAGCACAGGGCAAATCCGCACCACCCCTGTTACTGTGGCAAATGGGCAAACTATTAGGGTGGCTGCAGTTGGCATAGGTGT
TTCTGGAGAAGCTTTTGCAAACTCATCTTCTCTTTCTCTGAGGTGGGAGTTGAGCAGCTGTGAAGGACTGGCATATTGGTGGGCTGATGCTTATGAGTCG
CAGAGATTGAAATCCAGCTGGGAGAGGTTCTTAGTCTTGCAAAATGAATCAGGAGAATGTATTGTTCGTGCCACTGTTATTGGTCTTGATGATGCCCTCG
GTAGTCATTATTTGGCTCAGTTGCGAAGTCTAGAGAATGTTCTTACAGATGCTGTAAAGTTGCAGCTTGTTTCTTCTTTGAGAGTGAATCCAGAGTTCAA
CTTGTTGTATTTCAATCCCGATGCTAAGGTAAATCTTTCAATTGCTGGAGGGAGCTGTTTATGGGAAGCTGTTGTGAATGATTCTCAAGTGGTAGAAGTT
GCTCAGCCCCCACCAGGCTTACAATGCTTTCAGTTGAGGCTCCTTCCTAAAATGTTGGGGACTGCACTTGTGTCAATATCTGATATTGGGCTTATTCCTC
CTACTTCAGCTTCTGCTGTGGTTCAAGTTGCTGATGTGGACTGGATTAAAATAGTATCAGGAGAACAAATAAGTCTCATGGAAGGACAGTCACAGCCCAT
ACATTTAATGGCTGGGATTAAGGATGGGAATGCTTTTGACTCTAATCAGTATGCGTACATGAAAATTCATGTACACATCGAGGATCCTATAGTTGAACTT
GTTGACAAGAATGGTATGCCTAGTGATGCCGGTGGATATGTTAATGCTCCAAACTTTACGATCAATGCTAAAGATCTTGGATTCACAACACTTTATGTTA
GTGTTAGACAGCAATCTGGACATGAAATATTGAGTCAATCAATAAAGATAGAAGTATATGCGCCCCTGAGAATTCATCCAGATGATATTTTCCTGGTACC
AGGTGCTTGTTACATGCTTGCTATGAAAGGGGGCCCAACAGTTGGCGTATATATTCAGTATGCTTCAATGGATGATGGAATAGCAACAGTGGACAAATCT
TCAGGACGACTTTGTGCAATTTCTCCTGGAAACACTACAATCCTTTCCTCGGTGTTTGGGAATGGAGGTGTTGTGGTATGTCAAGCACATGGTAGCATTT
ATGTTGGAGTTCCTTCTTTGTCAATGTTGAGTGCACAAAGCGATAAGCTTGATGTCGGCCGTGAGATGCCGATATATCCATCATTTCCTGAGGGTGATTT
ATTTTCATTTTATGAGCTCTGCAGAAGTTACAAGTGGACCATAGAGGATGAAAAGGTGTTGAGTTTCAACATGGCAGAAAGCTCAAATGTTGAAAAGCAC
TGGTTCCCATTGGATGATGAGCAGGAACTTGATTTTATTAAAGTCTTGCATGGAAGATCTGCAGGCAAGGCAAATATCACAGTTACATTCTCTTGTGATT
TCGTTTCCGCTTCGTTCTCACAGTCGAGGCTATATGATGCTTCTTTATCACTATTGGTTGTGCCTCCTCTCCCTCTTGCTCTTGGAGTTCCAATGACATG
GCTTCTTCCTCCTCATTATGTTACATCTAGCCTTTTGCCTTCATCTTTGGAATCGCATGGTCAACAAGATGCTCAGAGTCGTAGAGGAACAATTATCTAT
TCGTTATTAAGTTGTGAAAAGAATGAAGTTTGGAAAAAAAATGCAATTTCTATAGATGGAGATAGAATAAAAACAACTGAGAGTAACAATCTTGCATGCA
TACAAGCCAAAGATCGAACCACTGGAAGAACTGAGATTGCATCTTGTGTCAAGGTCGCTGAGGTGGCTCAAATAAGAATAATGAATAGGGACCCCCCTTT
CCATGTAATTTATCTCGCTGTTGGTGCCAACCTTGATCTTCCAATAAGCTATTTTGATGCTTCAGGGAATCCCTTTTATGAAGCATATGATGTCATTTCC
TATCATGCTGAAACCAACTATCATGAAATTGTCTCCGTTGTGCATACAAGAAATGGCAATGGGACTATCCATCTGAAGGCAATGCAAACTGGTAGAGCTC
TGGTGCGAGTATCAATGAACAGTAATCCACGAAAATTTGACTATATACTGGTTTCAGTTGGTGCTCACATATATCCCCAAAACCCTGTCCTTCAACATGG
GAGTTCTCTTAACCTCAGTGTAATAGGCATTAATGATCAAGTTTCTGGCTGCTGGCATAGCGCTGATGAAAGAGTTGTTTCTGTAGACATGCGATCTGGA
AAAATTGAAGCATGTGGAATAGGTTCAACCCATGTATTTTTCAAATGCCCAAGCATGAAATTGCAAACAGAAATCACAGTGCTATCAGGAAATATTGTTT
CTATTCATGCCCCAAAAGAGATCCTAACAAACATTCCTTACCCAGCAAAAGGATATAGCTTCCCTTTGAGCTTAAGTGATACTTACAACAAGCTTGAAAC
TCTTGGAAACAGTAAGGGAGTATCATATGACTGCAAGGTTGATCCTCCTTTTGTGGGGTATGCAAAGCCATGGATGGATCTTGAATCTGGGAATTCCTAT
TGCCTGTTCTTCCCATATTCACCAGAGCACTTGGTGCATTCTGTACCCAGGTTGAAGGACATGAGACCATATGTATCTATTTCTATAAACGTATCACTCA
GAGAAGCCAGCCATGTCTCTGGATCTGCATCAGCAATTTTTATTGGAGGGTTTTCAATATTAGAAATGGACAAGAGTCCAATGCAACTGAACCTGACACC
AGATTCCAACAAGACTACTATAACCATCTTGGGAAATACAGATGTTGAAATCCACTGGCTTGATCGGGATTCAATGAAGATCGCTCTTGTCCACAAGGAA
GATTTCGGGATAGGTAGCCATGCACAATATGAGGTCGAAGTGTTGAGAGCCAAAAGATTGAAAGATAGAATCACCATCACTCTCCCAGCCAATGGTCAGA
GAATGGAAATACAAGTTACTTATGAACCTGATGCAAAAACGGGGGCAAAAACTAGCTTTGCAACCTTCATTGGGCTACTAGTAGCTGGATGTGTGATCCT
TTGTGTAATTTATCTAGGCTGTTCCTTGCTTATCTCTGAGATATCCAGCATACATGTGCCCTCTACCACCCCTGCCACTCCACCTAGCGCAGCACCACAG
ACACCTGCTCGTGGCAGTCCTGTCTTGACTGAACAGTCGCCTCGAACACCTCAGCCATTTGTAGATTATGTAAGGAAGACAATAGATGAAACTCCCTACT
ACAAGCGAGAAGTTAGGAGGAGGTCTAATCCACAGAATACTTACTAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.001G345100.1 pacid=42791539 polypeptide=Potri.001G345100.1.p locus=Potri.001G345100 ID=Potri.001G345100.1.v4.1 annot-version=v4.1
MKVLRFAAFLVAFTTLIVVGKTTSHLSPGPHVTDVNILLPPKMTRPVEYRLQGSDGCFKWSWDHHDILSVIPEYNLSYSHCSTSARLRSIAPFTGRKETA
IYAADVHTGIVIRCKVFIDNISRIQIFHNSIKLDLDGLATLRARAFDREDNVFSSLAGLQFMWQLMPETDGLPHHLVHVPLRDSPLSDCGGLCGDLNIQI
ELEDSGVFSDLYVVKGVEIGHENVSVHLLEPQFKHLADKIVLTVAEAMSLEPPSPVLVLIGAAFRYTLKVIRGNILQVVALPSPHHRWSVLNSSVAEVDH
LSGFAQALSLGVTSVIVKDTRVAGHMQVSSLNVVLPDTLCLFIMPLPVSGDPVDGLKAIPSLARWFVVSGRQYIIQMKVFLGGPDAQEIYITESDDLKLH
HEQSEYWTIFMLSEDIVVKHGWRNSRILRAISLGQGKLTASLTYFSGHRERKKVLSAAQEIMVCDQVKFSLDGASGTHQTILLPWTPTIYQEVELKATGG
CAKTSSDYKWFSSDTSTVSISPFGVLQAKKPGKATVRVVSIFDSSNHDEVVIEVSVPSSMIMLQNFPVETVVRSHLPAAVTMKASDGSYYYTCDAFHLFV
KWKVGSESFVIVNATEEARIAEKLGAFHLYESVYGPPCSWAYIYASGPGQTMLQATLSKEHHLDQSFHGSIILKASSRIAAYPPLTVHQVVDGSQFGGYW
LDLGHVEASNHLEPLESLYLVPGTSLDIMLLGGPAPWDKDVDYLETVEISGDKHASSKDGIHARRISGSYQSMYRVSCQTLGIFDLVFKRGNLVGDDHSL
PAIAEVALSLTCSFPSSIALIVDEPVNRHDAIRFASLADRSTGQIRTTPVTVANGQTIRVAAVGIGVSGEAFANSSSLSLRWELSSCEGLAYWWADAYES
QRLKSSWERFLVLQNESGECIVRATVIGLDDALGSHYLAQLRSLENVLTDAVKLQLVSSLRVNPEFNLLYFNPDAKVNLSIAGGSCLWEAVVNDSQVVEV
AQPPPGLQCFQLRLLPKMLGTALVSISDIGLIPPTSASAVVQVADVDWIKIVSGEQISLMEGQSQPIHLMAGIKDGNAFDSNQYAYMKIHVHIEDPIVEL
VDKNGMPSDAGGYVNAPNFTINAKDLGFTTLYVSVRQQSGHEILSQSIKIEVYAPLRIHPDDIFLVPGACYMLAMKGGPTVGVYIQYASMDDGIATVDKS
SGRLCAISPGNTTILSSVFGNGGVVVCQAHGSIYVGVPSLSMLSAQSDKLDVGREMPIYPSFPEGDLFSFYELCRSYKWTIEDEKVLSFNMAESSNVEKH
WFPLDDEQELDFIKVLHGRSAGKANITVTFSCDFVSASFSQSRLYDASLSLLVVPPLPLALGVPMTWLLPPHYVTSSLLPSSLESHGQQDAQSRRGTIIY
SLLSCEKNEVWKKNAISIDGDRIKTTESNNLACIQAKDRTTGRTEIASCVKVAEVAQIRIMNRDPPFHVIYLAVGANLDLPISYFDASGNPFYEAYDVIS
YHAETNYHEIVSVVHTRNGNGTIHLKAMQTGRALVRVSMNSNPRKFDYILVSVGAHIYPQNPVLQHGSSLNLSVIGINDQVSGCWHSADERVVSVDMRSG
KIEACGIGSTHVFFKCPSMKLQTEITVLSGNIVSIHAPKEILTNIPYPAKGYSFPLSLSDTYNKLETLGNSKGVSYDCKVDPPFVGYAKPWMDLESGNSY
CLFFPYSPEHLVHSVPRLKDMRPYVSISINVSLREASHVSGSASAIFIGGFSILEMDKSPMQLNLTPDSNKTTITILGNTDVEIHWLDRDSMKIALVHKE
DFGIGSHAQYEVEVLRAKRLKDRITITLPANGQRMEIQVTYEPDAKTGAKTSFATFIGLLVAGCVILCVIYLGCSLLISEISSIHVPSTTPATPPSAAPQ
TPARGSPVLTEQSPRTPQPFVDYVRKTIDETPYYKREVRRRSNPQNTY
|
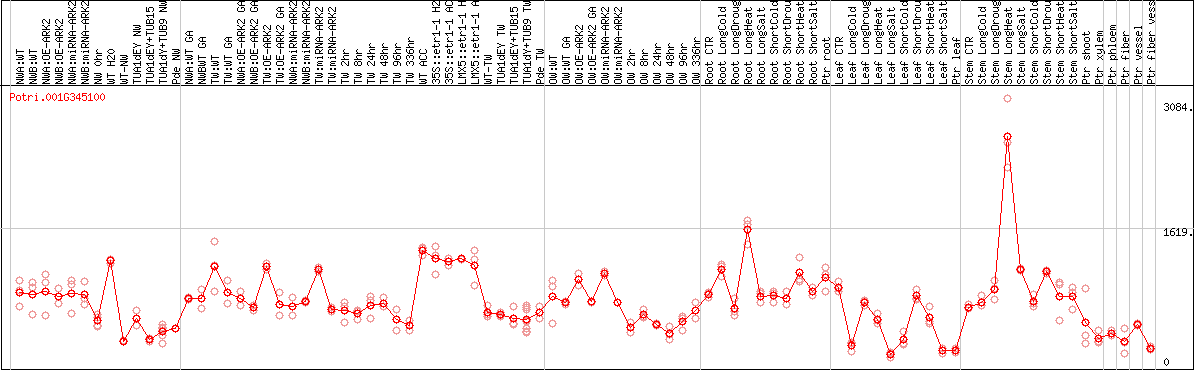
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G345100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
