Potri.001G346000 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G346000.1 pacid=42789513 polypeptide=Potri.001G346000.1.p locus=Potri.001G346000 ID=Potri.001G346000.1.v4.1 annot-version=v4.1
ATGGACTCTTACACTCCTTTACTCCAGAAAACACGGGTTCCACAACCCTCGCTCCAAAAATTCGCGGTGATTTCAATCTTCTCGAAGCTACGATCCGCGC
CGAGTTACCTCGACCCGGATTCCGAACCCGGTAGAGAAGCTATCTCTCAATGCCTCCGCTCCGCTTCTCCATCCGTCGTCGACCAGTCTGTACGCGAGCT
CTGTCGCCTCCTATTGGACTCCAGATTGGACCTCTCCCGAACTTTGCTGGAGCTCCAGTCTGCACTCGAAGGGTCCGACCCAAAATTTGTTGGTTTATTT
GTTAAAGCTCTTGGATTTGTCGTTCGGGTCGGGTTTCAAAGGAATCATGGGTCGTGGCGTTTTTCTTCTATTGAGAATCACCCCTTTGTTATGATTTTGT
CATCAAGAACTGAGGTTCAGTCTGAATTGGTTCAGCAAGTCTTACTATTTTTCGGACAGAATAGAAGATCGGGAATGGTGGAAATTTGTGAATTCTTGAG
ACCCTTTTTGAATTTTTCGATTCTACGCGTGCCCTTTTCGAATTCTTCGTCATCTTTGTTTGCAAGGCAGTTGATATCATCAATGGCTTCGTTTTGTTGT
TCTTTCCCTGATGAGGCGATTCCGGTTTTAAAGCTACTGATAGGTTGCCTTAAGCATGCGTCACATAAGAATTCAGATGAGCTTAAGAACTCATATTATT
TTCTGGAATCCATAGTTGATGCATATACGGTCGTTTTGAGACACTTGGTTGGAACCGGACTGCTGGTTACTGAGGCTCAACTGTTTGGTGTGGAACTGTC
AGATGCTATTCTTTCATTATTGACATGTCATCACCGGCATGCTGGTAGTAGTGAACCAATTGTTGAACTAGTGAAGCGGCTGTTTGTTGCCCAGAAAAAT
CTTTCTTTGCAATATATGCCTGAAATATCATCAACACTGCTATCCTTATTTGTTGTCCTGATTCAATCTGATCTTGAATACGAACAACTCTCTTTATTGA
AACTCCTGAACTTCCTTTTAAAATGGAAAAGTGAAAAGGAATATGAGGTTGATAGAGTCAAATGTGCTACGAGCGAAGAGCTTTTGTTCACATTTCCTGT
CATCAACCTATTATCATCTACATCTAGATCTATTAAAGGAGAAGCAGCAGAATTGCTTGTGACGCTGGAAAAGGTCCTAGTAGAACTGTCAAAAGCACCA
AAGGCTGGACTAGCCAAAGAAGGGGGATTCCCCCCTATCAGCTCTCTAGGATCGATTGCTTATAGGCTTTTGCGATGCTTGTGGTTTCAGGACCAGTTTT
TACTGCCCACTTCCTTCCTGAATTTTGCTTCCAGTGGCAAAACGGATGTCAAAGTAATGCACCAGAAACCAAGACATTGGGCTTCTCAGTTAAGAGAATA
CATTTTGTCGATTGTAGACAGAAGAAAATCCTCCCTATCTGTTTCTCAGTCTCAAGAACGTTTCACAAGGGAACTGCCCCCTTTACTCGGTGCCATTACT
GGTGTTTTGGTGATGCACCGGTCATTTGGAGATACGGCAATTGATTTGCTGGGTGCTATTGGCATTGTGGACCCCAAACAGGGAGTGCCACTGCTGCTAG
CAATTCTATTTTATAGTAACATCTTCACCAGTAAAGACATTAGTTACCAGAATATGTTGCCAAAACTTCTGGCATTACTTCCATCACTTGCTTCACATTC
TGTGATGATTCCCCTCATAATTCAGACAATCTTGCCGATGCTTCAGAAAGATGGAAAACCTGTGTTGTATGCCACCGGCGCTCGTTTGCTTTGTCAGACC
TGGGCAATCAATGACCGTGCCTTTGGGAGTTTGCAAGCAATATTGCTTCCGAAGGGGCTCACTGAATTCAAGCATGAGAGAAACATCTTAATAAGTTTGG
CTGCCTCCATTCGGGACATTTGCAGAAAGAACCCTGATAGGGGGGTGGATCTTATCTTATCTGTTTCGGCTTGCATTGAGAGCCAAGATCATATTATTAA
AGCTCTTGGATTTCAGAGCCTTGCACATCTATGTGAAGCTGATGTAATTGATTTTTATACTGCCTGGGATGTCATTGGTAAGCATGCGGTAGACTACACC
ACAGACCCGGCTCTTGCACAAAGCATATGCCTCCTTCTGAGATGGGGTGCAATGGATGCTGAAGCATATTCAGAAGCTTCAAGGAATGTCTTGCAAATTT
TATGGGGTATTGGAACTGCTGTACATGTCAGTCATGCACTTGAGTGGGCAAGGGCAAGGATCTTTGCCTTTGAGGCCCTGAGCCAATATGAGGTTACACA
TATACAAATAGGAATTCCAGATTTTAAGACAGTGAACACGGATTTGCTTTTACGTGAGACGAATCTTGATGTGCTCACGGCTATGGAAGGATTTCAAGTC
AAGATCATAACTCATGAACATGTGAATCGTCGAAGGTTGGTGAAGGAGAAAAAAATTGCTGGAAGTAAGATTGAGAAGTTGCTAAATGTATTTCCACAGG
TTCTCGTCTCAGGGATAAAAGGTAGTGCTGGGCAATTACCTGGGGCAGCTCTGTTATGCCTTTCCTTTACCCCTAAAGATGTGAACAGCCAATGCTTGTC
AAGAGTATCTGTAGATTTCCATGCTGGATATGAGAGTGCACTGGTGGAAATAGCTGCTTCTCTTCAATTATCAAGAAATATTTTCACTGCCCTTCTTTCA
TTGCAATCTTGGAAGTCCTTCATGCGGCGTTGGATAAGGGCAAATATTTCATCCCTTGATGCTAAAGCACCATCTGTTTCATTAGATAAAACTTCTAAAG
CTGCTACCGATATCCTGAAGCGTGTGATGCGACTTGCTGAAGAGTCTATCCCAAGTTCTGCTGAAAATATTGCTCTGGCTATTGGTGCACTCTGTGTGGT
GTTGGCTCCATCTACCCATACTGTCAAATCAACTGCTTCAAAGTTTCTGCTAAACTGGTTGTTCCAGAATGAGCATGATCATCGCCAATGGTCAGCCGCA
ATATCTCTTGGATTGGTTTCAAGTTGTCTTCATGTAACTGATCACAAACAGAAATTTGAGAATATCACTGGACTTATAAAGGTTTTGCATGGCAGCAAAA
GCATCCTCGTCAAAGGAGCTTGTGGGCTTGGCTTGGGCTTTGCATGCCAAGACCTTCTTACCAGGTTTGAAGCTGCTGATAATGTTGATTTGGATAAAGA
AAAGTACAAGGCACAGGAGGTTGACTTATTAGGGAAGATTCTTAGGACACTACTTCTGATGACAAGCCAGCTCAGTAATGCTTCATATGATATTCTAGAA
AGTCTTCCTCCATTTTTCAGCATGGGTGCTAATGATATGGAAATCAATTTGACATCTGACCAGTTGCTTGAGAAGTGTGATGACTTGGAGGAAGATCCTT
GGGGTGTTGCTGGTCTTGTCCTTGGCTTGGGGATCTCTTTCAGTGCAATATATAGAGCTGGGGCACATGATGCTATGCTGAAGATAAAGGACTTGATTAT
ATCATGGATCCCACATGTAAACTCTCTAGTAACGAACTCATCTTTTTCCAGTGAAGGAAGGGAGAAAGCTTTATCAGTAGGATCCTGTCTTGCACTTCCC
AGTGTTGTGGCATTCTGCCGGAGAGTGGAAATGATCAATGATAATGAGCTGGATCAACTATTAAAGGGCTATCATGAGCTCATATCTGAGTTATTATCTG
TTAAAAAATCTGGTACTTTCCATCAGAGCTTGATGTTGGCATCATGCATTGGTGCCGGAAGTCTTATCGCTTGCATTTTGAATGAAGGTGTGCACCCTCT
TGAAGCTGAATTTGTAAAAGGTTTACTGGAGATGTTTAGAAAATGCTACTGTAGTTCTTTCCCTCCTATCATCCATTTGGGTGGGATGCTTGGAGTTGTT
AATGCAATGGGAGCTGGTGCAGGGATTTTGGTTCATGCCCATCACTTCTCAGCCTCAATCAAGACTGCTTGTGAACAAAAGGAATCTTCTCATATTTTGG
GCCCTTTACTTTCAAGCCCTTTTTGTGAGCCACATTTGACAACATTGGTGCAAGAGATATTTCTCATTGCGCAAAATTCTGATGATCTTAAAATGCAACA
AAATGCAGCTTGGGCCGTTTCATTCCTTCGAAATGGTTTGTGGTCAAAGGAACTTCTAAATGCTGAAAGCAATGATCAGACTGATGTTGTTGATAGCAAA
ACAATTTCTCACAATTTTCCTGAGGACAATTTAGTTATGAAGCTGACTATATGGTTAATGCATTTAAATAACTCCGGGGCAGGGGCTATTGCACATGTGG
GCACTGTTGTAACAGTTTTAAGATGCCTTTCCCGAGCTCCTAGATTGCCAACAGTGGATTGGGGATTGATCATTAGACGCTGCATGAGATATGAGGCTCA
AGTGTCTGAGGTGTTGTTGCCAGATTCAGCTCTTAAGAGGGGAGCCCTTCGGGAGGAATGTGTACAATTCTCAATAGCTCATGCGAATCAATTTGATCCC
CTCCTGACTTTCCTTGACGAGCTATCTGATTTGACTAGGTTCAGGACACTAGAGCTGAATCTACAGTCATGTCTGCTCTTTCATCTTGCAGGCCTGATAA
AAGTATTCTCAGGTTCCAGGCTAGAGAAACTGCTTGATGATATTGCTGAATACTTTGGCTCAGATATTTTATACCAAGGGTATAGCTCTGATCAGAAGAG
CTCATTGAGAATCTCATGCTGGGTGGGTCTCTATCAGTGTCTAGAAGAAGCTGTCCTTAGCTCTGTAGAGTACATATCAAACCTTGAAAAGTGCATAGAG
GTGCTGTTTCACTTGTTGCCTGCATCAGAATCTGCTGCCTTCACAGGAGTGGATCTGCCAAACGCTGCAGAGGAATGGCGTGTGGCAGTTCAATGCTTGG
CAAAGGCTCAAGGGGATTGGTTATTGGATTTCCTGCAGGTTCCATTAGGGGATCTTGTGCAAGGAGGTAGCCAGTCAAATGAAGTCCTAAAGAAGATTCT
AGCGAAAGTGAAGCTAGTTAGGATGGGTTCCATCCCATTAACCGAGCTTGGAAGATTAAAAGCTTATATGCTGAACTCTAAATCAAAGGATATTTGGAAT
TTGCATGCTGAAGTGGTAGCAGCTTTGCAATATGCAGATGGTAGTGTCAAAAGACAGTGGCTTGTTGATGCAGTGGAGATTAGTTGTGTCTCGAGCTATC
CTTCCATAGCCTTAAAGTTTCTCGGTCTGCTATCTGGTAGCTGTTGCAAATATGGGTCTCTTCTGACTTTGGACCAGCTATCCGTGTTGAGTGATCTACC
AGTTACCCTCCCTTCCCTTGTGACCGAACCTAGTTGGGAAGTTGTTGCGGAATCCATTGTTTCTACCCTTTGGACATCAACCGAACGCATATACTACTTG
GTTACTGACAAAGGACCCCCTGACAATACAAATAGCACGCAACCTATTGACGGAAGTGAAAAGGATATTGCCTCCTTTCTATTGCATGTGATGTATCATA
CCTGTACATGTTTAAAAGAGTATTTGCCTTTGGAGAAGCAGCTTAGGCTTGCCAATATGCTGGTCACTTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G346000.1 pacid=42789513 polypeptide=Potri.001G346000.1.p locus=Potri.001G346000 ID=Potri.001G346000.1.v4.1 annot-version=v4.1
MDSYTPLLQKTRVPQPSLQKFAVISIFSKLRSAPSYLDPDSEPGREAISQCLRSASPSVVDQSVRELCRLLLDSRLDLSRTLLELQSALEGSDPKFVGLF
VKALGFVVRVGFQRNHGSWRFSSIENHPFVMILSSRTEVQSELVQQVLLFFGQNRRSGMVEICEFLRPFLNFSILRVPFSNSSSSLFARQLISSMASFCC
SFPDEAIPVLKLLIGCLKHASHKNSDELKNSYYFLESIVDAYTVVLRHLVGTGLLVTEAQLFGVELSDAILSLLTCHHRHAGSSEPIVELVKRLFVAQKN
LSLQYMPEISSTLLSLFVVLIQSDLEYEQLSLLKLLNFLLKWKSEKEYEVDRVKCATSEELLFTFPVINLLSSTSRSIKGEAAELLVTLEKVLVELSKAP
KAGLAKEGGFPPISSLGSIAYRLLRCLWFQDQFLLPTSFLNFASSGKTDVKVMHQKPRHWASQLREYILSIVDRRKSSLSVSQSQERFTRELPPLLGAIT
GVLVMHRSFGDTAIDLLGAIGIVDPKQGVPLLLAILFYSNIFTSKDISYQNMLPKLLALLPSLASHSVMIPLIIQTILPMLQKDGKPVLYATGARLLCQT
WAINDRAFGSLQAILLPKGLTEFKHERNILISLAASIRDICRKNPDRGVDLILSVSACIESQDHIIKALGFQSLAHLCEADVIDFYTAWDVIGKHAVDYT
TDPALAQSICLLLRWGAMDAEAYSEASRNVLQILWGIGTAVHVSHALEWARARIFAFEALSQYEVTHIQIGIPDFKTVNTDLLLRETNLDVLTAMEGFQV
KIITHEHVNRRRLVKEKKIAGSKIEKLLNVFPQVLVSGIKGSAGQLPGAALLCLSFTPKDVNSQCLSRVSVDFHAGYESALVEIAASLQLSRNIFTALLS
LQSWKSFMRRWIRANISSLDAKAPSVSLDKTSKAATDILKRVMRLAEESIPSSAENIALAIGALCVVLAPSTHTVKSTASKFLLNWLFQNEHDHRQWSAA
ISLGLVSSCLHVTDHKQKFENITGLIKVLHGSKSILVKGACGLGLGFACQDLLTRFEAADNVDLDKEKYKAQEVDLLGKILRTLLLMTSQLSNASYDILE
SLPPFFSMGANDMEINLTSDQLLEKCDDLEEDPWGVAGLVLGLGISFSAIYRAGAHDAMLKIKDLIISWIPHVNSLVTNSSFSSEGREKALSVGSCLALP
SVVAFCRRVEMINDNELDQLLKGYHELISELLSVKKSGTFHQSLMLASCIGAGSLIACILNEGVHPLEAEFVKGLLEMFRKCYCSSFPPIIHLGGMLGVV
NAMGAGAGILVHAHHFSASIKTACEQKESSHILGPLLSSPFCEPHLTTLVQEIFLIAQNSDDLKMQQNAAWAVSFLRNGLWSKELLNAESNDQTDVVDSK
TISHNFPEDNLVMKLTIWLMHLNNSGAGAIAHVGTVVTVLRCLSRAPRLPTVDWGLIIRRCMRYEAQVSEVLLPDSALKRGALREECVQFSIAHANQFDP
LLTFLDELSDLTRFRTLELNLQSCLLFHLAGLIKVFSGSRLEKLLDDIAEYFGSDILYQGYSSDQKSSLRISCWVGLYQCLEEAVLSSVEYISNLEKCIE
VLFHLLPASESAAFTGVDLPNAAEEWRVAVQCLAKAQGDWLLDFLQVPLGDLVQGGSQSNEVLKKILAKVKLVRMGSIPLTELGRLKAYMLNSKSKDIWN
LHAEVVAALQYADGSVKRQWLVDAVEISCVSSYPSIALKFLGLLSGSCCKYGSLLTLDQLSVLSDLPVTLPSLVTEPSWEVVAESIVSTLWTSTERIYYL
VTDKGPPDNTNSTQPIDGSEKDIASFLLHVMYHTCTCLKEYLPLEKQLRLANMLVT
|
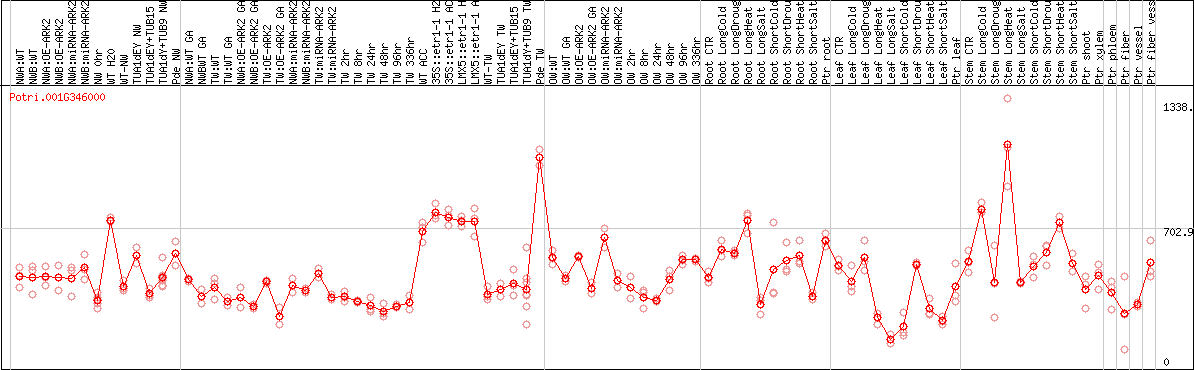
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G346000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
