Potri.001G350200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G350200.12 pacid=42791063 polypeptide=Potri.001G350200.12.p locus=Potri.001G350200 ID=Potri.001G350200.12.v4.1 annot-version=v4.1
ATGAAGAGAATGAGAGATGATATTTTCTCAGCCCCAGCCCCAGCCCCTGCCCCTGCCCCTGAATTTAAACGGCCCTTAACTTCGACACGTGGAGAATCCT
ATGGGCAAACACAAATCCCTGGCGGCGGTGGTGGCGGAGGTAGTGGTAATTCGCAGAAACTAACTACCACCGACGCCTTACAATATCTAAAGGAAGTAAA
GGACATGTTTCAAGACCAGAAGGAAAAATATGACATGTTCCTCGAAGTCATGAAAGATTTCAAGGCTCAAAGAACTGACACTTCCGGTGTCATTGTAAGA
GTAAAGGAGTTATTCAAAGGTCATAATAACTTGATATTTGGATTCAACACCTTCTTGCCCAAGGGATATGAAATAACCCTAGATGAGGATGAGGCAGCTC
CGCCAAAGAAAACTGTTGAATTTAACCAAGCTATCAATTTTGTAAACAAAATAAAGAAACGTTTCCAAAATGATGAACGTGTTTATAAATCATTTCTAGA
CATTTTGAATATGTACCGGAAGGAACACAAGGACATAAATGAGGTCTACAGTGAGGTTTCAGCTCTTTTTGAGGATCATCATGATTTGCTTGATGAGTTT
GCTAGATTTTTGCCAGATACTTCAGCAACACCCATTACACATACTGTTCCATATGCTCGGAATTCTAACCAGCATTACAATGAACGGAATTCTACTGCAC
CCTCCGCACGACAAACACAAATAGACAAGCAGCGTCGGCGTGACAAAGTTTCTTCTTCCCATGCTGAGCGTGACCTTAGTGTTGATCGACCAGAGATGGA
AGATGACAAAGGAATGGTGAAAGTGCATAAGGAGCAGAGAAAACGTGCTGACAAGGAGAATTGGGATAGGAGAATTCATGATCAGGATGATAGGGAGCCT
GAGCATGATAGCAGTAGAGACTTCAGCTTGCAACCTTTTCTTGAGAAAAGGAATTCTTCCCAGAAGGTGGAAGGTTTTGGAAATTCTAACTTTGCTTGTT
ATGATGACAAAGACAACATAAAGAGCATCTACAACCAAGAATTTGTTTTCTTTGAGAAAGTCAAGGAGAAGTTGGGAAATGGAGATGATTATCAAGCATT
CTTGAAGTGCCTTAACATTTATAACCAAGGGATTATTAAAAAGAACGAGTTACAAAATTTGGTGACTGATTTACTTGGAAAGTATCCTGATCTTATGGAA
GAGTTCATTGATCACCTGGAGTGTCATGGGCATATTGATGGGTTCCTTGCTGGTGTTACGAGCAAAAAATCCCTGGGTAACGATGGACAAGCATCCAGAT
CTTTGAAACTAGAGGACAAGGAAAAGGAGCAGAAGCAAATAGACGGAGCTAAAGAAAAGGAAAGATGCAGGGAGAAGTATATGGCTAAATCCATTCAGGA
GCTTGACCTTTCTAACTGTGAACGTTGTACTCCTAGTTATCGGTTTCTACCAGATGATTATCCAATATCTTCAGCAAGCCAGAGATCAGAGCTTGGTGCT
CAAGTGTTGAATGATCACTGGGTCTCTGTAACATCAGGAAGTGAGGACTACTCTTTTAAACATATGCGTAGAAATCAGTTTGAAGAAAGTCTGTTTAGAT
GTGAAGACGATAGATTTGAGCTGGACATGTTGTTGGAATCTGTGAGCTCCACTACCAAGCGTGCAGAAGAATTGTTTAATGGCATAAACGAAAATAAAGT
AGAAACCTCAATCCATATTGAAGACCACTTTACCGCTCTAAACTTAAGGTGTATAGAGCGGTTATATGGTGACCATGGTCTTGATGTGATGGAAATATTG
CGGAAAAACCGAAGTCTTGCATTGCCTGTTATTTTAACTCGTCTGAAGCAGAAACAAGAGGAGTGGACTAGGTGTCGTACTGATTTTAACAAGGTTTGGG
CTGAAATTTATGTGAAAAACCATTACAAATCACTAGATCATCGCAGCTTCTACTTCAAGCAACAAGAATCAAAGAACTTGAGCACAAAATCATTGGTGGT
TGAGATCAAGGAGCTGAAAGAAAAACAGCAGAGAGAGGATGGTGTTCTTCTTGCTTTCGCCACTGGAAAAAGACAACCCTTGGTTCCGAATTTGAAATAT
AATTACCCAGATAAAAAAATTCATGAAGACTTGTATAAACTTGTCCAATATTCTTGTAAAGAGGTTTGCTCAACCAAAGAACAATTAAATAAAGTTATAA
GACTTTGGACTAACTTCGTGGAACCAATGCTGGGTATTGTTTCTCATCCTGATGGCTCGGAGAGTTGTGAAGGTGAAGGGAAACCAAAGCATCCTCTCAT
GAATTGCACTTCATCAAGCATAGCAGAAAAGGATGGAAGTCCTAATGCTGTTCCTGCAATATCAACTTTTAAGCAAGCAAAATCACCTAGCAATGGAGAT
GAAAACATGTTGCAAGAACTAGGAAACCTTTGCAAGCTCAGTTTGAAAAGTTCAGATAAATTAGCTAAGGAAGATAGCTTATGTGAACTTGATCATGTTG
GTAAGGAAGATGGAGCGCTTAATGTGCTTAGACCAGAAAGAGAGCAGAAAGATAAAGTTGTAACTGACAGAGTATCTGGATTTAACATACAAGGAGTAGC
AGATACCAAAACATCATTTATGATAGGATCAGAATGTGGTCATGAAAGGAACAGTGCTGGGGAGATAGCAGGTTCAGGTTCAAATGTGTCTGTGCCAGGT
GGTGATGCTATTGACCGTCAACTCAATGCTGGTATTGATGCTGGACCTTCTTCAGAGGGTGTGATTGTTGTAAAATCAGTTTTACCTGCAAATGAAGGAG
TAAGGGATGGTGCTAAAAATGATAGATGTCATGAAGAATCCACTGGACCATCCAAAATTGAGAAGGAAGAAGGCGAGTTATCCCCTAATGGTGATTTTGA
AGAGGATAATTTCGATGCCTATGGAGATACTGGCTTGCAGGCTATTGCCACGGGAAAGAATAGTATTGGATGCATGCGGCATGGATCTGGAAATGATGAG
GACTTACACACTCAGGATATCGTAGAAGGACATGATGCTGATGATGAGGATAGTGGGAATGTTTCTGAGGCTCGTGATGCTGCATCAAGCAGCGAGTCTG
CTGGTGATGAGTGCTCCCGGGAAGAGCTTGAGGATGATGAAGTAGAACGTGATGATGCGAAAGCTGAGAGTGAAGGTGAAGCTGAGGGGATGGTTGATAC
ACAGTATAATGGAGGAGATGCACCATTCCCAGAACACTCTCTCATGTCTGTTAAGCCCCTTGCAAAGCATGTTCCAACAGATTTAGTTAACGAGAAAAGG
AGAGATTCCTGGGTTTTCTATGGAAATGATGATTTTTATGTGCTTTTCAGGCTTCATCAGATCCTCTATGACCGTATTTTATCAGCAAGAGTGAATTCAT
CTGGTGCTGAAATCAAATGGAGAACTTCTAAGGATGCTAGTTCTCCAGATCCTTATGCCAGATTTATGAGTGCACTATACAGTTTACTAGATGGATCTGT
TGATAATGCGAAGTTTGAGGATGAGTGCAGAGCTATTATTGGAAATCAGTCATATGTGTTATTCACACTGGACAAGTTAATATATAAATTAGTCAAACAG
CTCCAAACTGTTGCAACTGATGAGATGGCTAGTAAGCTTCTTCAATTGTATGAATACGAAATATCCCGGAAATCTGAGAGCTTCAATGATTTGGTGTATT
ATGACAACATGCGGTTTTTCCTTCATGAGGAGAACATATATCGGTTGGAATTTTCATCTGCTCCATCTGTTTTATCCACCCAGCTGATGGACAACGCAAC
TGAGAAATCTGAAGTGTTGGCAGTTTGCATGGATCCAACTTTTTCTGCATATTTGCATAATGATTATCTCTCTGTCCATCCTATAAAAATGGAGTCGCAT
GACATTACACTGCTGAGAAACAAGCGGAAATATGCTGGCCTAGATGAATTTTCTGCCTTAAGCATGGCCATGGAAGGCGTTAAAATGTTCAATGGTTTGG
AGTGTAAGGTTGCCTGCAATTCATGCAAGATTTCCTATGTCCTAGACACGGAAGATTTCTTTTTTCGAACAAGAAGGAAAAGAAGAAATTCACCTCAAGG
AAGATCTTTGTATCATGATAAGGTTCAAGCAAGAGTACAACGCTTCCGGAGATTCTTATCAGCTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G350200.12 pacid=42791063 polypeptide=Potri.001G350200.12.p locus=Potri.001G350200 ID=Potri.001G350200.12.v4.1 annot-version=v4.1
MKRMRDDIFSAPAPAPAPAPEFKRPLTSTRGESYGQTQIPGGGGGGGSGNSQKLTTTDALQYLKEVKDMFQDQKEKYDMFLEVMKDFKAQRTDTSGVIVR
VKELFKGHNNLIFGFNTFLPKGYEITLDEDEAAPPKKTVEFNQAINFVNKIKKRFQNDERVYKSFLDILNMYRKEHKDINEVYSEVSALFEDHHDLLDEF
ARFLPDTSATPITHTVPYARNSNQHYNERNSTAPSARQTQIDKQRRRDKVSSSHAERDLSVDRPEMEDDKGMVKVHKEQRKRADKENWDRRIHDQDDREP
EHDSSRDFSLQPFLEKRNSSQKVEGFGNSNFACYDDKDNIKSIYNQEFVFFEKVKEKLGNGDDYQAFLKCLNIYNQGIIKKNELQNLVTDLLGKYPDLME
EFIDHLECHGHIDGFLAGVTSKKSLGNDGQASRSLKLEDKEKEQKQIDGAKEKERCREKYMAKSIQELDLSNCERCTPSYRFLPDDYPISSASQRSELGA
QVLNDHWVSVTSGSEDYSFKHMRRNQFEESLFRCEDDRFELDMLLESVSSTTKRAEELFNGINENKVETSIHIEDHFTALNLRCIERLYGDHGLDVMEIL
RKNRSLALPVILTRLKQKQEEWTRCRTDFNKVWAEIYVKNHYKSLDHRSFYFKQQESKNLSTKSLVVEIKELKEKQQREDGVLLAFATGKRQPLVPNLKY
NYPDKKIHEDLYKLVQYSCKEVCSTKEQLNKVIRLWTNFVEPMLGIVSHPDGSESCEGEGKPKHPLMNCTSSSIAEKDGSPNAVPAISTFKQAKSPSNGD
ENMLQELGNLCKLSLKSSDKLAKEDSLCELDHVGKEDGALNVLRPEREQKDKVVTDRVSGFNIQGVADTKTSFMIGSECGHERNSAGEIAGSGSNVSVPG
GDAIDRQLNAGIDAGPSSEGVIVVKSVLPANEGVRDGAKNDRCHEESTGPSKIEKEEGELSPNGDFEEDNFDAYGDTGLQAIATGKNSIGCMRHGSGNDE
DLHTQDIVEGHDADDEDSGNVSEARDAASSSESAGDECSREELEDDEVERDDAKAESEGEAEGMVDTQYNGGDAPFPEHSLMSVKPLAKHVPTDLVNEKR
RDSWVFYGNDDFYVLFRLHQILYDRILSARVNSSGAEIKWRTSKDASSPDPYARFMSALYSLLDGSVDNAKFEDECRAIIGNQSYVLFTLDKLIYKLVKQ
LQTVATDEMASKLLQLYEYEISRKSESFNDLVYYDNMRFFLHEENIYRLEFSSAPSVLSTQLMDNATEKSEVLAVCMDPTFSAYLHNDYLSVHPIKMESH
DITLLRNKRKYAGLDEFSALSMAMEGVKMFNGLECKVACNSCKISYVLDTEDFFFRTRRKRRNSPQGRSLYHDKVQARVQRFRRFLSA
|
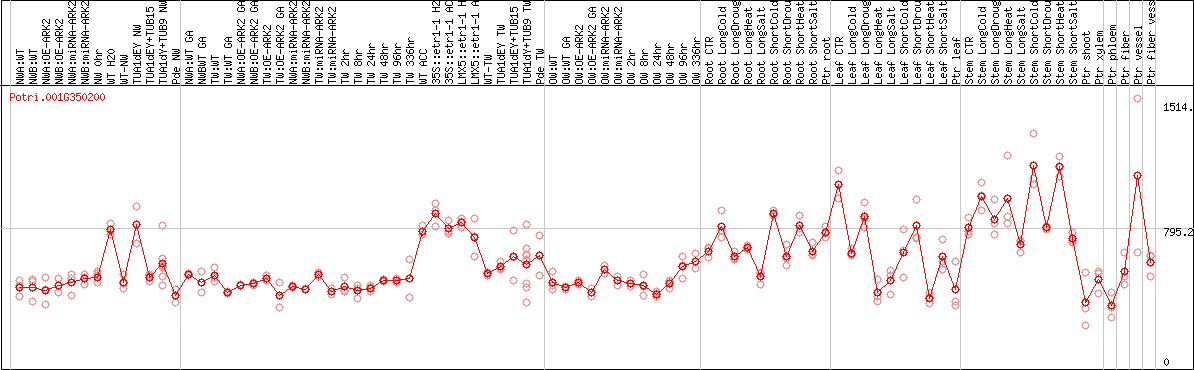
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G350200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
