Potri.001G354900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G354900.1 pacid=42791903 polypeptide=Potri.001G354900.1.p locus=Potri.001G354900 ID=Potri.001G354900.1.v4.1 annot-version=v4.1
ATGAGGGAAGATACAGAAGGTGCATCAACAAACAGTATTGCTAATGGCCAGAAAACCACGAATGGAGAAGATCAGAAGGTTGCATTCCACAAGCTGTTCA
CGTTTGCAGATAGGCTTGACGTGGTTTTGATGATCGTTGGCACGTTAAGTGCCATTGCTAATGGCCTTGCTCAGCCTCTGATGACACTCATTTTTGGCCA
GCTTATAAACTCCTTCGGCTCCAGTGATCGTTCAAACGTTGTTAAGGAAGTTTCCAAGGTAGCACTAAATTTTGTGTACTTGGCAATTGGATCGGGCATC
GCTTCTCTTTTACAGGTGTCCAGTTGGATGGTTACTGGAGAAAGACAGTCTACCCGCATTCGAAGTCTGTACTTGAAAACAATACTGAGACAGGACATTG
GCTTCTTCGACTCTGAAACATCCACTGGTGAAGTCATTGGAAGAATGTCTGGTGACACTATTCTCATTCAAGATGCCATGGGTGAAAAGGTCGGGAAGTT
CATACAACTACTAGCAACATTTTTCGGTGGCTTTGCGATTGGCTTCATAAAAGGCTGGCTTCTTGCATTAGTTCTGCTTTCCAGTATTCCTCCTCTTGTA
ATTGCTGGTGGAGTCATGGCTTTGATCATGACAAAAATGTCAAGCCGTGGACAGGTTGCTTATGCAGAGGCTGGAAACATTGTAGAGCAAACAGTTGGAG
CCATACGAACAGTTGCATCTTTCACGGGGGAGAAGCATGCAATTGAAAAATATAACAGTAAGCTGAAAATTGCGTATAACTCTGCCGCCCAACAAGGGTT
GGCCTCAGGCTTGGGTCTCGGTACAATGTTGTTCATTGTATTTGGCACTTATGCACTTGCCATATGGTATGGATCCAAATTGATCGTCGAGAAAGGTTAT
AACGGTGGACAAGTTATGACTGTAATTATATCAATTATGACTGGAGGAATGTCATTGGGTCAGACTTCTCCATGTTTGAATGCTTTTGCATCAGGGCAAG
CTGCGGCATATAAGATGTTTGAGACCATTGAACGGAAGCCAAAAATCGATCCCTATGACACTAGTGGGATGGTAGTGGAAGATCTGGATGGAGAAATTGA
ACTCAGAGATGTATATTTCAGATACCCAGCTAGGCCTGAGGTTCAAATCTTCTCTGGATTTTCACTGCAAGTTCCAAGTGGCACAACTACAGCTTTGGTT
GGGCAAAGTGGAAGTGGAAAGTCAACTGTTATTAGCCTTGTTGAGAGATTTTATGATCCTGATTCAGGTGAAGTACTTATCGATGGTGTTGACTTGAAGA
AGCTGAAGCTTAGCTGGATAAGGGAGAAGATTGGACTAGTTAGCCAGGAACCTATTTTATTTGCAACCAGCATAAAGGAAAATATAGCTTATGGGAAAGA
AAATGCAACTGATCAGGAGATTAGAACAGCAATCCAACTAGCCAATGCTGCAAAATTCATTGACAAAATGCCTGAGGGACTTGACACCATGGTTGGTGAG
CATGGTACTCAGCTGTCTGGTGGCCAAAAGCAAAGAATTGCAATTGCAAGGGCCATATTGAAGAATCCAAAGATTCTTCTTCTTGATGAAGCAACAAGTG
CACTTGATGCGGAGTCTGAACGCATTGTCCAAGATGCGTTGGTGAAAATTATGTGCAACCGAACAACTTTGGTTGTTGCACATCGTTTGACAACTATCAG
GAATGCGGACATGATTGCTGTGGTTCATTTGGGAAAAATTGTAGAGAAAGGAAGTCATGAGGAGCTGACAAAAGATCCGGAGGGGGCTTACTCCCAGTTA
ATTCGCCTCCAAGGAGGAGCTATGGATTCTGAAGAATCTCAGGACATTGATGCAGATATGTCGAGATCTAGTTTGGACAGAGACAGGTCCATTTCCAGTC
CTAGGAGCCAGAAACATTCTGTTCAGGGATCCATAAGTAGAGGTTCATCAGGAAGTCGGCGCTCCTTCACACTCAACACTGTTGGCTTTGGCATGCCCGG
TCCTACCAGTGTTCATGATGATGAATTCGAACAGAACAATGAAAGAAATGTAAAGCCTAAAGAAGTTTCTATTAAACGACTTGCCTACCTCAACAAGCCC
GAGCTTCCTGTTTTATTTCTCGGCACCGTTGCTGCAGTTATACATGGTGTGATTTTCCCAGTGTTTGGCCTTCTACTATCAAAAGCCATTAACATGTTTT
ACGAACCTCCAAAGGAGATTCGAAAAGATTCCAAGTTCTGGGCAGTGCTGTATTTGGGATTGGGTTTTATTACCTTTGCAGCCCTTCCACTGCAATATTA
CCTCTTTGGAATTGCTGGTGGTAAGTTGATTGAACGAATTCGTTCCAAAACATTTGAAAAAGTAGTGCACCAAGAGATTAGCTGGTTTGATGACCCTACA
AATTCAAGTGGTGCAATTGGTGCAAGGTTGTCAACGGATGCTTCTACAGTTAGGAGGCTTGTTGGTGATTCCCTGTCCTTAATTGTCCAAAATATATCAA
CAATCTTGTCAGCTTTGGTTATAGCATTTTCAGCCAACTGGATGTTGACCTTGATAATTATTGCTATATCTCCCTTGCTATTTATCCAAGGGTACATGCA
AGCAAAGTTTATGAAGGGGTTCAGTGCAGATTCAAAGATGATGTATGAACAAGCTAGTCAAGTGGCTAATGATGCGGTTGGCAGCATTAGAACTGTTGCT
TCTTTTTGTGCCGAGAAGAAGGTAATGGAGTTGTATCAAAAGAAATGTGAAGGTCCCACGAAGCAAGGTGTTCGGCTTGGGTTTGTTAGTGGCATTGGTT
ATGGCTTATCATTCTTTATACTCTATTGCACCAATGCATTTTGTTTCTATATTGGAGCTATCTTCGTCCAAAATGGGAAAACAACATTTGCAGATGTTTT
CAGGGTTTTCTTTGCGTTGACTATTGGAGCACTGGGGGTTTCCCAATCTAGTGGCCTGGCTCCAGACACTGCCAAGGCCAAAGATTCAGCAGCTTCAATA
TTTGCAATTCTTGATAGAAAGCCCAAAATTGACTCAAGTAGAGATGAAGGGCTCACTCTACCACATGTTAATGGTGATATTGAGATTGAACATGTTAGCT
TTAAATACCCGATGAGACCACATGTTCAAATCTTCAGAGACATGTCCCTTAGTATTCCCTCTGGAAAGACTGTTGCTCTAGTTGGGGAGAGTGGTAGCGG
GAAGTCAACAGTTATCAGCCTCATAGAGAGGTTTTATGATCCTGATTCCGGTCACGTATATTTGGACAGTGTGGAAATCAAGAAGTTCAAACTAAACTGG
TTGAGACAACAAATGGGTTTGGTCAGCCAAGAACCAATACTGTTCAATGAAACTATCCGCGCAAATATAGCTTATGGAAAGCATGGAGAGATTGCCGAGG
AAGAGATAATCGAAGCTACAAGAGCATCAAATGCGCACAACTTCATTAGCACACTACCACAAGGCTATGACACCAAAGTAGGTGAACGCGGGATTCAACT
GTCGGGCGGACAGAAGCAGAGAATAGCCATCGCCAGGGCAATCCTAAAGAATCCGAAAATTCTCTTACTCGACGAGGCAACGAGTGCGCTGGATGCAGAG
TCAGAGCGTATAGTACAAGAAGCATTAGATAGAGTTATGGTGAACAGAACAACCGTTGTTGTTGCTCATCGCCTTGCCACAATTAAAGGTGCTGATGTCA
TTGCTGTAGTGAAGAATGGTGCGATTGCCGAGAAGGGAAAGCATGATGTGCTCATGAAGATCACTGACGGGGCTTATGCATCCTTGGTCGCCCTTCATAT
GAGTGCAACTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G354900.1 pacid=42791903 polypeptide=Potri.001G354900.1.p locus=Potri.001G354900 ID=Potri.001G354900.1.v4.1 annot-version=v4.1
MREDTEGASTNSIANGQKTTNGEDQKVAFHKLFTFADRLDVVLMIVGTLSAIANGLAQPLMTLIFGQLINSFGSSDRSNVVKEVSKVALNFVYLAIGSGI
ASLLQVSSWMVTGERQSTRIRSLYLKTILRQDIGFFDSETSTGEVIGRMSGDTILIQDAMGEKVGKFIQLLATFFGGFAIGFIKGWLLALVLLSSIPPLV
IAGGVMALIMTKMSSRGQVAYAEAGNIVEQTVGAIRTVASFTGEKHAIEKYNSKLKIAYNSAAQQGLASGLGLGTMLFIVFGTYALAIWYGSKLIVEKGY
NGGQVMTVIISIMTGGMSLGQTSPCLNAFASGQAAAYKMFETIERKPKIDPYDTSGMVVEDLDGEIELRDVYFRYPARPEVQIFSGFSLQVPSGTTTALV
GQSGSGKSTVISLVERFYDPDSGEVLIDGVDLKKLKLSWIREKIGLVSQEPILFATSIKENIAYGKENATDQEIRTAIQLANAAKFIDKMPEGLDTMVGE
HGTQLSGGQKQRIAIARAILKNPKILLLDEATSALDAESERIVQDALVKIMCNRTTLVVAHRLTTIRNADMIAVVHLGKIVEKGSHEELTKDPEGAYSQL
IRLQGGAMDSEESQDIDADMSRSSLDRDRSISSPRSQKHSVQGSISRGSSGSRRSFTLNTVGFGMPGPTSVHDDEFEQNNERNVKPKEVSIKRLAYLNKP
ELPVLFLGTVAAVIHGVIFPVFGLLLSKAINMFYEPPKEIRKDSKFWAVLYLGLGFITFAALPLQYYLFGIAGGKLIERIRSKTFEKVVHQEISWFDDPT
NSSGAIGARLSTDASTVRRLVGDSLSLIVQNISTILSALVIAFSANWMLTLIIIAISPLLFIQGYMQAKFMKGFSADSKMMYEQASQVANDAVGSIRTVA
SFCAEKKVMELYQKKCEGPTKQGVRLGFVSGIGYGLSFFILYCTNAFCFYIGAIFVQNGKTTFADVFRVFFALTIGALGVSQSSGLAPDTAKAKDSAASI
FAILDRKPKIDSSRDEGLTLPHVNGDIEIEHVSFKYPMRPHVQIFRDMSLSIPSGKTVALVGESGSGKSTVISLIERFYDPDSGHVYLDSVEIKKFKLNW
LRQQMGLVSQEPILFNETIRANIAYGKHGEIAEEEIIEATRASNAHNFISTLPQGYDTKVGERGIQLSGGQKQRIAIARAILKNPKILLLDEATSALDAE
SERIVQEALDRVMVNRTTVVVAHRLATIKGADVIAVVKNGAIAEKGKHDVLMKITDGAYASLVALHMSAT
|
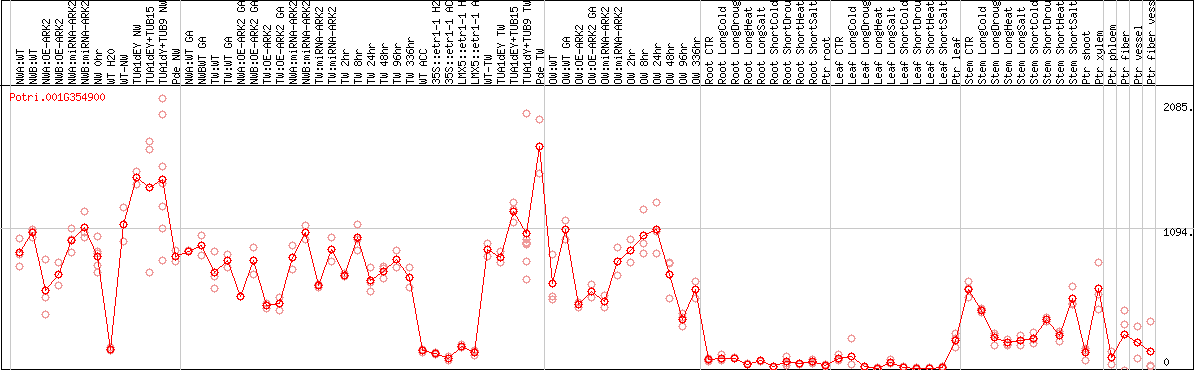
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G354900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
