Potri.001G362200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G362200.2 pacid=42789695 polypeptide=Potri.001G362200.2.p locus=Potri.001G362200 ID=Potri.001G362200.2.v4.1 annot-version=v4.1
ATGCACCTCTTTGATTCAACAAAGCTTGGTCTCTTTTCCAGTCTTTCAAACTCATCCTCTTTAATGTATTCAGGTGCTAGTTTTCTCCTTAATCCCATTT
TCCTGCGTGGGTTTTCTGGTTCATTGCACCTAGTTTTGTTACTTGCCTTAGTCGTCTCATATGCGTGGAAGAAACTCATGAGGGGTGATGGTGGTGAGGG
TTCTGGAGAGAGGTTTAGGAACAACAAGAGGTTTCTGTGTTCTAAGCAAACTCTTTTATGTTGTTTAGGTGTTTCAGTGTTTAATCTTTTCTTGTGTTTG
CTGAGCTACTTTTATTGGTATAAAAATGGATGGTCTGATGATAAACTAGTGAACCTTTTTGATTCAGTGCTTAGAACATTCTCTTGGGGTGCACTTTCTG
TCTATTTGTATACCCTTAATTCAGGTGAGAAAAGGTTCTCATTTTTATTGAGAGTTTGGTGGGTTTTTTATTTCTCAATTTCTTGTTATTGTCTTGTTGT
AGATTTTCTTTTTTATCACAAACATGAGTCCTCAGAGGTACGGTATTTAGTATCTGATATAGTCTCTGTGTTCACTGCTTTGTTTCTTTGTTATGTTGGG
TTCTTGAGACATGAGACTAAGGATAGCCTTCTCGAGCAACCCTTTATGAATGTTGATATTTCTAGCGACACAAGTAATACTCCTACTTTGGAATCAAATA
AGTCAAGGGGGGGTGATGCTGTGACGACTTATGCAAATGCTGGTCCTTTCAGTATTCTTACCTTCTACTGGATGAATTCTCTAATTGCTTTTGGCAATAA
GAAGACTTTGGACCTTGAGGATGTTCCTCAACTAGACAGTCTTGATAGTGTGTTTAGAGGATTCCCAGCTTTTAGAAATAAGCTCGAGTCACATAATGGT
GCTTCTAGTAGAGTTACCCCTTTCAAGCTCGCCAAGACATTGTTCTTGTCAGCTTGGAAGGAAATTCTATGGACAGCTTTGTTGGCAATAATATACACCT
CAGCTTCCTATGTTGGTCCATACCTTATCGACGCTTTTGTTCAATGCCTTGATGGACGAGGAGAATACAAAAATCAGGGATATATTTTAGCTTCAACCTT
TTTCGTTGCAAAGCTTGTAGAGTGCATCTCACAGAGGCACTGGTTCTTTAGGTTGCAACAAATAGGATTTAGGCACCGTGCAGTAACAACCACAATGATT
TACAACAAAGGTTTGACCCTCTCCAATCAGTCAAAGCAGGGCCAAACTAGTGGTGAAATCATCAATATCATGACTGTTGATGCAGACAGAATTGGTGAGT
TTAGTTGGTACATGCACGATCCATGGTTGATCCTATTGCAAGTTGGTTTGGCCTTATTGATACTGTACCAAAATTTGGGGCTTGCATCAGTTGCTGCTTT
TGTTGCAACAATAGTTGTCATGTTGATAAACTATCCTCTGGGTAGATTACAAGAAACGTTTCAAGACAAGTTAATGGACTCAAAAGATAAAAGAATGAGG
GCAACCACAGAGATTTTAAGGAATATGAGAATTCTTAAGCTTCAGGGGTGGGAAATGAAGTTTCTGTCTAAGATTCTTGAACTCAGAGACGTAGAGACAG
GATGGTTGAAGAAATATGTTTATAATTCGGCAATGGTCATGTCTGTCTTCTGGCTAGCTCCTACTCTTGTGGCTGTGGCCACTTTTGGTGCTTGTATGCT
TATAGGCATCCCACTTGAATCAGGGAAAATCTTATCTGCACTTGCAACTTTCAGGGTCCTTCAAATGCCAATCTATAGTCTTCCTGATACACTTTCTATG
ACAGTTCAGACCAAGGTTTCTCTTGACCGAATTGCCTCTTTCCTTTGTCTGGATGACTTGCAGACTGATGTTTTGGAGATGCAACCAATTGGAACCTCTG
ATACAGCAATTGAGATTGCTGATGGAAATTTCTCCTGGGATTTGTCTTCACCTAGTGCAACATTAAAGGATGTTAACGTCAGAGTGCTCCATGGCATGAG
AGTTGCCGTTTGCGGAACAGTTGGTTCAGGCAAGTCAAGCTTACTTTCCTGCGTTTTGGGGGAACTGCACAAGATCTCAGGGACTCTCAAGTTATGTGGG
ACAAAGGCCTATGTTGCCCAGTCTGCCTGGATACAGAGTGGCAAGATTGAAGAGAACATATTGTTTGGTAAGGAGATGGACAGAGAAAGATATGAAAGGA
TACTGGAGGCATGTTCCCTGAAGAAGGACCTTGAAATTATGTCATTTGGTGATCAAACAGTTATTGGTGAGAGGGGAATCAACTTAAGCGGTGGTCAGAA
GCAGAGAATTCAGATAGCACGGGCTCTGTACCAAGACGCTGATATCTATCTATTTGATGACCCTTTCAGTGCTGTGGATGCTCATACAGGATCTCATTTA
TTTAAGGAAGTTTTACTTGGTCTTTTGAGTTCTAAAACAGTTATTTATGTTACTCATCAAGTTGAATTCTTACCTGCGGCTGATCTTATCTTGGTAATGA
AAGATGGAAAGATTGCACAAGCGGGTAAATATGATGACATTCTCTGTTCAGGATCTGACTTTATGGAACTTGTAGGTGCCCACGAGGCAGCTTTATCAGT
GCTTGATTCCAAGCAGAAAGGACCGGCATCAGATACTGATGGGATTGTGCAGAAACAAGGAAAAAACGATTATGAAAATGGTAAATCAGATAAGGTAGCT
GAGCCAAAAGCACAGATCATTCAAGAAGAAGAAAGAGAGACGGGTAGTGTTGGATTTCCTATCTACTGGAACTATATCACAACAGCATATGGAGGAGGTC
TTGTGCCCTTTATATTATTGGCACAGATTCTCTTTCAGGTCCTTCAAATTAGTAGCAACTACTGGATGGCATGGGCAGCTCCTGTGTCAAAGGATGTTAA
ACCTGTTGTCAGTGGATCTACATTAATAATCGTCTATGTCTTGTTCGCTATTGGAAGTTCGATTTGCATTTTAGCTAGAGCCACGCTTCATGCAACAGCA
GGGTACAAAACTGCAACTCTACTCTTCAATAAAATGCATCTGTGCATATTTCGTGCTCCCATGTCCTTTTTCGATGCTACCCCTAGTGGACGAATTATAA
ATAGAGTTTCTACGGATCAAAGTGCAGTTGAAAATCGAATTCCATATCTAATTGGGGCACTTGCCTTCTCAGTTATCCAGCTTCTAGGAATCATTGCAGT
GATGTCACAGGTAGCATGGCAAGTTTTTATCGTTTTCATCCCTGTGATTGCTGCCTGCATCTGGTATCAGAGATATTACATATCTTCTGCAAGAGAGCTT
TCACGTTTGGTTGGAGTATGCAAAGCTCCAGTTATACAACATTTTGCTGAAACAATTTCGGGAGCAACAACTATCAGGAGCTTCGATCAGCAATCAAGAT
TCCAAGAAACAAATATGAAAGTTACAGATGCGTATTCTCGGCCCAAATTTCACATTGCTGGTGCAAGGGAGTGGCTTTGCTTCCGCCTGGATATGTTCTC
TTCTGTTACTTTTGCTTTCTCTTTGGCTTTCTTAGTCTCCTTCCGAAGCAATATTGATCCAGCCATTGCAGGCTTGGCTGTGACATATGGACTCAACCTA
AACATGTTACAAGCTTCAGTGATATGGAATTTATGTGATTGTGAGAACAAAATCATATCTGTAGAGAGAATTATTCAATACATGTCTATTCCTAGTGAGC
CGCCTCTTTTAGTAGAAGCAAACAGGCCAGGCCGTTCTTGGCCATCCCATGGTGAAGTTGATATCAACAATCTGCAGGTCCGTTATGCTCCACAACTGCC
ACTTGTGCTACGCGGTATTACATGCACCTTTCCAGGGGGAAAGAAAACTGGTATTGTAGGGAGAACAGGCAGTGGAAAATCAACTCTCATACAGACTCTT
TTTCGTATTGTCAAACCTGCTGCTGGTCAGATTATGATTGATGGACTTGATATCTCATCGATTGGACTGCATGATTTAAGGTCAAGACTAAGCATTATTC
CTCAGGATCCAACCATGTTTGAAGGGACTATACGAAGTAATCTGGACCCACTTGAAGAGCACACTGATGAACTGATCTGGGAGGCTCTTGACAAGTGCCA
ACTTGGAGACGAAGTTAGGAAGAAAGAAAGAAAGCTAGACTCCACAGTCAACGAGAACGGAGAGAACTGGAGTATGGGTCAGAGGCAGCTAGTTTGTCTT
GGGCGTGTGCTACTCAAGAAAAGTAAGGTGTTAGTCCTTGATGAAGCTACTGCTTCGGTTGACACATCCACCGATAATCTTATCCAGCAAACCCTCAGGC
AACACTTCTCTGACTGTACAGTTATAACCATTGCACATCGGATAACTTCTGTTCTTGACAGTGACATGGTTTTGCTTCTGAGTCACGGGCTCGTTATGGA
ATATGATTCTCCAACAAGATTGTTAGAAAACAAGTCATCATCTTTCGCACAACTTGTTGCAGAGTACACAATCAGGTCAAATGCCAATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G362200.2 pacid=42789695 polypeptide=Potri.001G362200.2.p locus=Potri.001G362200 ID=Potri.001G362200.2.v4.1 annot-version=v4.1
MHLFDSTKLGLFSSLSNSSSLMYSGASFLLNPIFLRGFSGSLHLVLLLALVVSYAWKKLMRGDGGEGSGERFRNNKRFLCSKQTLLCCLGVSVFNLFLCL
LSYFYWYKNGWSDDKLVNLFDSVLRTFSWGALSVYLYTLNSGEKRFSFLLRVWWVFYFSISCYCLVVDFLFYHKHESSEVRYLVSDIVSVFTALFLCYVG
FLRHETKDSLLEQPFMNVDISSDTSNTPTLESNKSRGGDAVTTYANAGPFSILTFYWMNSLIAFGNKKTLDLEDVPQLDSLDSVFRGFPAFRNKLESHNG
ASSRVTPFKLAKTLFLSAWKEILWTALLAIIYTSASYVGPYLIDAFVQCLDGRGEYKNQGYILASTFFVAKLVECISQRHWFFRLQQIGFRHRAVTTTMI
YNKGLTLSNQSKQGQTSGEIINIMTVDADRIGEFSWYMHDPWLILLQVGLALLILYQNLGLASVAAFVATIVVMLINYPLGRLQETFQDKLMDSKDKRMR
ATTEILRNMRILKLQGWEMKFLSKILELRDVETGWLKKYVYNSAMVMSVFWLAPTLVAVATFGACMLIGIPLESGKILSALATFRVLQMPIYSLPDTLSM
TVQTKVSLDRIASFLCLDDLQTDVLEMQPIGTSDTAIEIADGNFSWDLSSPSATLKDVNVRVLHGMRVAVCGTVGSGKSSLLSCVLGELHKISGTLKLCG
TKAYVAQSAWIQSGKIEENILFGKEMDRERYERILEACSLKKDLEIMSFGDQTVIGERGINLSGGQKQRIQIARALYQDADIYLFDDPFSAVDAHTGSHL
FKEVLLGLLSSKTVIYVTHQVEFLPAADLILVMKDGKIAQAGKYDDILCSGSDFMELVGAHEAALSVLDSKQKGPASDTDGIVQKQGKNDYENGKSDKVA
EPKAQIIQEEERETGSVGFPIYWNYITTAYGGGLVPFILLAQILFQVLQISSNYWMAWAAPVSKDVKPVVSGSTLIIVYVLFAIGSSICILARATLHATA
GYKTATLLFNKMHLCIFRAPMSFFDATPSGRIINRVSTDQSAVENRIPYLIGALAFSVIQLLGIIAVMSQVAWQVFIVFIPVIAACIWYQRYYISSAREL
SRLVGVCKAPVIQHFAETISGATTIRSFDQQSRFQETNMKVTDAYSRPKFHIAGAREWLCFRLDMFSSVTFAFSLAFLVSFRSNIDPAIAGLAVTYGLNL
NMLQASVIWNLCDCENKIISVERIIQYMSIPSEPPLLVEANRPGRSWPSHGEVDINNLQVRYAPQLPLVLRGITCTFPGGKKTGIVGRTGSGKSTLIQTL
FRIVKPAAGQIMIDGLDISSIGLHDLRSRLSIIPQDPTMFEGTIRSNLDPLEEHTDELIWEALDKCQLGDEVRKKERKLDSTVNENGENWSMGQRQLVCL
GRVLLKKSKVLVLDEATASVDTSTDNLIQQTLRQHFSDCTVITIAHRITSVLDSDMVLLLSHGLVMEYDSPTRLLENKSSSFAQLVAEYTIRSNAN
|
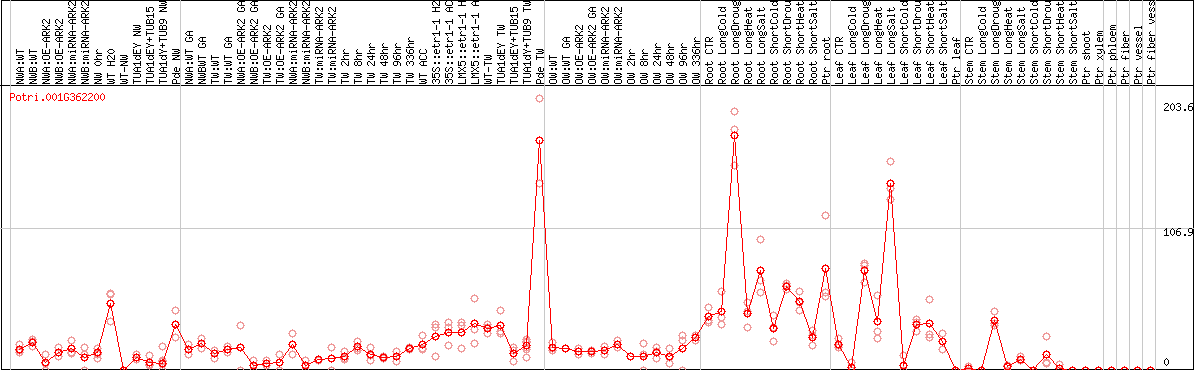
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G362200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
