External link
Symbol
Pt-ZAC.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G21160 506 / 0
ZAC, AGD12
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
AT4G05330 496 / 3e-178
AGD13
ARF-GAP domain 13 (.1)
AT3G07940 349 / 2e-119
Calcium-dependent ARF-type GTPase activating protein family (.1)
AT1G70790 136 / 5e-39
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G73580 134 / 2e-38
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT5G37740 133 / 5e-38
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G70800 132 / 2e-37
EHB1
ENHANCED BENDING 1, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G17980 132 / 2e-37
AtC2
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G66360 131 / 5e-37
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT2G01540 130 / 6e-37
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G098500
573 / 0
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.003G198301
345 / 1e-116
AT3G07940 503 / 7e-178
Calcium-dependent ARF-type GTPase activating protein family (.1)
Potri.T125706
345 / 2e-116
AT3G07940 503 / 8e-178
Calcium-dependent ARF-type GTPase activating protein family (.1)
Potri.001G026400
162 / 1e-47
AT3G07940 211 / 4e-66
Calcium-dependent ARF-type GTPase activating protein family (.1)
Potri.008G131500
150 / 1e-44
AT3G17980 202 / 5e-67
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.012G047100
143 / 8e-42
AT3G17980 251 / 2e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.010G110900
141 / 5e-41
AT1G70790 233 / 3e-79
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.004G089500
133 / 7e-38
AT3G17980 245 / 4e-84
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.017G125500
132 / 2e-37
AT3G17980 249 / 6e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038539
465 / 4e-166
AT4G21160 491 / 4e-176
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Lus10023265
446 / 5e-158
AT4G21160 495 / 3e-177
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Lus10016347
355 / 2e-117
AT3G07940 478 / 8e-164
Calcium-dependent ARF-type GTPase activating protein family (.1)
Lus10024165
350 / 3e-117
AT3G07940 491 / 1e-171
Calcium-dependent ARF-type GTPase activating protein family (.1)
Lus10039538
350 / 6e-117
AT3G07940 490 / 4e-171
Calcium-dependent ARF-type GTPase activating protein family (.1)
Lus10002760
345 / 5e-113
AT3G07940 480 / 2e-164
Calcium-dependent ARF-type GTPase activating protein family (.1)
Lus10009684
143 / 8e-41
AT3G17980 254 / 1e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10009053
142 / 1e-40
AT3G17980 255 / 1e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10038745
133 / 5e-38
AT5G47710 241 / 1e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10033585
133 / 5e-38
AT3G17980 244 / 1e-83
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
CL0154
PF01412
ArfGap
Putative GTPase activating protein for Arf
Representative CDS sequence
>Potri.001G372000.1 pacid=42789929 polypeptide=Potri.001G372000.1.p locus=Potri.001G372000 ID=Potri.001G372000.1.v4.1 annot-version=v4.1
ATGAGCCGTTTGTCAGAGCTTCAGCAGGTTGCCTCAGGTAAAAGAAGATTAAAAGATCTATTGCTCCAAAGTGATAATCGCTTTTGCGCTGATTGTGGTG
CTCCAGATCCAAAATGGGCATCAGCAAATATTGGGGTTTTCATATGCTTAAAATGCTGTGGTGTGCACAGAAGTCTTGGTACACATATCTCTAAGGTTTT
ATCCGTGACGTTGGATGAATGGTCTGATGATGAAATTGATGCCATGATTGAAGTTGGAGGAAACTTGTCTGCTAATTCAATTTATGAGGCCTTTTTACCT
GAAGGAGTTTCAAAGCCAGGGCCAAATTCCAGTAATGAGGAGCGTACGAGGTTCATCAGGTCAAAGTATGAGCTTCAAGAATTTTTGAAACCTAGTTTGC
GGATCACATCGGGGAAGACTTCCTCCTCTTCTCTCAAGTCAAGTTTATCCACAAACCTTTTTGATAGTTTTCAAATTCCGAGTGTCTCACAGAATTTGGA
TGGCATAGTAGAGTTTATGGGAATATTGAAGGTTAAAGTTATAAAAGGTACAAATTTAGCTATTAGAGATATGATGTCAAGTGATCCCTATGTGATCGTT
GCTCTTGGGAAACAGACTGCTCAAACCACCGTGATGAAGAGCAACTTGAATCCAGTCTGGAATGAAGAACTCATGCTATCTGTTCCACAGGATTTTGGGC
CCATAAAGTTGAGCGTTTTTGATCATGACACATTTTCTGCCGATGACATAATGGGGGAAGCAGAGATTGACATCCAGCCATTGATCACATCAGCAATGGC
ATTCGGGGATCCAGAAATGTTTGGAAATATGCAGATTGGAAAATGGCTGAAATCAAACGACAACGCCCTCATAGATGATAGCATCATAAACATTGTTGAC
GGGAAGGTGAAACAGGAGATCTCATTGAAACTCCAGAATGTTGAATCTGGGGAACTACAAGTAGAACTGGAGTGGATGCCTCTTGACCAGTAA
AA sequence
>Potri.001G372000.1 pacid=42789929 polypeptide=Potri.001G372000.1.p locus=Potri.001G372000 ID=Potri.001G372000.1.v4.1 annot-version=v4.1
MSRLSELQQVASGKRRLKDLLLQSDNRFCADCGAPDPKWASANIGVFICLKCCGVHRSLGTHISKVLSVTLDEWSDDEIDAMIEVGGNLSANSIYEAFLP
EGVSKPGPNSSNEERTRFIRSKYELQEFLKPSLRITSGKTSSSSLKSSLSTNLFDSFQIPSVSQNLDGIVEFMGILKVKVIKGTNLAIRDMMSSDPYVIV
ALGKQTAQTTVMKSNLNPVWNEELMLSVPQDFGPIKLSVFDHDTFSADDIMGEAEIDIQPLITSAMAFGDPEMFGNMQIGKWLKSNDNALIDDSIINIVD
GKVKQEISLKLQNVESGELQVELEWMPLDQ
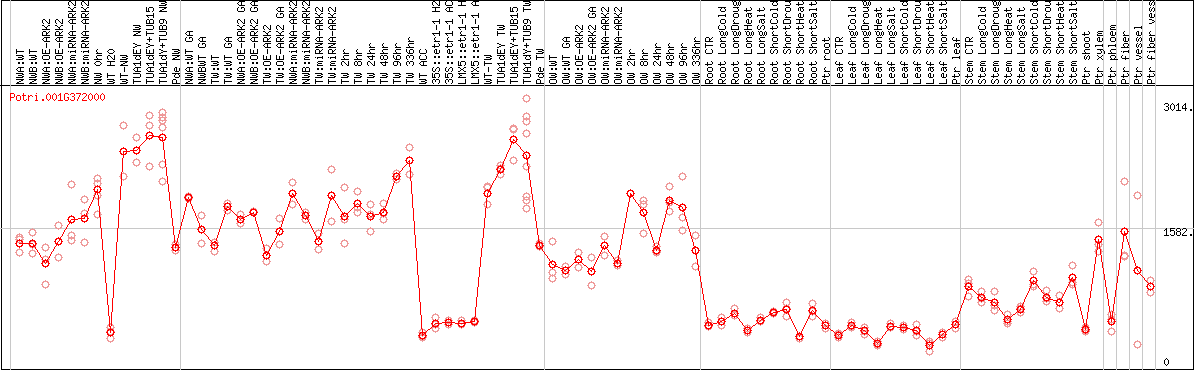
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G372000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.