PSCLRR52.53 (Potri.001G389100) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PSCLRR52.53 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G389100.1 pacid=42789453 polypeptide=Potri.001G389100.1.p locus=Potri.001G389100 ID=Potri.001G389100.1.v4.1 annot-version=v4.1
ATGAGTTGCTCAGGACGGGCTGGTCAAGTCATATTTTGCCTCCTGATATTGATGTTGTATTGTCAACTAACTGTGTCACAATTATGCCCCAAAGATGAAA
GCTATGCTTTACTCCAACTTAAGGATGGATTTGAGTATGATGAATCCGCTTCGTTGGATTGCAGCCCTCCTTATCCTAGGATGGCCTCATGGAAATCTGG
AACAGATTGCTGTTCGTGGGATGGAGTCGCATGCCATGGGGTCACAGGTCATGTGATCGCTCTTGACCTTAGTTGCAGTGGTCTTAGAGGCAACCTCAGT
TCCAACAGCAGTCTCTTTCACCTTTCACACCTTCGGAGACTCAACCTTGCTTTCAATTATTTTAACCGCTCCTCCATTCCACCTGAGTTTGGTATGTTTT
CAAGTTTGACACACCTTAACCTTTCTAGCACATGGTTCTCAGGTCAAGTCCCAACTGAAATATCTCACTTGTCCAAATTGATCTCTCTTGATCTTTCTTT
GAATGAACCATTGATACTCGAAGCACCAGCCATGAAAATGATTGTCCAAAACCTTACTCTAGTAAGAGAGATTTTTCTTGATTACATAAACATGTCTTCA
GTTGATCTTGGTTCCTTGATGAACCTATCTTCTTCATTAACTTCTCTCAGCCTCAATTTATGTGGATTGCAAGGACAATTCCCGGAAAACATTTTCCACC
TACCAAACCTCCAGCTCCTGAGTTTATTGTTGAATAGCGATCTATACGGGCGTTTACCTGTGTCCAATTGGAGCAGTTCCCTTGAACTGTTGAAGCTTGG
CTCCACAAGTTTCTCAGGAGGATTGCCAGAAATAATTGGCAATCTTGACTCCATCAAGGTGTTGGATCTTGGAAACTGTGCTTTCTACGGATCAGTTCCA
GCTTCGCTTGGGAATCTTCAACAGCTCAATCAATTAGATCTTTCGAATAACAACTGGACAGGGCAGATCCCGGATGTTTTCGGCAACCTCAGCAAACTAA
ACAGCCTATCCCTTCAAGTCGGTAATTTCAGTGGTATGTTGCCATCATCTGTCTTTAACCTCACTGAACTCTTACGTTTAGACCTGTCTCAAAACCAACT
AGAGGGAACTCTTCCTGATCACATATGTGGCCTTGACAATGTTACTTATCTTGACCTATCTTATAACTTGCTCAGTGGCACAATACCATCTTGTTTGTTC
GGTCTCCCATCATTGGTCTGGTTTAACCTTAATAATAACCACCTCACTGGAGAACTTGGTGAACATTGGTCAAAGTCTTTGCTAGAAATAAGACTGGAAA
GTAACAAGATAAATGGTCTTATTCCACCATCAATCTCTGAATTGGTGAATCTCACCAATTTTGACGTTTCATCAAACAACTTGAGTGGCATTGTGGACTT
AAACCTGTTTTCAAACATGAAAAACCTTTGGGGACTTGATCTTTCACACAACAGTCTGTCTGTCGTCACTAATAATAATAGGAACTCAACCTGGCCCCAA
TTCTACAAACTGGCATTGTCGTCTTGCAACATAATTGAGTTCCCAGACTTCTTGAAAATTCAAAATCAGCTCAACTTTTTGTCTCTATCCCACAATAGAA
TTCATGGAGAAATTCCCAAATGGCTGTCAGCAAAGGGGATGCAGTCTTTACAATATTTGGATCTTTCTCACAACTTTCTGACCATTGTAAACGAACTTCC
ACCCAGCTTACAATATCTTGACCTCACTTCCAACTTGCTTCAACAACCATTTCCAATTCTTCCACAATCCATGTACATTCTTTTAATAGCAAATAATAAA
CTCACTGGAGAAATCCCACCTTGGATTTGCAATATCACCACCTTCCAGATCATAAACTTATCTAATAATAGTTTGAGTGGCAATATTCCACAGTGCTTGG
GGAACTTCAGTACTGAACTATCCGTGTTGAATTTGAGGAGCAACAGCTTCCATGGCACCATTCCAGGATCATTTACAGAGGGAAATAAAATAAGAAGTCT
TGATTTGAATGGCAATGAGTTGGAAGGATCGTTACCACTATCCTTAGCTAATTGCAAAATGTTGGAAGTCCTCGATCTGGGAAACAATTACATAAATGAT
AGTTTTCCCCTTTGGCTGCAAACTCTTCCGAAGCTACAGGTTCTGGTGCTCCGCTCTAACAGACTACATGGTTCTATAGGCAATCCCACTGCGATATCTC
CTTTCTCTAGTCTAAGAATCATTGATCTCTCGCACAATGAGTTCATCGGCCTTCTGCCGACACAGTACATTGCAAATTTTCAAGCTATGAAGAAAGTGGA
TGGAGAAGTTAAAGCTACACCAAAGTATATAGGGGAAATATATTATCAGGATTCTATAGTTTTGACGATGAAAGGGACAGAGATTCCAATGGAGAGAATA
CTAACTATCTTTACAACCATTGACCTATCAAGCAACCGATTTGAAGGACAGATTCCAAAGGAGGTTGGACTTCTTAGTTCACTCATAGTCCTTAATATTT
CAAGAAACAGCGTCACTGGTCAAATACCATCATCACTTGGGAACCTGACAGCACTTGAATCACTGGATCTCTCATCAAATGGGCTTGGTGGAGGCATTCC
TTCACAATTAACAAGGCTGACCTTCCTTGCAGTACTAAACCTCTCATATAATCAGCTTGTAGGCCCTATTCCTCATGGCAGTCAATTTGACACCTTTCAG
AATGATTCATATGTTGGAAACTTGCGGTTGTGCGGATTTCCGTTATCTGTGAAATGTAGTGGTGATGTGGCACCACAACCCCCACCATTCCAAGAGAAAG
AAGATCCTGCAAGCTTATTCAATTGGAAGTTTGCCATGATAGGTTATGGATGTGGGCTGGTGATTGGATTGTCCGTGGGATATATTGTATTCACGACAGG
AAAGCCACAATGGTTCGTAAGGAAGGTCGAGGTTGAACAAAAGAAATGGTTAAGAAGACGGACAAAGAGGAATATATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G389100.1 pacid=42789453 polypeptide=Potri.001G389100.1.p locus=Potri.001G389100 ID=Potri.001G389100.1.v4.1 annot-version=v4.1
MSCSGRAGQVIFCLLILMLYCQLTVSQLCPKDESYALLQLKDGFEYDESASLDCSPPYPRMASWKSGTDCCSWDGVACHGVTGHVIALDLSCSGLRGNLS
SNSSLFHLSHLRRLNLAFNYFNRSSIPPEFGMFSSLTHLNLSSTWFSGQVPTEISHLSKLISLDLSLNEPLILEAPAMKMIVQNLTLVREIFLDYINMSS
VDLGSLMNLSSSLTSLSLNLCGLQGQFPENIFHLPNLQLLSLLLNSDLYGRLPVSNWSSSLELLKLGSTSFSGGLPEIIGNLDSIKVLDLGNCAFYGSVP
ASLGNLQQLNQLDLSNNNWTGQIPDVFGNLSKLNSLSLQVGNFSGMLPSSVFNLTELLRLDLSQNQLEGTLPDHICGLDNVTYLDLSYNLLSGTIPSCLF
GLPSLVWFNLNNNHLTGELGEHWSKSLLEIRLESNKINGLIPPSISELVNLTNFDVSSNNLSGIVDLNLFSNMKNLWGLDLSHNSLSVVTNNNRNSTWPQ
FYKLALSSCNIIEFPDFLKIQNQLNFLSLSHNRIHGEIPKWLSAKGMQSLQYLDLSHNFLTIVNELPPSLQYLDLTSNLLQQPFPILPQSMYILLIANNK
LTGEIPPWICNITTFQIINLSNNSLSGNIPQCLGNFSTELSVLNLRSNSFHGTIPGSFTEGNKIRSLDLNGNELEGSLPLSLANCKMLEVLDLGNNYIND
SFPLWLQTLPKLQVLVLRSNRLHGSIGNPTAISPFSSLRIIDLSHNEFIGLLPTQYIANFQAMKKVDGEVKATPKYIGEIYYQDSIVLTMKGTEIPMERI
LTIFTTIDLSSNRFEGQIPKEVGLLSSLIVLNISRNSVTGQIPSSLGNLTALESLDLSSNGLGGGIPSQLTRLTFLAVLNLSYNQLVGPIPHGSQFDTFQ
NDSYVGNLRLCGFPLSVKCSGDVAPQPPPFQEKEDPASLFNWKFAMIGYGCGLVIGLSVGYIVFTTGKPQWFVRKVEVEQKKWLRRRTKRNI
|
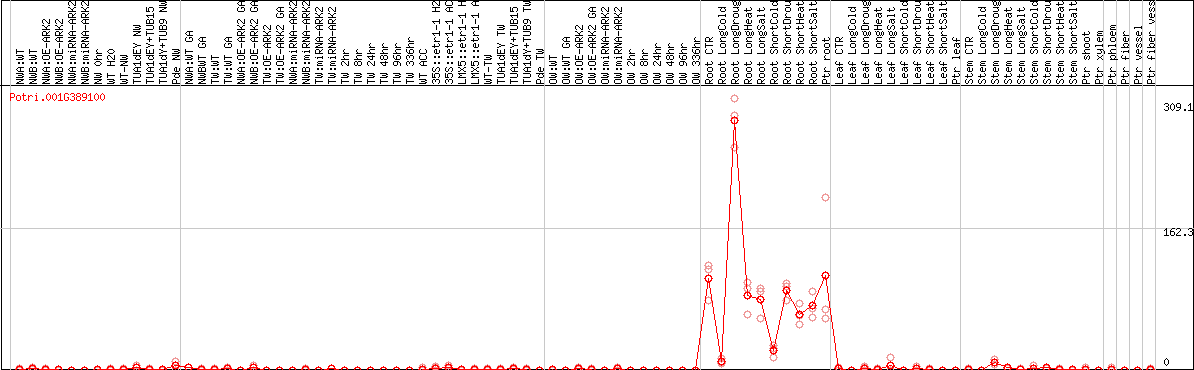
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G389100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
