Potri.001G392801 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G392801.4 pacid=42792131 polypeptide=Potri.001G392801.4.p locus=Potri.001G392801 ID=Potri.001G392801.4.v4.1 annot-version=v4.1
ATGACCAGTTCAGCAATTGATGATTTTCGGGGTGATGGCTTTGAGGGGTCACATCAAGAGCGCTGTATTTTTGCAGATATTTTCTTTGGCAAAGATACTG
GTGGGACAGGTAAGAGGAGTATTGATGCTGGAGTTATTAATTTAAAGAGCCAGGACTGTAAGATTGCAGATCCTTCGCTGCATACTAACAATGAATACTC
AGCTGTCAGTACTCTTTCTTCTCCTATAAGTTTGTCAATCGAGGATTCTGATGTGAATGAGAATTCTGTAGGGGCTACTGCTTCAGGCTGTTTTACAGAA
AGATTTACCTTTGTGGAGGGCACTAGCCAGAATAAGACTGTCAAGCGAATGAAGTTTGCAGTTGATGAACCTTCGGACACTGAACCTGATACATTGAAAG
TTCTGACTTCATCCTTACTGCATAAAGAGATTGTTAGTGGTACAGCTGCTGCAGATATGGATTCACTGAGCCAAACAGTACTATTACATTTAGTCGAATC
ATCCTCTCAGGGTGTTGTATCCACCAGCTATCTGTTAAAGCAACATGCAAAAATAGACAGAAAAGGTGATGCACGCGAGCCAGATGTTCTGAAGTGCAGT
TTGCCAAACTCAGATGGTGTTGCTGGCAAAGCTATTGCTTCACCTGTTTCCCAAGAGAGCTATGCAACAAGGATATTGCTTGCAAGACCTGTAGATGTTG
TTGGGAAACCTGGATCCCCTTTAAATGCAGAGGAAAGAGCGAAGGCATTCAATTCGCCAGGATTGGATGTCTCCATTATCTCGAAGACAGATTCTAAGAT
GGATCCTCGTCCTTTTCTCCAAAGTCATATCACCCGCTTACTTTCAGCTTTGGGTTGGTGTATTGGAAAGCGCAAGAGACCCAGTCGAAAATATATGGAG
TCTATTTATCAATCACCTGAGGGGAGATTAATTCGTGATTTCCCTAAAGTGTGGAGGCTGTGTGGTCAAATTTTGTTCGCAAATGGATACAAGGTGGTGC
AGGAAGGCAATGGTAAGGAATGGGCTGATATAAGTCACTTTTGGTCTGATCTCTCAGATACATTGACTACTATTGAGAAAGACATGGATAAATCAGATCT
TGCCAAAGCCTTGGCACATCAGTGGAGTATTTTAGATCCCTTTGTTAATGTTGTGTTCATTGATCGGAAGGTTGGCGTACTGAGAAAGGGATGTATGGTT
AAAGCTGCACCAAGCTTAGTGAATGATAGAAATGTGAAAAGTGAAGATGGCATTGGAAAGGAGTCTGCTCAGAAAAATTTGTTGGCTTGGCATAGTGATT
CTTGTCCGGTCACTAAAAGTGCATCAACAATTTGTGAAGGAAGCTGCCATGACTGTGATGTACAATCTGGCAACAGGACTTTCTCGGAATGCGGACAAGA
ATCAATGGTGAAAGGTCAGATGGATGTATCTATTCATATGAATGACAGAGAAGGCATGTGTTCTGATGGGATGGGAAATCAATCTTGCAGCATGTGTAAG
GACAGGATAGGCGGTGTGGATATGACTGACTGGCTAACTTGTGGGACAGATTGCACTTCTGCCCAGTTGTGTAGTTGCCAATGTAATGATTTTCCTGTTA
CTGCTGGGAACATTATAGTGCATGGGGCGTCTGAATCTGTTTCCCCTCATCAAGACAGCAACTTGGTTGATCCAGATGATGGCACTGGTCATATGGACTC
CATCCACCATGATGAGCCAACTTCTGCCCAAGTTGTGACTTCAGGAGTTTTGCAGCAATCCAGATTCAGTGAAGAGGAGGGATTACAGTGCATTCAAGCT
TCGAGGTTCAAAACAAGGGATAAAGCAGCTATGAAAAAGATACGTAGAAAGTCCAGAAAAATATCAGAAATCAGATCAACCACATTAAGCCAAAGTGAAA
ATATTGATGTGCTTGGCAATCCATTAGAGTCAAACAAAGTTGAAGAGAAGCTTATCAAAAGAACTAAAAAGATCTGCATGAAGTCCTCCCCTTTGGATAA
TTGCCTACATCAGGGGGTGAAAAATGGTACAAAATTAAAGAGTACGCATGGCAATAGTTATGGTCCCAAATACAAACAAAAGAAAACAACCGGCTGCCAA
ATTGATGATGATGATTTGTTGATTGCAGCCATTATCAAGAACAAAGATTTTAGCCCAGGTGCCACCAGATCTATTTCTAAAAAGAAATCCTGCATATTAA
GAGCTGGCTCAAAGCGCAAACGGAAAAAGGGCGGCTGCAGGCTACTCCCAAGAAATCTGGGCAAACTGGGAAAGCATTATGTGGGTGGTAAGTGGTCAAG
AATGGGATCTAGGACGGTTTTGTCATGGTTGATAGATGCTGGGGTCTTGTCTGTAAAAGATGTAGTCCAGTACCGAAATCTAAAAGACGATTTTGTGATT
AAAGATGGTGTGGTTACCAAGGATGGCATCATGTGTAAGTGCTGCAATATGGTGCTTTCGGTTACCAAGTTTAAGAGTCATGCTGGGTTCAAGCTTAACC
GACCATGTTCAAACCTTTTCATGGAATCTGGAAAGCCTTTTACTTTATGTCAGCTGCAAGCATGGTCTGCTGAATATAAGAGCAGGAAAAGTGGTACCCA
AGTGGTTCGTGCTGATGAAGATGATAAGAATGATGATTCATGTGGGCTGTGCGGTGATGGTGGGGAGTTGATATGCTGTGATAATTGCCCCTCTACTTTT
CATCAAGCTTGCTTGTGTACAGAGGATCTTCCTGAAGGCAGTTGGTATTGCCCGAATTGCACCTGTTGGATATGTGGGGATCTGGTCAATGATAAAGAGG
CTTCAAGTTCTGTTGGTGCTTATAAATGTTTGCAGTGTGAGCACAAATATCATGGGGCATGCCAACAAGGAAAACAAACACATGAAGGACTGGTTTCTGA
TGCTTGGTTTTGTAGTGGAAGCTGTCAAGAGGTGTACTCTGGTCTTCATTCTCGTGTTGGGATCAATAATCCAATAGCTGATGGCTTTTGTTGGACTCTT
CTCAGATGCATTCATGAAGATCAAAAGGTTCTTTCTGCTCAAAGACTTGCCCTCAAGGCAGAATGCAACTCAAAATTAGCTGTTGCTCTTACCATTATGG
AGGAATGCTTTCAATCTATGGTAGACCCAAGAACAGGCATAGACATGATACCCCATGCTTTATACAATTGGGGGTCAGATTTTGCTCGTCTAAATTTCTT
TGGGTTTTACACTGTGGTGTTGGAGAAAGATGATGTGCTTGTATCTGCAGCATCTGTCAGGGTGCATGGAGTAACAGTTGCAGAGATGCCCCTCATTGCA
ACTTGCAGTAACTACCGCCGTCAGGGGATGTGTCGACACCTTATGACTGCTATTGAAGAGATGCTAATTTCCTTCAAGGTGGAAAAGCTTGTTATATCCG
CCATCCCAGATTTAGTTGAGACATGGACTAAGGGTTTTGGCTTCATACCGGTGAGTAAAGATGAAAAGCAAAGTCTGAACAAGATAAACTTCATGGTGTT
CCCTGGAACAATATTGTTGAAAAAACAGTTGTACAAAACAAAAGAAGCAGACACACAATCAGGCTTGGGTGATGCAGCACCTTTAACTGAAGTAGATATT
TGCCCCATGGAGGATCATGTCACTGAGTTGGTGCAGCAATCCAATGAGAACAGCTACCTTGATGAAGTTGGTATCTCAGCAGAGCTCGAACATGGTGAAA
GCCAGAATTTGCAAGAATCTGAACCTAGCAGTGAAAGGCAAGGTAACTACCTTGATGAAGTTGGTATCTCAGCAGAGCTCAAACATGTTGAAAGCCAGAA
ATTGCAAGAATCTGAACCAAGCAGTGAAGGGGAAACTCATGATGGTGCTGAGGGACTAGGACGAGCACCACGCATGGTAACTAACCTTTCGTCTGAAGTA
GGACTCTGTTCTGATGGAATGCCCTTTGTCGAGTCTGACAGAAAATTTTTCCCTAACAAACATGCTGCTAAAAAGGAAGACAAGAAGACGGAAACCCAAG
GTAGTGATTTGCAGGAACAACTTTCAAAGCTGTCCCGTGGAGACTTAGTTCCAGCCCTAGGACGAGGCCCAGGTGAAGTGGCCTGTAATGTTCAATGTAT
TAGTTATGTTCCTTCTCCGGACACACCATCACAGCAAGTCTGTGAGTTAAATGAGAAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G392801.4 pacid=42792131 polypeptide=Potri.001G392801.4.p locus=Potri.001G392801 ID=Potri.001G392801.4.v4.1 annot-version=v4.1
MTSSAIDDFRGDGFEGSHQERCIFADIFFGKDTGGTGKRSIDAGVINLKSQDCKIADPSLHTNNEYSAVSTLSSPISLSIEDSDVNENSVGATASGCFTE
RFTFVEGTSQNKTVKRMKFAVDEPSDTEPDTLKVLTSSLLHKEIVSGTAAADMDSLSQTVLLHLVESSSQGVVSTSYLLKQHAKIDRKGDAREPDVLKCS
LPNSDGVAGKAIASPVSQESYATRILLARPVDVVGKPGSPLNAEERAKAFNSPGLDVSIISKTDSKMDPRPFLQSHITRLLSALGWCIGKRKRPSRKYME
SIYQSPEGRLIRDFPKVWRLCGQILFANGYKVVQEGNGKEWADISHFWSDLSDTLTTIEKDMDKSDLAKALAHQWSILDPFVNVVFIDRKVGVLRKGCMV
KAAPSLVNDRNVKSEDGIGKESAQKNLLAWHSDSCPVTKSASTICEGSCHDCDVQSGNRTFSECGQESMVKGQMDVSIHMNDREGMCSDGMGNQSCSMCK
DRIGGVDMTDWLTCGTDCTSAQLCSCQCNDFPVTAGNIIVHGASESVSPHQDSNLVDPDDGTGHMDSIHHDEPTSAQVVTSGVLQQSRFSEEEGLQCIQA
SRFKTRDKAAMKKIRRKSRKISEIRSTTLSQSENIDVLGNPLESNKVEEKLIKRTKKICMKSSPLDNCLHQGVKNGTKLKSTHGNSYGPKYKQKKTTGCQ
IDDDDLLIAAIIKNKDFSPGATRSISKKKSCILRAGSKRKRKKGGCRLLPRNLGKLGKHYVGGKWSRMGSRTVLSWLIDAGVLSVKDVVQYRNLKDDFVI
KDGVVTKDGIMCKCCNMVLSVTKFKSHAGFKLNRPCSNLFMESGKPFTLCQLQAWSAEYKSRKSGTQVVRADEDDKNDDSCGLCGDGGELICCDNCPSTF
HQACLCTEDLPEGSWYCPNCTCWICGDLVNDKEASSSVGAYKCLQCEHKYHGACQQGKQTHEGLVSDAWFCSGSCQEVYSGLHSRVGINNPIADGFCWTL
LRCIHEDQKVLSAQRLALKAECNSKLAVALTIMEECFQSMVDPRTGIDMIPHALYNWGSDFARLNFFGFYTVVLEKDDVLVSAASVRVHGVTVAEMPLIA
TCSNYRRQGMCRHLMTAIEEMLISFKVEKLVISAIPDLVETWTKGFGFIPVSKDEKQSLNKINFMVFPGTILLKKQLYKTKEADTQSGLGDAAPLTEVDI
CPMEDHVTELVQQSNENSYLDEVGISAELEHGESQNLQESEPSSERQGNYLDEVGISAELKHVESQKLQESEPSSEGETHDGAEGLGRAPRMVTNLSSEV
GLCSDGMPFVESDRKFFPNKHAAKKEDKKTETQGSDLQEQLSKLSRGDLVPALGRGPGEVACNVQCISYVPSPDTPSQQVCELNEK
|
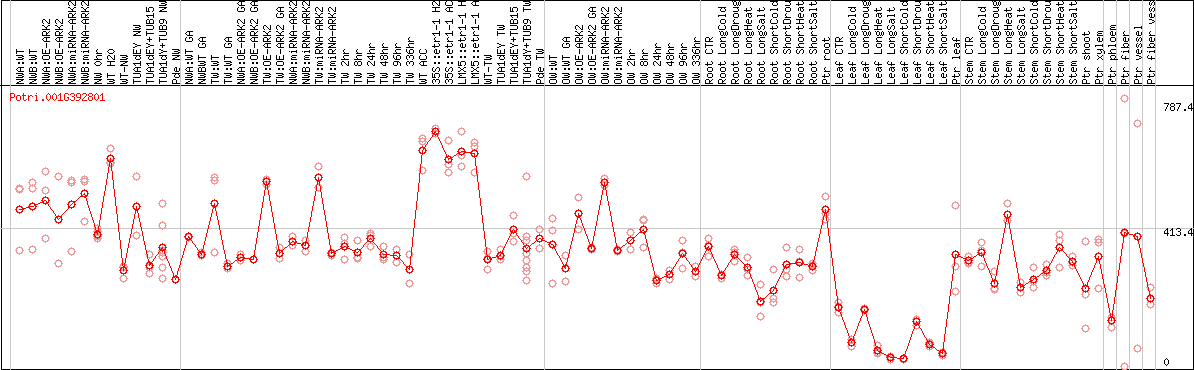
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G392801 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
