External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G27450 436 / 3e-157
Aluminium induced protein with YGL and LRDR motifs (.1)
AT3G15450 376 / 2e-133
Aluminium induced protein with YGL and LRDR motifs (.1.2.3)
AT5G43830 232 / 2e-76
Aluminium induced protein with YGL and LRDR motifs (.1)
AT3G22850 219 / 1e-71
Aluminium induced protein with YGL and LRDR motifs (.1)
AT5G19140 193 / 2e-61
AtAILP1, AILP1
Aluminium induced protein with YGL and LRDR motifs (.1.2)
AT5G10240 52 / 1e-07
ASN3
asparagine synthetase 3 (.1.2)
AT5G65010 52 / 2e-07
ASN2
asparagine synthetase 2 (.1.2)
AT3G47340 48 / 3e-06
AT-ASN1, DIN6, ASN1
DARK INDUCIBLE 6, ARABIDOPSIS THALIANA GLUTAMINE-DEPENDENT ASPARAGINE SYNTHASE 1, glutamine-dependent asparagine synthase 1 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G122100
493 / 2e-179
AT4G27450 428 / 7e-154
Aluminium induced protein with YGL and LRDR motifs (.1)
Potri.011G049900
368 / 4e-130
AT4G27450 383 / 6e-136
Aluminium induced protein with YGL and LRDR motifs (.1)
Potri.004G040600
367 / 1e-129
AT4G27450 366 / 2e-129
Aluminium induced protein with YGL and LRDR motifs (.1)
Potri.008G158400
235 / 1e-77
AT5G43830 388 / 3e-138
Aluminium induced protein with YGL and LRDR motifs (.1)
Potri.010G081600
232 / 2e-76
AT5G43830 391 / 3e-139
Aluminium induced protein with YGL and LRDR motifs (.1)
Potri.008G203400
176 / 1e-54
AT5G19140 397 / 4e-142
Aluminium induced protein with YGL and LRDR motifs (.1.2)
Potri.001G278400
54 / 4e-08
AT3G47340 1018 / 0.0
DARK INDUCIBLE 6, ARABIDOPSIS THALIANA GLUTAMINE-DEPENDENT ASPARAGINE SYNTHASE 1, glutamine-dependent asparagine synthase 1 (.1.2.3)
Potri.009G072900
53 / 6e-08
AT3G47340 1027 / 0.0
DARK INDUCIBLE 6, ARABIDOPSIS THALIANA GLUTAMINE-DEPENDENT ASPARAGINE SYNTHASE 1, glutamine-dependent asparagine synthase 1 (.1.2.3)
Potri.005G075700
43 / 0.0001
AT5G10240 1039 / 0.0
asparagine synthetase 3 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033418
370 / 1e-130
AT4G27450 369 / 2e-130
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10034883
367 / 1e-129
AT4G27450 365 / 6e-129
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10018484
307 / 8e-106
AT4G27450 305 / 3e-105
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10011191
291 / 2e-99
AT4G27450 303 / 2e-104
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10006624
229 / 2e-75
AT5G43830 374 / 2e-132
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10039382
228 / 1e-74
AT5G43830 372 / 6e-132
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10041181
225 / 1e-73
AT5G43830 369 / 1e-130
Aluminium induced protein with YGL and LRDR motifs (.1)
Lus10010505
183 / 2e-57
AT5G19140 392 / 4e-140
Aluminium induced protein with YGL and LRDR motifs (.1.2)
Lus10034038
181 / 2e-56
AT5G19140 389 / 3e-139
Aluminium induced protein with YGL and LRDR motifs (.1.2)
Lus10021899
89 / 5e-20
AT1G04210 609 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF12481
DUF3700
Aluminium induced protein
Representative CDS sequence
>Potri.001G403000.1 pacid=42787705 polypeptide=Potri.001G403000.1.p locus=Potri.001G403000 ID=Potri.001G403000.1.v4.1 annot-version=v4.1
ATGTTGGCAATTTTTCACAAAGCTTTTGCTCACCCACCTGAGGAGCTTAATAGTCCAGCATCTCAAAATGTTACTAAGAAGCCTAAGCTTCCTGAGGAAA
CACTCAATGATTTCCTTTCTCACCGCCCTCAGAAAACTTTCTCCATGAACTTTGGTCAAGCTGCTGTGCTTGCCTATGCTCCCCAAGATAACCCTTTCTC
TCCTCAACAAAAGCTGTTTTGTGGTTTTGATGGTATATACTGCCTTTTCTCTGGAAGCTTGAACAACTTATGCACACTCAATAGGCAGTATGGATTGACA
AAGGGGACTAATGAGGCCATGTTTGTGATAGAAGCTTATAAGACTCTTCGTGATCGAGGTCCTTATCCAGCTGATCAAGTTGTTAAGGATCTTGATGGGA
GCTTTGCTTTCGTTATCTACGATAGCACGGCTGGAAGTGTTTTTGCTGCACTGGGTTCTGATGGTGGAGTTAAGCTGTACTGGGGTATTGCAGCAGATGG
CTCTGTGGTGATATCTGATGATTTGGAAGTCATAAAAGAAAGCTGTGTCAAATCGTTTGCTCCCTTCCCAACAGGATTCATGTTCCATAGTGAAGGAGGC
TTGATGAGCTTTGAGCATCCAATGAACAAAGTTAGAGCAATGCCAAGGACTGACAGTGAAGGATTTCTCTGTGGGGCAAATTTCAAGGTTGATGTCTACA
CAAGGATCAACAGCTTACCCCGTAGAGGAAGTGAAGCTAATTGGACCGAGTGGCAATCACACAGCTAA
AA sequence
>Potri.001G403000.1 pacid=42787705 polypeptide=Potri.001G403000.1.p locus=Potri.001G403000 ID=Potri.001G403000.1.v4.1 annot-version=v4.1
MLAIFHKAFAHPPEELNSPASQNVTKKPKLPEETLNDFLSHRPQKTFSMNFGQAAVLAYAPQDNPFSPQQKLFCGFDGIYCLFSGSLNNLCTLNRQYGLT
KGTNEAMFVIEAYKTLRDRGPYPADQVVKDLDGSFAFVIYDSTAGSVFAALGSDGGVKLYWGIAADGSVVISDDLEVIKESCVKSFAPFPTGFMFHSEGG
LMSFEHPMNKVRAMPRTDSEGFLCGANFKVDVYTRINSLPRRGSEANWTEWQSHS
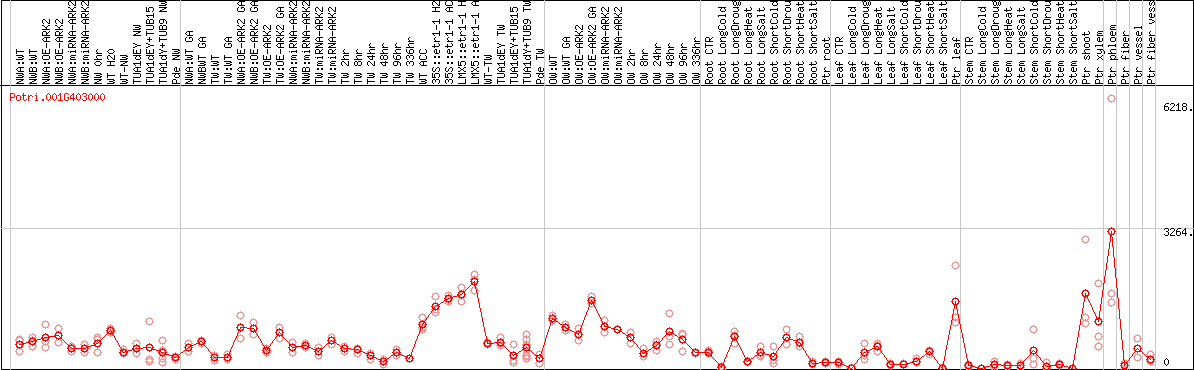
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G403000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.