Pt-CIP7.2 (Potri.001G403700) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-CIP7.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G403700.1 pacid=42791491 polypeptide=Potri.001G403700.1.p locus=Potri.001G403700 ID=Potri.001G403700.1.v4.1 annot-version=v4.1
ATGGATTCAAGAACTTTTCTTGATCATGCTCTCTTTCAACTCACACCCACTAGAACCAGGTGTGATCTGGTGATTTATGCTGGTGGTGTGAATGAGAGAT
TGGCATCTGGGCTTTTAGAACCTTTCCTTCAACACTTAAAAACTGCTAAAGATCAGATATCAAAAGGAGGGTATTCCATTTCTCTTCGTCCTCTGTCTCC
TAATGCCTTCTGGTTCACTAAAGCCACTCTCCAAAGATTTGTGAGGTTTGTCAGCTCACCAGAAGTTCTTGAGAGATTTGTGACAATAGAGACAGAAATA
GAGCAGATTGAGAGTTCAGTTCAATCAAATGAGCTATTAAATGGAGATGCAGAAGGTGCTGCTGGGAATTACCAAAAGTCAACTGTCTCATCTAAGTCCA
AAGGAAATCAAAATGGAAGCAGTGATGGTGTACAAGAAGAAAATTCCAAGGTTCGTCTTCAACGTGCTTTGGAAACTAGGAAAGCTGTTCTTCACAAAGA
GCAAGCAATGGCATATGCCCGTGCCTTGGTCACTGGATTTGAACCAGACTTTATAAATGATCTTATCTGCTTTGCTGATGCTTTTGGAGCTTCACGTTTA
AGGGAAGCATGCATAAATTTCATGGAACTATGCAAGAAGAAGAATCAAGATCGGCTTTGGATGGATGAAATAGCAGCAATGCAGGCATCGCAGCTGGAAT
TGCCTTATCTGGGAACTTCTGGTATTGTACTTTCCGTTGAGGAAAATTACCCTGGTCAAATTGGTGGTCTCTCTGGTGGGAAGCAGAATAGTTCAATGGA
TGCATCAGACTCGGCTACAAGTCCTGGAAGTTTAGAACAAAACCCAGATAGTGGTTTTCCACCATCAGCTCAGATGCAATCAACTGATGGGAAAGCTCAT
ATGCCAATGCCATGGCCAAACCATCATCCTCAGTTCATGCACAATTTTCAAGGTCCTGGATTTCAACAAATGCCCCCGTATCAAGGCTACCTTTTTCCTG
GTATGCGGGTTGGGTCTCCATATTTTCCAGGGAACATGCAATGGCCTCCAAATGTTGATGATTCTAGCCTTGGTCGAGATTGGGAAACTGATGATCGTGA
AAATCGTAAATCATCCTCCAGGAGCAAGAAGAAGTCTTCTCATCGAAAAGAACGGCAAGCTTCTAGTCAAGATCAATCCACTGAGCCCAGTGATTCCAGC
TCAGAGACTGAATCAGATGAACATCTGCAAAGTGACAAAAAGAGGTCTTTGGTGGATAAGATGCACAGGAAAAGGCATGGGAAGAAGTCCTCAAGAAAGG
TTGTCATTCGCAACATCAATTACATTACTTCAATGAAGGATGGAGAAAAGGGCAGCATATCTGACTGCACATCTGATGAGGATGAATTTATTGATGGAGA
ATCTCTCAAGCAGCAAGTACAGGAAGCTGTTGGTTCGTTGGAGAGACGGCATAAATCAACCTCTCGGCAACACAAGAAATCACAGCGCAGCACTATAGAT
GGTTCAAATGATGCTATTGATCAGGAAGGCAAGAATATAATGGCAAATAATCTTGATGGAGAAAAAGGAAAGGACCATTGGGGTGCCTTTCAGAGCCTTC
TCATGCAAGAAAGAGAGCCGAACTCTTTCGGAATTGAACCTGATCCTCCACAGATTCAGAGAGATGACATTACAGCTAAGAGCTATGAGGAGGGAAGGTC
ATTGGAGTTTAACCTGGGATCTGAGGGAATAAGGAAGCAGCGAGCATTATCAGATGATTCATTCATCGCAACAAAGAGGGAGTCTGGTAATGAAGGTGAA
TCTCGTATTGAAAACTTTGAAGCTGGTGCAAATGCCCACCCAATGATAAAGAAAAGGGATAGCACATATGAGGAGTTGTTATTTTCTCAAAGAGCTGGAG
AATCGGGGAACTATCCTATCATTGCTGATTACTCCACTGAATCTCCAATACCCAAAAGTAAAAAAGAAGGGGATTGGTTTATTAGCAGCCAACTGGACAG
GTCAGTAAACATGGATGATCACAGAGACCATAAAGCATTCAGTTGTGATTATGATTCATCATTAACTGGTGAGCATTTCCAAACTGAGAAAAATAAGAAA
GATGTATTAGTGGATGACTCATTCATGATCCAAGCTCGACCTTTAGTAGATGATCAGTCTGATTCACTATTACGAACAGACATAAGCATTGCTCCAGATG
TAGTTGAAGCTACTCAATATGAGAATGGTAGAACAGAGATTTCGCTTGATAAATCTAAGGTATTTGATGTCCATGAACCAGATGACCTTTACATGGTGCT
TGGGCGGGATTCAGTTGCTGAGCATGCTCTATCATCCTGGACTCCAGAAATGGATTATGAAACAAATGCTGTTCAGGATAAGCTTCCTTCTAACAGTATG
GACACCAATGGCAAAAAAAGTGGAAATCCTGGTAAAAAAGTTGCTGGTAAAGAAGCCAGGTCAAAAGTTCCTAATGGATCTCTTGGTAGGAGCAAATCTG
ACATCATGTCAAGGACGAAGAAACCAACTTCAGCAAGCAGAACCACACTTCTGAAGAGCAAATCTGAGAAGGAAGAAGAAAACAGGAAGAGAATGGAGGA
ATTATCAATTGAACGTCAGAAGAGAATTGCAGAAAGGAGTTCTGGTGGCAGTGGTCCAGCGACATCTAAGAGAATCCCTGCAGGGAAAGTACCTACTGCT
ATTTCTATAAAGAATGAGAAGCCTAAAACCCAATCTCCAAGCCAAGATACGAAGAAACCAGTCTTTAGGAGTTCCACTATTGATCGCCTTGCTACTGCTC
GGGCAACCCCAAAGTTGTCATCAACTGAATCAAAAGCTGCCCAACCCAAAAAGGCAACCTTGAAGGCAAATGTTTTATCACAAAAGGCTGCTGGTGCTGG
AAATACTAGTCCAAACACGGTCAAATCTGACATCAACCGAAAGAAGGATGGAACAATAGCCACAGCAGAGAAGCCTGTGGATTTAATACCGACACAAGCA
TCTCAATCTGCAGAAGGAATTAATGATTTCAGAGATATCAAGGAGTTGCAGAGTGTTTCTTCCGCAAAAAATAAGGCAGGAAATATGATTTCAGGAGATA
GCTTAGATGATAAAGGCTGCAATGGTGATTCTCTCCACAAAGACTCATCTGCAGGTGACGAGGGATTCTCTAAAGTAGCTCCAGTTGTTTGTGAATACAT
AGAAACACCTGGTGATCATGGTGAGTACACATCTGAAACAACAATACATCATGTGCCAGAGTCACCAAATAAGGCTCTGAACCTATGCGCTGTCAATATC
AGAGAAAATGGTGGATTCAGTGAAATTTTGGAGTTACCTGAGAAATCTGAAATAGAAATATCGACCCCACCACCTGATGAAATAAACCCAGAACCAATCC
ACTCCAGGAAGAAATGGAACAGTGATGAGAACTCTCCTAAAGCTGCAAAAGGTTTTAGAAAGCTTCTTTTGTTTGGAAGGAAAGGCCGAGCCACTGCTGC
AAACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G403700.1 pacid=42791491 polypeptide=Potri.001G403700.1.p locus=Potri.001G403700 ID=Potri.001G403700.1.v4.1 annot-version=v4.1
MDSRTFLDHALFQLTPTRTRCDLVIYAGGVNERLASGLLEPFLQHLKTAKDQISKGGYSISLRPLSPNAFWFTKATLQRFVRFVSSPEVLERFVTIETEI
EQIESSVQSNELLNGDAEGAAGNYQKSTVSSKSKGNQNGSSDGVQEENSKVRLQRALETRKAVLHKEQAMAYARALVTGFEPDFINDLICFADAFGASRL
REACINFMELCKKKNQDRLWMDEIAAMQASQLELPYLGTSGIVLSVEENYPGQIGGLSGGKQNSSMDASDSATSPGSLEQNPDSGFPPSAQMQSTDGKAH
MPMPWPNHHPQFMHNFQGPGFQQMPPYQGYLFPGMRVGSPYFPGNMQWPPNVDDSSLGRDWETDDRENRKSSSRSKKKSSHRKERQASSQDQSTEPSDSS
SETESDEHLQSDKKRSLVDKMHRKRHGKKSSRKVVIRNINYITSMKDGEKGSISDCTSDEDEFIDGESLKQQVQEAVGSLERRHKSTSRQHKKSQRSTID
GSNDAIDQEGKNIMANNLDGEKGKDHWGAFQSLLMQEREPNSFGIEPDPPQIQRDDITAKSYEEGRSLEFNLGSEGIRKQRALSDDSFIATKRESGNEGE
SRIENFEAGANAHPMIKKRDSTYEELLFSQRAGESGNYPIIADYSTESPIPKSKKEGDWFISSQLDRSVNMDDHRDHKAFSCDYDSSLTGEHFQTEKNKK
DVLVDDSFMIQARPLVDDQSDSLLRTDISIAPDVVEATQYENGRTEISLDKSKVFDVHEPDDLYMVLGRDSVAEHALSSWTPEMDYETNAVQDKLPSNSM
DTNGKKSGNPGKKVAGKEARSKVPNGSLGRSKSDIMSRTKKPTSASRTTLLKSKSEKEEENRKRMEELSIERQKRIAERSSGGSGPATSKRIPAGKVPTA
ISIKNEKPKTQSPSQDTKKPVFRSSTIDRLATARATPKLSSTESKAAQPKKATLKANVLSQKAAGAGNTSPNTVKSDINRKKDGTIATAEKPVDLIPTQA
SQSAEGINDFRDIKELQSVSSAKNKAGNMISGDSLDDKGCNGDSLHKDSSAGDEGFSKVAPVVCEYIETPGDHGEYTSETTIHHVPESPNKALNLCAVNI
RENGGFSEILELPEKSEIEISTPPPDEINPEPIHSRKKWNSDENSPKAAKGFRKLLLFGRKGRATAAN
|
DESeq2's median of ratios [POPLAR]
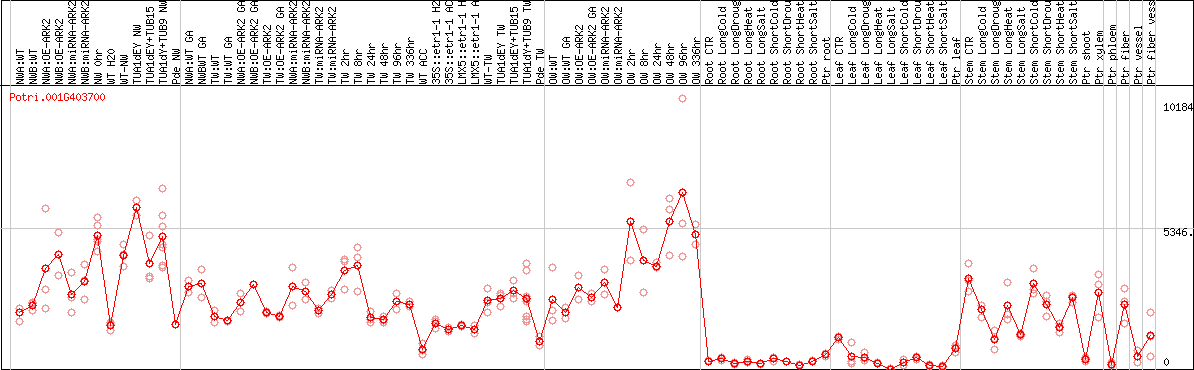
Coexpressed genes
Potri.001G403700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
