Potri.001G411400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G411400.1 pacid=42791638 polypeptide=Potri.001G411400.1.p locus=Potri.001G411400 ID=Potri.001G411400.1.v4.1 annot-version=v4.1
ATGGACACTCTTTCTTCAATGCTTATTACTGCGAATATCCTCCTCCTATTCTCTAGATTCTGTAATACAGCTGATACCCTTGCCTTTTCTCAATCTATTA
GTGATGGAGGGACAGGAACCTTAGTCTCGAAGGATGGAAGTTTTGAGCTTGGTTTCTTCAGTCCTGGAAGTTCCAGGAACCGTTATATGGGAATATGGTA
CAAGAACATCCCAGTTCGAACTGTTGTTTGGGTTGCAAACAGAAACAACCCAATCAATGATTCCTCTGGTTTCTTGTTGATAGACAACACAGGAAATTTT
GTACTTGTCAGCAACAACAATAGTACTGTTGTTTGGTCATCAAGCTTAACGAAAGCAGGCCGGAGAGCAATGGGAGAACTCCTGGATTCTGGGAATCTGG
TCTTAAGAGATGAGAAAGATACCAATTCTGGAAGCTACTTGTGGCAGAGTTTTGACTACCCATCTGATACTATGATTCCAGGAATGAAGCTTGGATGGGG
CTTAAGGACTGGACTTGATCGACGGCTATCAGCATGGAAGGGCCCTGATGATCCATCTCCAGGAGATTTTACTTGGGGAACACAACTACAGGGTAATCCC
GAGTTAGTTATGTGGAAGGGGTCCAAGAAGTACTGTAGAAGTGGTCCATGGAATGGTATCGGGTTCAGTGGTGCGCCGGAGTTGAGGAAAAATCCAGTGT
TTAATTTTGATTTTGTCGACGACGGGGAGGAAGTATATTACACATACAACCTCAAAAATAAATATGTATTTACGAGGGTAGTTATGAACCAAACTACCTA
TATTCGGCAGCGTTACACCTGGAATGAAATCAATCAGACATGGGTACTCTATGCAACTGTGCCTAAAGATTACTGTGACACCTACAATCTTTGTGGTGCC
TACGGAAATTGCATCACCAGCCAGTCGCCAGTTTGTGAATGTTTAGAAAAATTCACGCCTAAATCACCTGAGAGTTGGAATTCAATGGACTGGTCTCAAG
GTTGTGTACGAAATAAGCCACTGGATTGCCAGAAGGAAGATGGATTCGTTATATATGTTGGTTTGAAATTGCCAGATGCTACAAATTCTTGGGTTAACAA
AACTATGAATCTCAAGGAATGCAGGTCAGAATGCTTACAAAATTGTTCTTGTATGGCATATACAGCTGCAGATATTAAGGAAGGTAGTGGCTGTGCTATT
TGGTTTGGCGATCTGATAGATATTAGACAGTTTTCAGCTGCAGGGCAGGAAATATACATTCGCTTGAATGCTTCAGAATCGAGGGCTAAAGCTGCATCTA
AGATAAAGATGGCAGTGGGAATTGCACTATCTATTTTCGTGGCCTGCGGGATACTTTTGGTCGCCTATTACATCTTCAAAAGAAAGGCAAAACTCATAGG
AGGCAACAGAGAAGAAAATGATCAGATAGATAGTGGACCAAAGGAAGACTTGGAGCTACCCCTATTCCAGTTCACTACAATAGCCAAAGCCACCAACGGC
TTTTCATTCAACAACAAGCTAGGAGAAGGTGGTTTTGGACCTGTGTACAAGGGTACTCTAGAAGATGGTCAAGAAATAGCAGCAAAGACGCATTCAAGGA
GTTCTGGACAAGGAATAAATGAGTTCAAAAATGAAGTTATACTGATCACAAAGCTTCAGCACCGAAATCTTGTAAAACTTCTTGGCTGTTGCATTCAAGG
AGAAGAGAAAATTCTGGTTTATGAATACATGCCCAACAAAAGCCTCGATTCCTTCATTTTCGATCAAACAAGAGGCGAACTGTTAGACTGGTCCAAGCGT
TTCAGCATTATATGTGGTATTGCTAGAGGGCTTCTATATCTTCATCAGGATTCCAGATTGAGGATTGTACACAGGGATCTTAAAGCAAGTAACGTTTTAC
TTGATAAGGATATGAACCCAAAAATCTCTGATTTTGGTTTGGCCAGAATGTTTGGTGGAGACCAGACGGAAGGAAACACAACCAGAGTTGTTGGAACTTA
TGGTTATATGGCTCCTGAATATGCTACTGATGGGCTCTTCTCAGTGAAGTCTGATGTTTTTAGCTTTGGAATCTTAATGCTTGAGATAATAAGCGGAAAG
AAGAGTAGAGGATTTTATCATCCGGACCATAGCCTAAGCCTTATTGGACATGCATGGAGACTGTGGAAAGATGGAAAACCTTTGGACTTAATTGAGGCGT
TTCCAGGAGAGTCATGCAATTTATCAGAAGTGATCATGAGATGCATAAACATTAGTCTCTTGTGTGTGCAGCAGCATCCTGATGATAGGCCAAGCATGGC
CACTGTGGTTTGGATGCTAGGAGGTGAGAATACACTGCCTCAGCCAAAGGAACCAGGCTTTTTTAAGGGTAGCGGTCCGTTCGGACCATCTTCTTCATCA
AGCAATATTGAATTATCTTCGAACAATGAAATTACCACCTCACTCTTTTATCCTCGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G411400.1 pacid=42791638 polypeptide=Potri.001G411400.1.p locus=Potri.001G411400 ID=Potri.001G411400.1.v4.1 annot-version=v4.1
MDTLSSMLITANILLLFSRFCNTADTLAFSQSISDGGTGTLVSKDGSFELGFFSPGSSRNRYMGIWYKNIPVRTVVWVANRNNPINDSSGFLLIDNTGNF
VLVSNNNSTVVWSSSLTKAGRRAMGELLDSGNLVLRDEKDTNSGSYLWQSFDYPSDTMIPGMKLGWGLRTGLDRRLSAWKGPDDPSPGDFTWGTQLQGNP
ELVMWKGSKKYCRSGPWNGIGFSGAPELRKNPVFNFDFVDDGEEVYYTYNLKNKYVFTRVVMNQTTYIRQRYTWNEINQTWVLYATVPKDYCDTYNLCGA
YGNCITSQSPVCECLEKFTPKSPESWNSMDWSQGCVRNKPLDCQKEDGFVIYVGLKLPDATNSWVNKTMNLKECRSECLQNCSCMAYTAADIKEGSGCAI
WFGDLIDIRQFSAAGQEIYIRLNASESRAKAASKIKMAVGIALSIFVACGILLVAYYIFKRKAKLIGGNREENDQIDSGPKEDLELPLFQFTTIAKATNG
FSFNNKLGEGGFGPVYKGTLEDGQEIAAKTHSRSSGQGINEFKNEVILITKLQHRNLVKLLGCCIQGEEKILVYEYMPNKSLDSFIFDQTRGELLDWSKR
FSIICGIARGLLYLHQDSRLRIVHRDLKASNVLLDKDMNPKISDFGLARMFGGDQTEGNTTRVVGTYGYMAPEYATDGLFSVKSDVFSFGILMLEIISGK
KSRGFYHPDHSLSLIGHAWRLWKDGKPLDLIEAFPGESCNLSEVIMRCINISLLCVQQHPDDRPSMATVVWMLGGENTLPQPKEPGFFKGSGPFGPSSSS
SNIELSSNNEITTSLFYPR
|
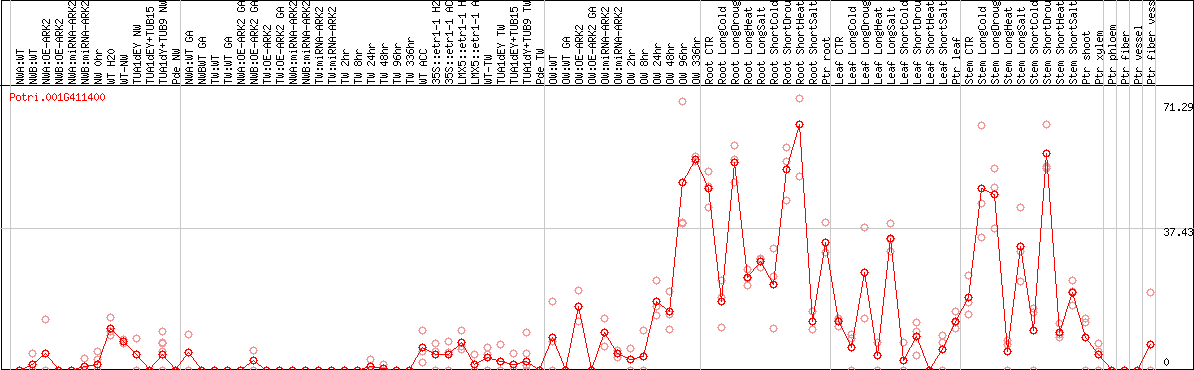
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G411400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
