Potri.001G413700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G413700.1 pacid=42789707 polypeptide=Potri.001G413700.1.p locus=Potri.001G413700 ID=Potri.001G413700.1.v4.1 annot-version=v4.1
ATGAAAGGCATTGCAAGACTTTTCTTCTTCTTCTTCTTCTTCTTCTTCACGGTTTCTAATGGTGCAGATATCGTAGCTGTAAACCAGACAATCTCAGATG
GTGAGACTATAGTTTCAGCTGGGAACAACTTTGAACTGGGATTCTTCAGCCCCAAAAGTTCAAGTTTGCGTTATGTAGGAATATGGTACAAGTTCTCTAA
TGAGACTGTGGTTTGGGTCGCCAACAGAGAAGCACCACTCAATGACACATCTGGAGTTTTGCAAGTTACCAGCAAGGGAATCCTGGTCCTCCATAATTCA
ACAAATGTTGTTTTGTGGTCTACTAACACATCAAGACAGCCCCAGAATCCAGTGGCTCAGCTTTTGAATTCAGGAAATCTTGTTGTTAGGGAAGCAAGCG
ATACCAACGAGGATCATTACTTATGGGAGAGTTTTGACTACCCAGGTAACGTGTTTTTGCCAGGCATCAATTTTGGCAGGAACTTGGTTACTGGCCTTGA
TACGTACTTGGTGTCATGGAAAAGCTCTAATGATCCTTCATTGGGTGATTCAACAACTAGGCTTGATCCTGGTGGATATCCACAGATTTATATCAGGGTT
GGTGAGAACATAGTCTTCCGATCAGGACCATGGAATGGTGTTAGATTCAGTGGGATGCCTAATTTGAAACCGAACCCGATTTACACTTATGGGTTTGTTT
ACAATGAGAAGGAGATTTGTTACAGGTATGATCTGACGGACAGTTCAGTTGTTTCGCATATGTTGCTGACCAATGAAGGAATTTTGCAGCGTTTCACCTG
GACTAATACCACAAGGACATGGAATCTGTACTTGACAGCCCAGATGGATAATTGTGATAGGTATGCAGTATGTGGTGCATATGGAAGTTGTAACATCAAC
AATTCTCCTCCTTGCGCGTGCTTGAAAGGGTTTCAACCAAAATCTCCTCAAGAATGGGAATCTGGGGAATGGTCTGGAGGATGTGTTAGGAAGAATGAAT
CAATTTGCAGGGCAGGAGAGGGGTTTCAGAAGGTACCAAGTGTGAAATTGCCAGATACAAGGACATCCTCGTTTAATTGGACAATGGACTTTGTAGAATG
CCGAAGGGTGTGCTTGATGAATTGTTCTTGTACAGCCTATTCAACTTTGAATATCACAGGTGGATCTGGATGCTTGCTTTGGTTTGAGGAGCTTCTTGAT
ATCAGAGAGTACACTGTAAATGGGCAAGATTTTTACATTAGATTATCTGCCTCTGATTTAGAGCCAACCAGCAGACCTAAGAGAAAGACGCGAGTGTGGA
TTATAGCCATCTGTTCATTGGTTGCTGGAGTTACCATTCTAGGCGTTGGCCTGCTGTTTCTGATGAGGAGGAAGCCTAAGACAGTAGGAAAGATGGTTTC
AATGAGAGAAAGAGACATAATTGATAGCACCGATAAAGATCTAGAGTTGCCAGTATTTGACTTCGCAACCATAGCAATTGCCACTGGCAATTTCTCAGAT
GACAATAAGCTTGGAGAGGGTGGATACGGACCTGTCTACAAGGGCACGCTTAAAGATGGGAAAGAGGTAGCTGTAAAGAGGCTTTCAAAGACTTCCACAC
AAGGACTTGATGAATTCAAAAACGAAGTTATATGCATTGCGAAACTTCAACATCGAAATCTTGTGAAACTTCTTGGTTGCTGCATTGAATCTGAAGAAAA
AATGTTGGTCTATGAATACATGCCCAACGGAAGCTTGGATACCTTCATCTTTGATAAAAATCAGAGCAAATTATTGGAATGGTCTATGAGGCACCACGTT
ATCAATGGGATTGGCCGTGGACTTCTTTATCTACATCAAGATTCCAGGCTAAGGATCATCCATAGAGATCTTAAAGCCAGCAATATATTGCTGGACTTTG
AAATGAACCCAAAAATCTCAGATTTTGGCATGGCCAGGAGTTTTGGTGGAAATGAAATTCAGGGAAACACAAAAAGAGTGGTTGGAACATATGGCTACAT
GGCTCCAGAGTACGCAATTGATGGGCTCTTTTCTATAAAGTCCGATGTATTTAGCTTTGGTGTTTTGGTGCTAGAGATAGTAAACGGGAAGAGAAACAGA
GGATTTTGTCATCCAGACCACAAGCACAACCTTCTTGGTCATGCATGGAGACTTTACAAGGAACAGAAGTCATTCGAGTTGATCGACGAATCTTTAAACA
ATACCTGCGATTTATCTGAAGTCATGCGAGTGATCCAAGTCGGTTTACTGTGTGTGCAACAAGCTCCGGAGGACAGGCCTACTATGTCAACTGTGGTGCT
AATGTTAACCAGTAACATTACATTGCCTGAACCAAAAGAGCCTGGATTTTTCACAGAAAGGAAGTTATTTGATCAAGAGTCTTCATCTAGCAAGGTGGAT
TCGTGTTCCGCAAATGAGATTACTATTACACTGTTAACTGCTCGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G413700.1 pacid=42789707 polypeptide=Potri.001G413700.1.p locus=Potri.001G413700 ID=Potri.001G413700.1.v4.1 annot-version=v4.1
MKGIARLFFFFFFFFFTVSNGADIVAVNQTISDGETIVSAGNNFELGFFSPKSSSLRYVGIWYKFSNETVVWVANREAPLNDTSGVLQVTSKGILVLHNS
TNVVLWSTNTSRQPQNPVAQLLNSGNLVVREASDTNEDHYLWESFDYPGNVFLPGINFGRNLVTGLDTYLVSWKSSNDPSLGDSTTRLDPGGYPQIYIRV
GENIVFRSGPWNGVRFSGMPNLKPNPIYTYGFVYNEKEICYRYDLTDSSVVSHMLLTNEGILQRFTWTNTTRTWNLYLTAQMDNCDRYAVCGAYGSCNIN
NSPPCACLKGFQPKSPQEWESGEWSGGCVRKNESICRAGEGFQKVPSVKLPDTRTSSFNWTMDFVECRRVCLMNCSCTAYSTLNITGGSGCLLWFEELLD
IREYTVNGQDFYIRLSASDLEPTSRPKRKTRVWIIAICSLVAGVTILGVGLLFLMRRKPKTVGKMVSMRERDIIDSTDKDLELPVFDFATIAIATGNFSD
DNKLGEGGYGPVYKGTLKDGKEVAVKRLSKTSTQGLDEFKNEVICIAKLQHRNLVKLLGCCIESEEKMLVYEYMPNGSLDTFIFDKNQSKLLEWSMRHHV
INGIGRGLLYLHQDSRLRIIHRDLKASNILLDFEMNPKISDFGMARSFGGNEIQGNTKRVVGTYGYMAPEYAIDGLFSIKSDVFSFGVLVLEIVNGKRNR
GFCHPDHKHNLLGHAWRLYKEQKSFELIDESLNNTCDLSEVMRVIQVGLLCVQQAPEDRPTMSTVVLMLTSNITLPEPKEPGFFTERKLFDQESSSSKVD
SCSANEITITLLTAR
|
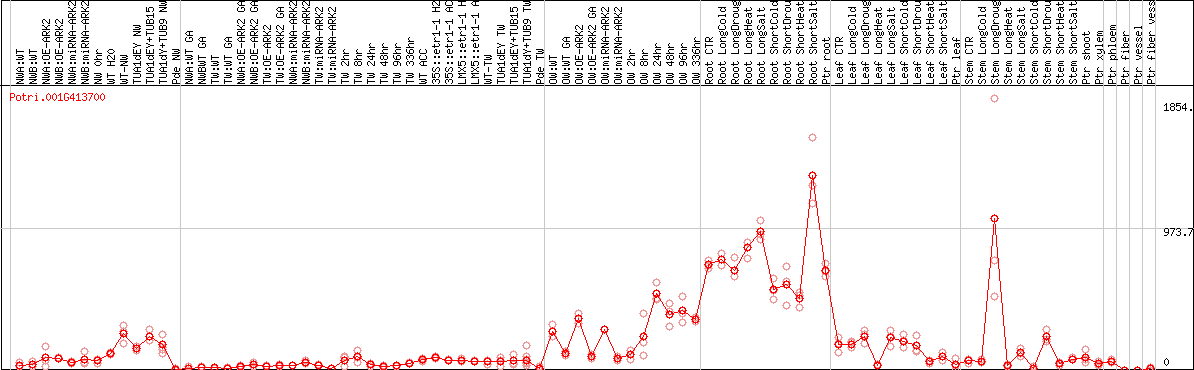
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G413700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
