Potri.001G418100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G418100.1 pacid=42790252 polypeptide=Potri.001G418100.1.p locus=Potri.001G418100 ID=Potri.001G418100.1.v4.1 annot-version=v4.1
ATGGACACTCTTTCTTCAATGCTAATTATTATTGCGAATCTACTCCTTCTATTCTCTAGATTTTGTAATACAGCCAATACCCTTACTTTGTCTCAATCTA
TTCGTGATGGAGGGACAAGAACCTTAGTTTCGAAAGATGGAAGTTTTGAGCTTGGTTTCTTCAGTCCTGGAAGTTCCAGGAACCGTTATGTCGGAATATG
GTACAAGAACATCCCAGTTAGAACTGTTGTTTGGGTTGCAAACAGAAACAACCCAATCAATGATTCCTCTGGCTTCTTGATGTTAGACAACACAGGAAAT
TTTGTACTTGTCAGCAACAACAATAGTACTGTTGTTTGGTCATCAAACTCAAAGAAAGCAGCGCAGAGTGCAATGGGAGAACTCCAGGATTCTGGGAATC
TGGTCTTAAGAGATGAGAAAGACGACAATTCTGGAATCTATTTGTGGCAGAGTTTTGATTATCCATCTGATACGTTGCTTCCAGGAATGAAGCTTGGATG
GGACTTACGGATTGGACTTGATCGGCGGCTATCAGCTTGGAAGAGCCCTGATGATCCATCTTCTGGAGATTTTACTTGGGGAACACAACTACAGAGTAAT
CCTGAGTTAGTTATGTGGAAGGGGTCCAAGAAGTACTATAGAAGTGGTCCATGGAATGGTATCGGGTTCAGTGGTGGGCTGGCGTTAAGGATTAATCCAG
TTTTTTATTTTGATTTTGTCGACGACGGGGAGGAGGTTTATTACACATACAACCTCAAAAATAAATCTTTAATCACGAGGATAGTTATGAACCAAACTAC
CTATTTTCGGCAGCGTTACACCTGGAATGAAATCAATCAGACATGGGTACTCTATGCAACTGTGCCGAGAGATTACTGTGACACTTACAATCTTTGTGGT
GCCTATGGAAATTGCATCATGAGCCAGTCACCAGTTTGTCAATGTTTAGAAAAATTCACGCCTAGATCACCTGAGAGTTGGAATTCAATGGACTGGTCTA
AAGGGTGTGTACGAAATAAGCCACTAGATTGCCAGAAGGGAGATGGATTCGTTAAATATGTTGGTTTGAAATTGCCAGATGCTACAAATTCTTGGGTAAA
CAAAACTATGAATCTCAAGGAATGCAGGTCAAAATGCTTACAAAACTGTTCTTGTATGGCATATACAGCTACAAATATAAAGGAACGCAGTGGCTGTGCT
GTTTGGTTTGGCGATCTGATAGATATTAGACAGTTTCCAGCTGCAGGGCAGGAAATATACATTCGCATGAATGCTTCAGAATCAAAGGCAAAAGCTGCAT
CAAACATAAAGATGGCAGTGGGAATTGCACTATCTATTTCCGTGGTCTGCGGGATGCTTTTGGTCGCCTATTACATCTTCAAAAGAAAGGCAAAACTCAT
AGGAGGCAACAGAGAAGAAAATGATCAGATAGATAGTGGACCAAAGGAAGACTTGGAGCTACCCCTATTCCAGTTCACTACAATAGCCAAAGCCACCAAC
GGCTTTTCATTCAACAACAAGCTAGGAGAAGGTGGTTTTGGACCTGTGTACAAGGGTACTCTAGAAGATGGCCAAGAAATAGCAGCAAAAACGCTATCAA
GGAGTTCTGGACAAGGATTAAATGAGTTCAAAAATGAAGTTATACTGATTACAAAGCTTCAGCACCGAAATCTTGTAAAACTTCTTGGCTGTTGCATTCA
AGGAGAAGAGAAAATTCTGGTTTATGAATACATGCCCAACAAAAGCCTCGATTCCTTCATTTTCGATCAAACAAGAGGCAAACTGTTAGACTGGTCCAAG
CGTTTCAGCATTATATGTGGTATTGCTAGGGGGCTTCTATATCTTCATCAGGATTCCAGATTGAGGATTGTACACAGGGATCTTAAAGCAAGCAACGTTT
TACTTGATAAGGATATGAACCCAAAAATCTCTGATTTTGGTTTGGCCAGAATGTTTGGTGGAGACCAGACGGAAGGAAACACAACAAGAGTTGTCGGGAC
TTATGGTTACATGGCTCCTGAATATGCTACTGATGGGCTCTTCTCAGTGAAGTCTGATGTTTTTAGCTTTGGAATCTTAATGCTTGAGATAATAAGTGGA
AAGAAGAGTAGAGGATTTTGTCATCCAGACCATAGCCTAAGCCTTATTGGACATGCATGGAGATTGTGGAAAGATGGAAAACCTTTGGGCTTAATTGAGG
CGTTTCCAGGAGAGTCATGCAATTTATCAGAAGTGATCATGAGATGCATAAACATTAGTCTCTTGTGTGTGCAGCAGCATCCTGATGATAGGCCAAGCAT
GGCCACTGTGGTTTGGATGCTAGGAGGTGAGAATACACTGCCTCAGCCAAAGGAACCAGGCTTTTTTAAGGGTAGCGGTCCGTTCCGACCATCTTCTTCA
TCAAAAAATACTGAATTATTTTCGAACAATGAAATTACTTCCTCACTCTTGTATCCTCGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G418100.1 pacid=42790252 polypeptide=Potri.001G418100.1.p locus=Potri.001G418100 ID=Potri.001G418100.1.v4.1 annot-version=v4.1
MDTLSSMLIIIANLLLLFSRFCNTANTLTLSQSIRDGGTRTLVSKDGSFELGFFSPGSSRNRYVGIWYKNIPVRTVVWVANRNNPINDSSGFLMLDNTGN
FVLVSNNNSTVVWSSNSKKAAQSAMGELQDSGNLVLRDEKDDNSGIYLWQSFDYPSDTLLPGMKLGWDLRIGLDRRLSAWKSPDDPSSGDFTWGTQLQSN
PELVMWKGSKKYYRSGPWNGIGFSGGLALRINPVFYFDFVDDGEEVYYTYNLKNKSLITRIVMNQTTYFRQRYTWNEINQTWVLYATVPRDYCDTYNLCG
AYGNCIMSQSPVCQCLEKFTPRSPESWNSMDWSKGCVRNKPLDCQKGDGFVKYVGLKLPDATNSWVNKTMNLKECRSKCLQNCSCMAYTATNIKERSGCA
VWFGDLIDIRQFPAAGQEIYIRMNASESKAKAASNIKMAVGIALSISVVCGMLLVAYYIFKRKAKLIGGNREENDQIDSGPKEDLELPLFQFTTIAKATN
GFSFNNKLGEGGFGPVYKGTLEDGQEIAAKTLSRSSGQGLNEFKNEVILITKLQHRNLVKLLGCCIQGEEKILVYEYMPNKSLDSFIFDQTRGKLLDWSK
RFSIICGIARGLLYLHQDSRLRIVHRDLKASNVLLDKDMNPKISDFGLARMFGGDQTEGNTTRVVGTYGYMAPEYATDGLFSVKSDVFSFGILMLEIISG
KKSRGFCHPDHSLSLIGHAWRLWKDGKPLGLIEAFPGESCNLSEVIMRCINISLLCVQQHPDDRPSMATVVWMLGGENTLPQPKEPGFFKGSGPFRPSSS
SKNTELFSNNEITSSLLYPR
|
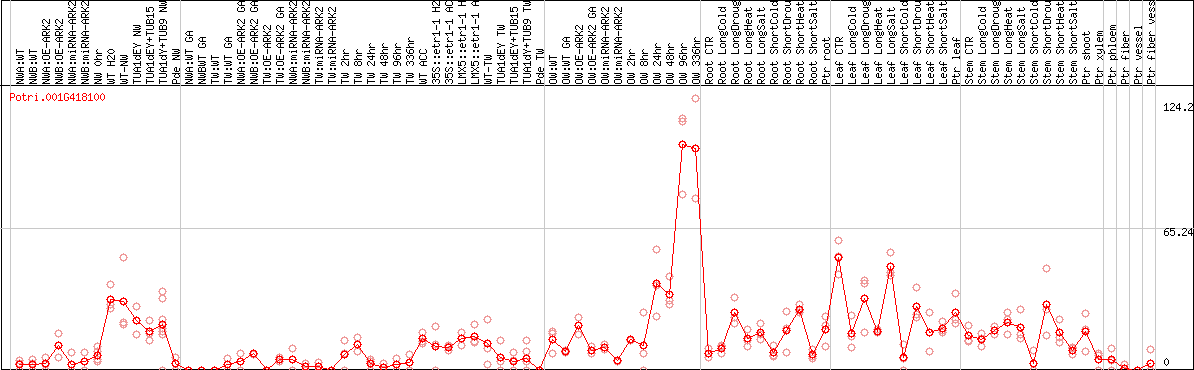
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G418100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
